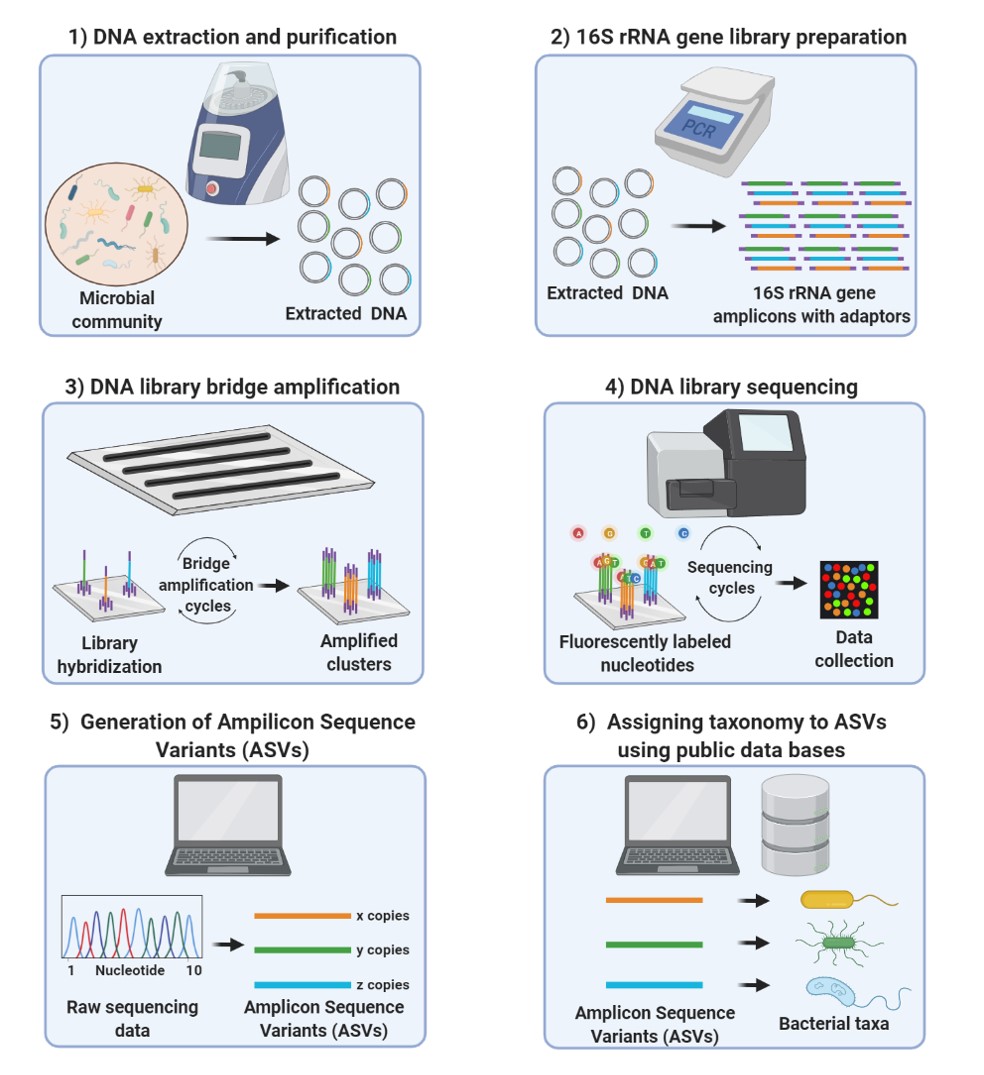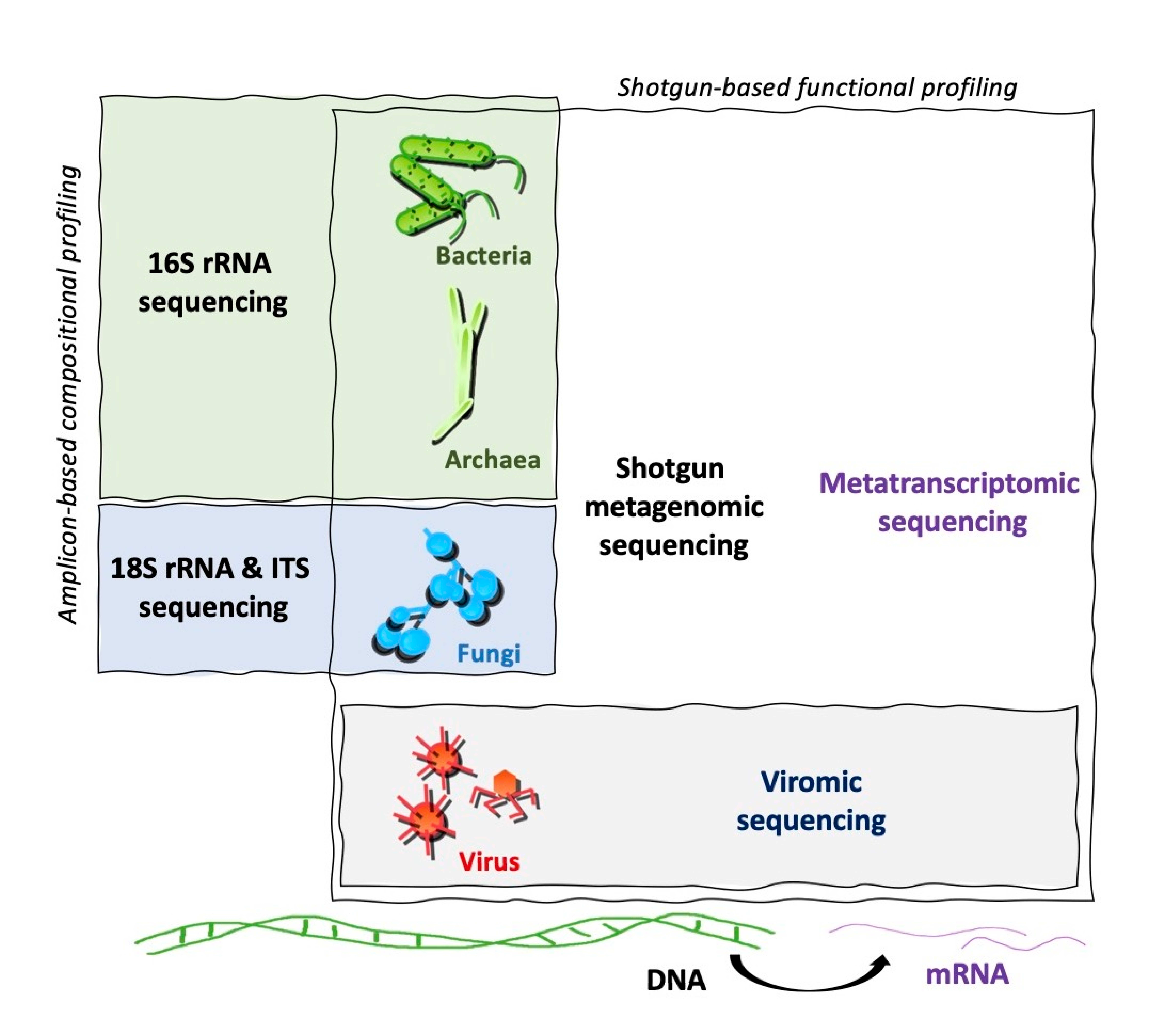
Biomolecules | Free Full-Text | An Introduction to Next Generation Sequencing Bioinformatic Analysis in Gut Microbiome Studies
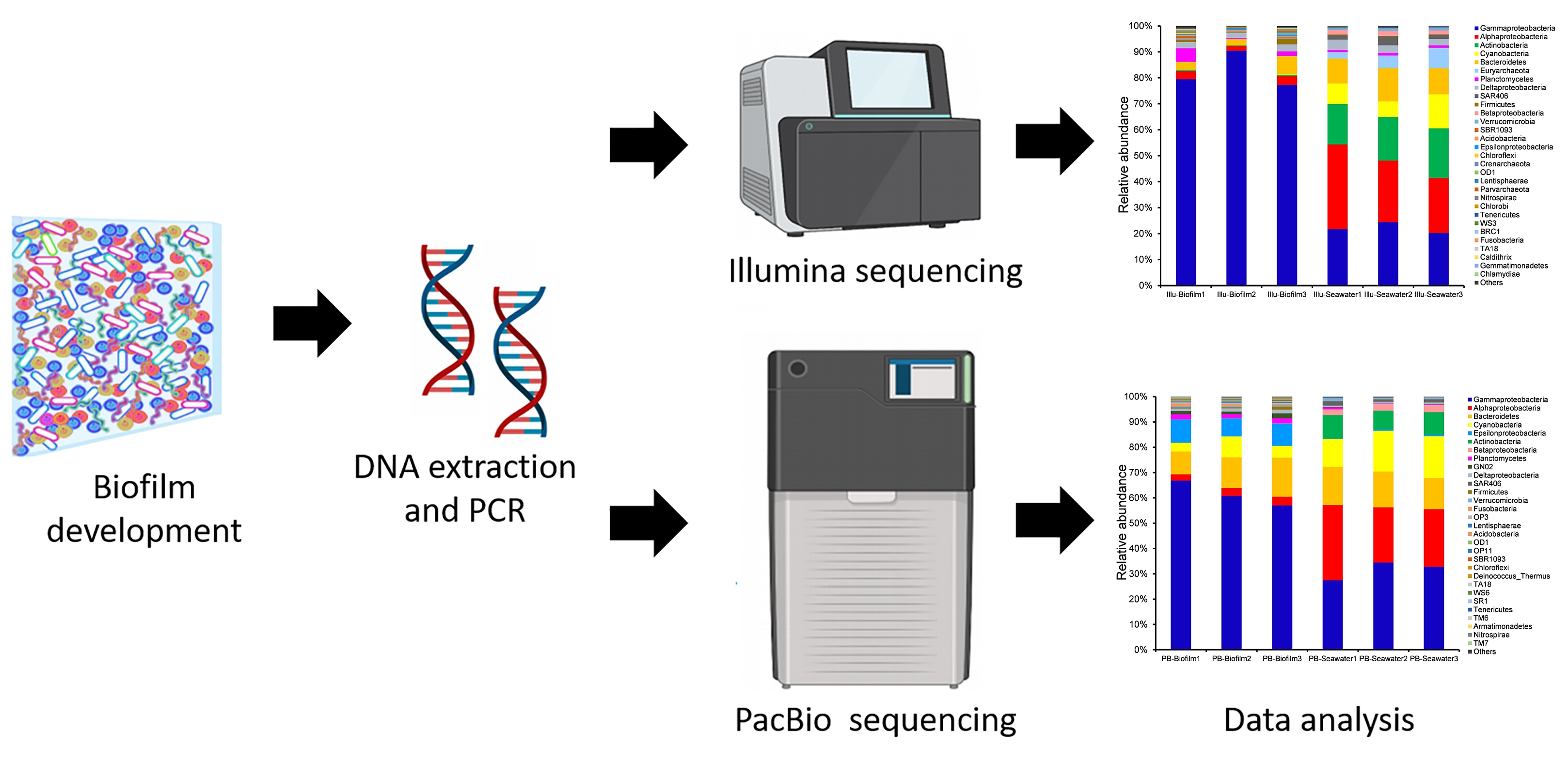
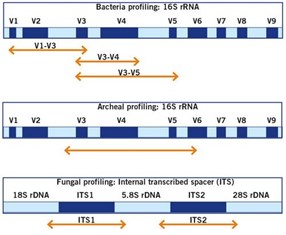
