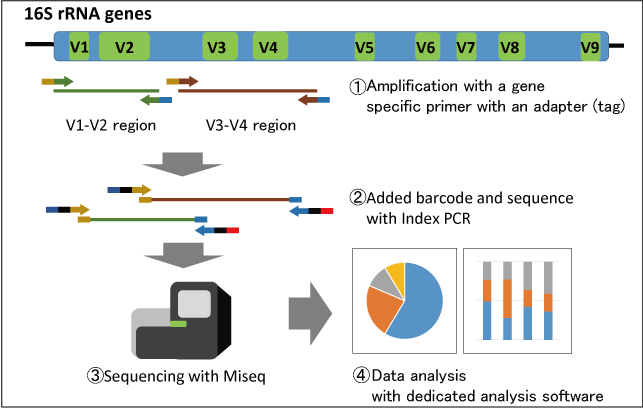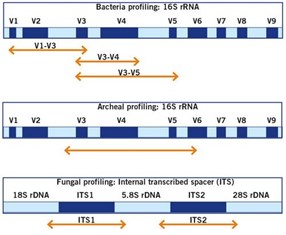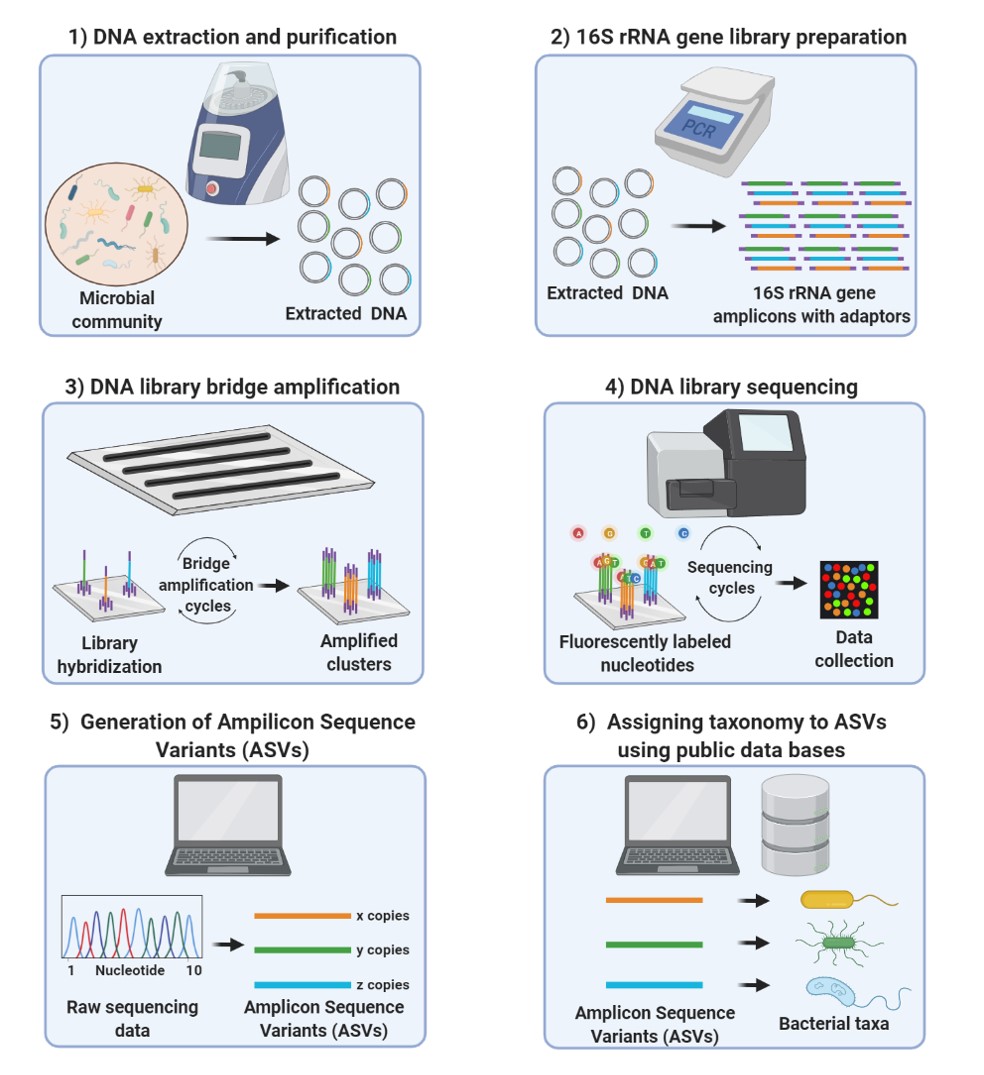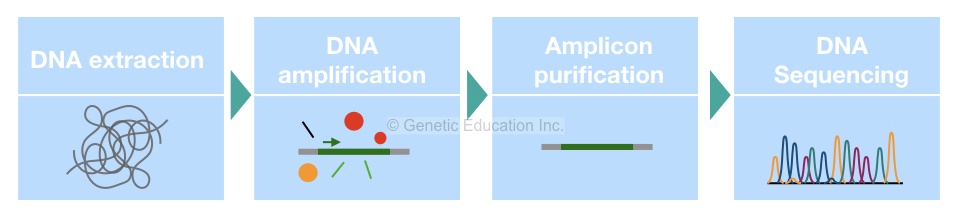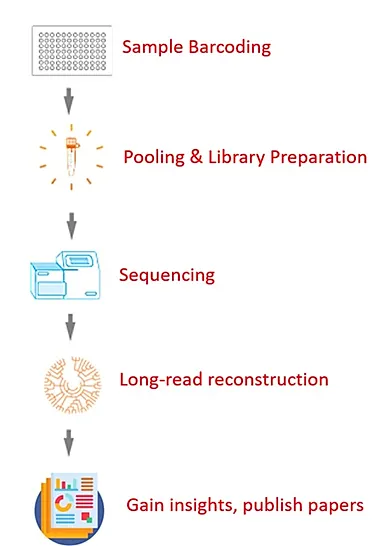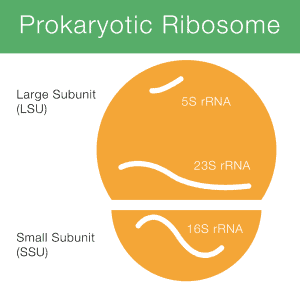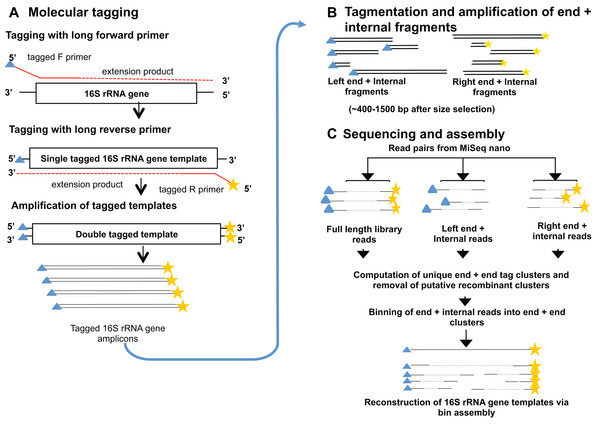