Genes | Free Full-Text | Microbial Richness of Marine Biofilms Revealed by Sequencing Full-Length 16S rRNA Genes
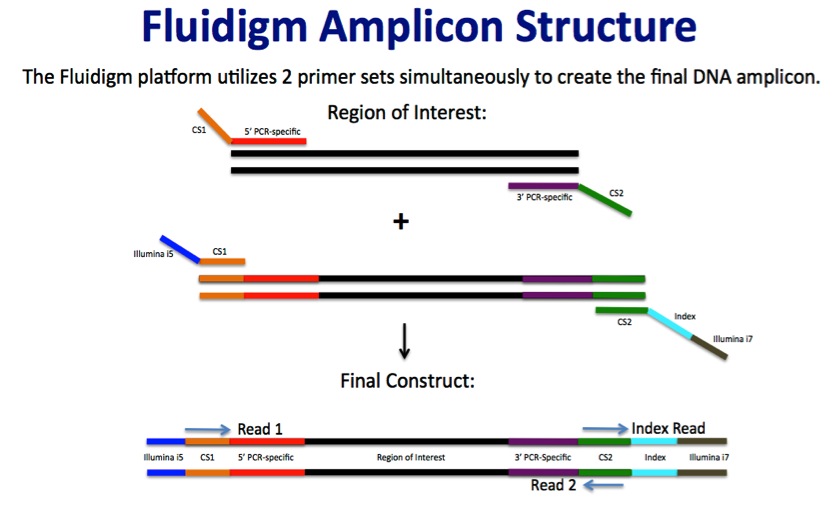
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech

Analysis of the mouse gut microbiome using full-length 16S rRNA amplicon sequencing | Scientific Reports
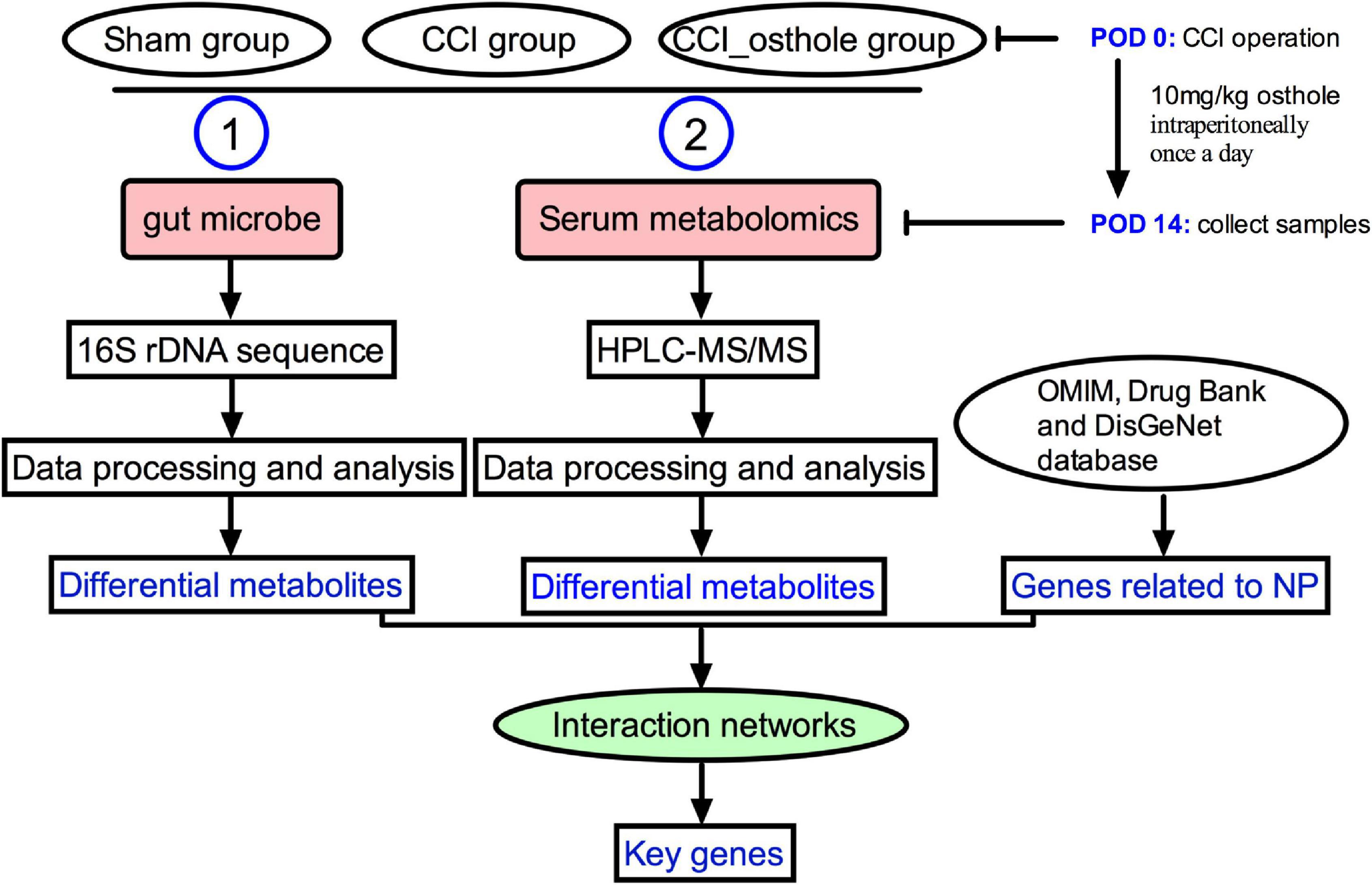
Frontiers | Integrated 16S rRNA Gene Sequencing and Metabolomics Analysis to Investigate the Important Role of Osthole on Gut Microbiota and Serum Metabolites in Neuropathic Pain Mice
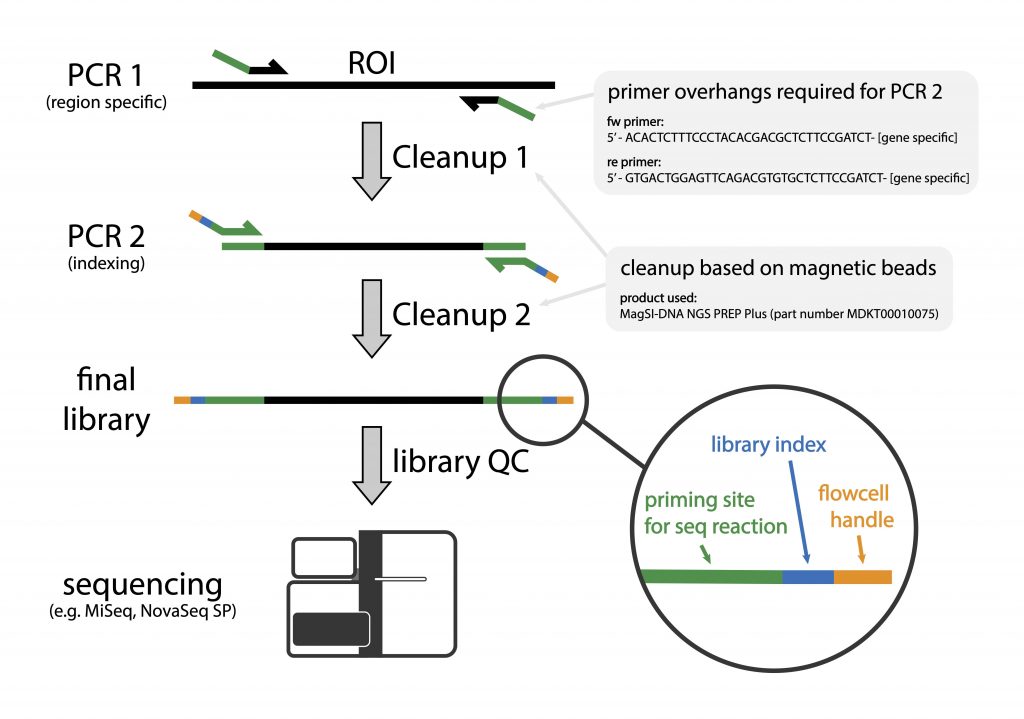
Highly multiplexed gene amplicon sequencing | Joint Microbiome Facility of the Medical University of Vienna and the University of Vienna
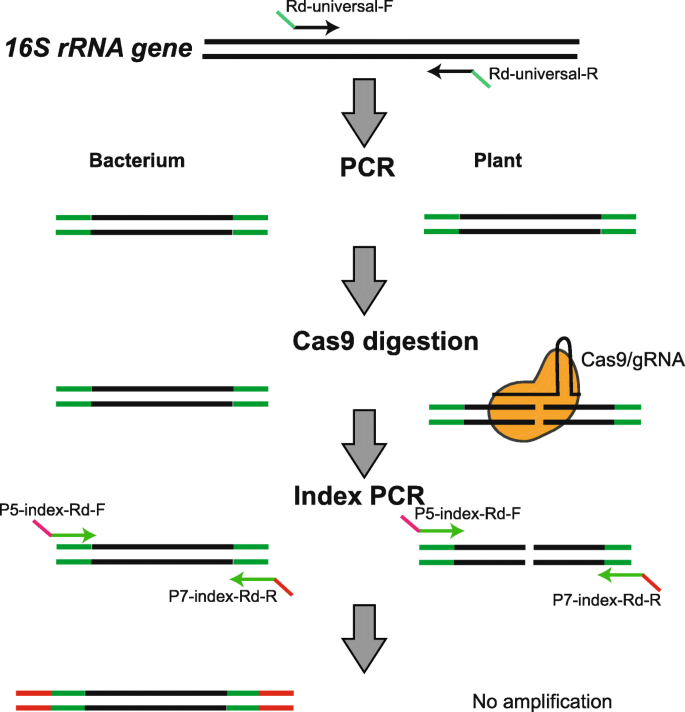
Engineering CRISPR/Cas9 to mitigate abundant host contamination for 16S rRNA gene-based amplicon sequencing | Microbiome | Full Text

DNA extraction protocols cause differences in 16S rRNA amplicon sequencing efficiency but not in community profile composition or structure - Rubin - 2014 - MicrobiologyOpen - Wiley Online Library

Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research - ScienceDirect

Efficient Nucleic Acid Extraction and 16S rRNA Gene Sequencing for Bacterial Community Characterization | Protocol
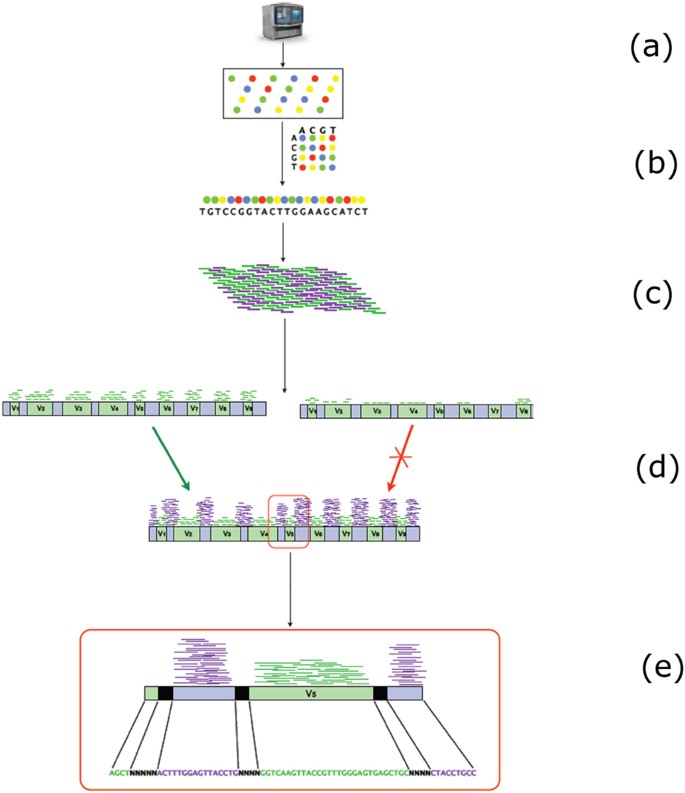
Direct 16S rRNA-seq from bacterial communities: a PCR-independent approach to simultaneously assess microbial diversity and functional activity potential of each taxon | Scientific Reports
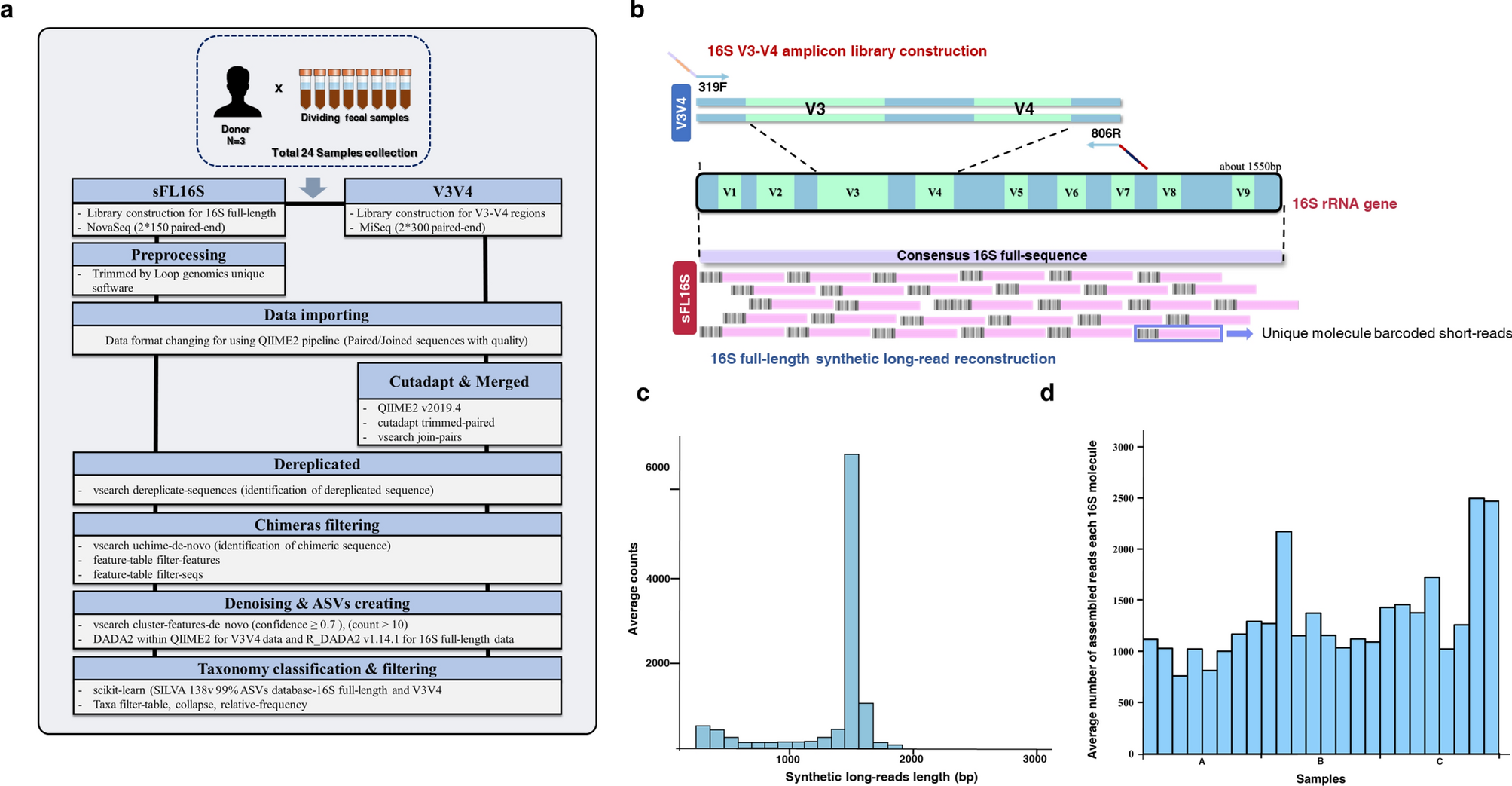
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports
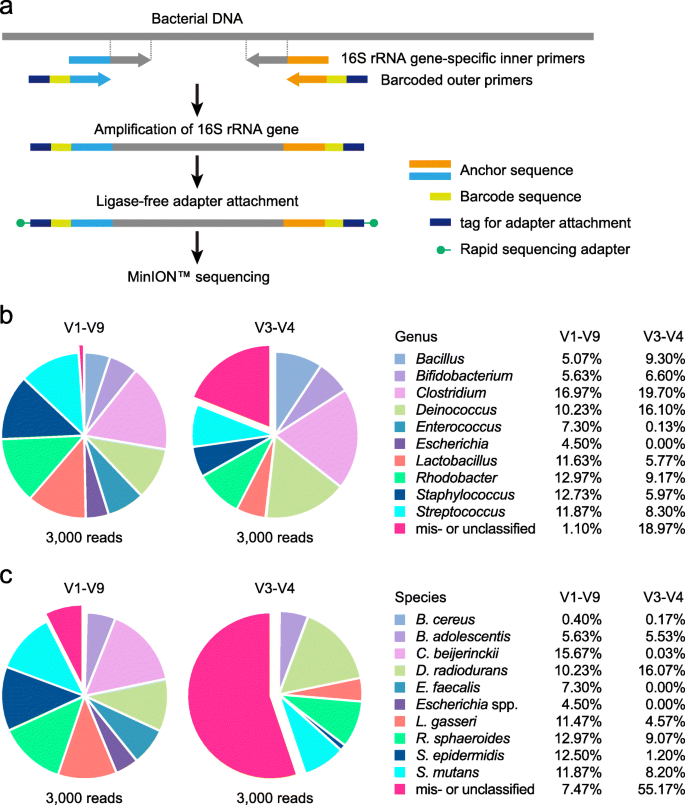
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text
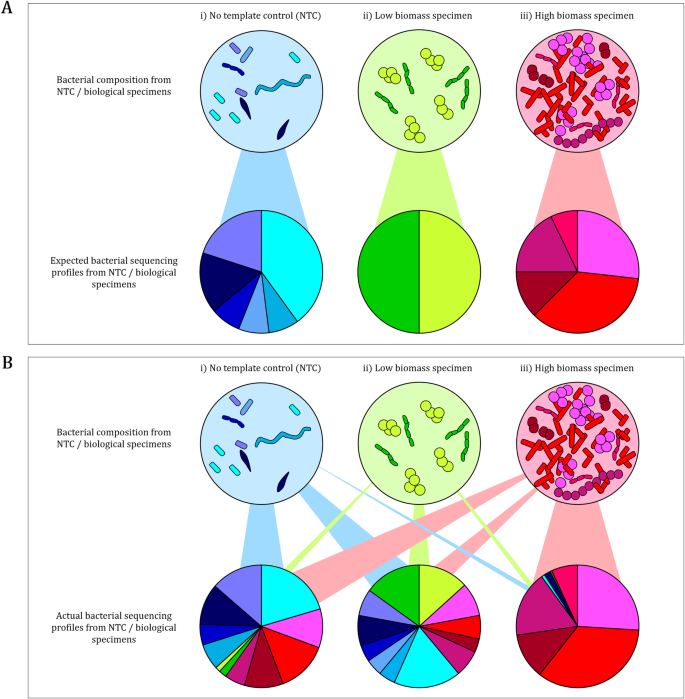
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text

![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)