Method of interest: precision sequencing of near full 16S rRNA genes on a MiSeq (from @Cath_B & @koadman) Â – microBEnet: the microbiology of the Built Environment network
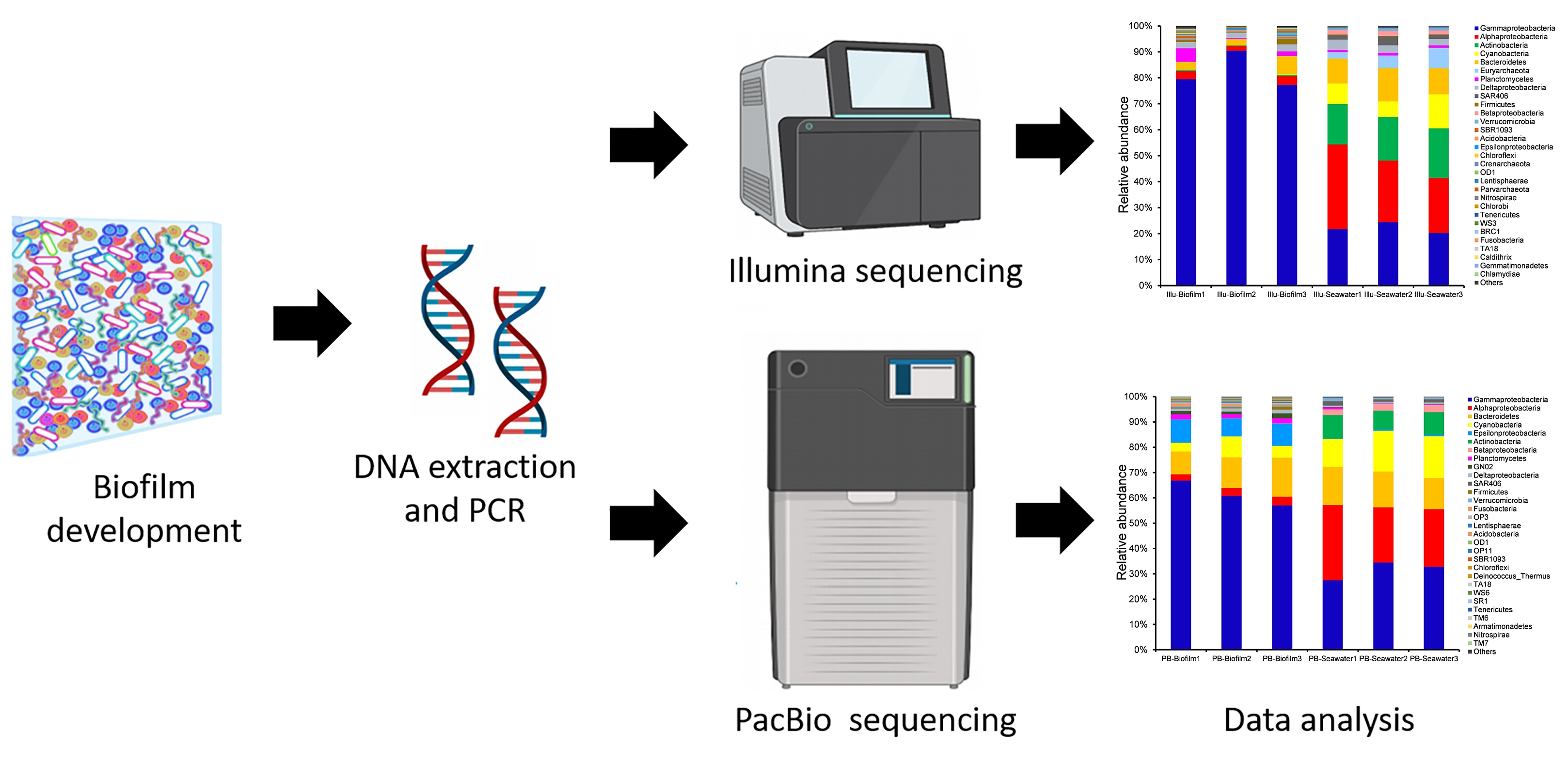
Genes | Free Full-Text | Microbial Richness of Marine Biofilms Revealed by Sequencing Full-Length 16S rRNA Genes
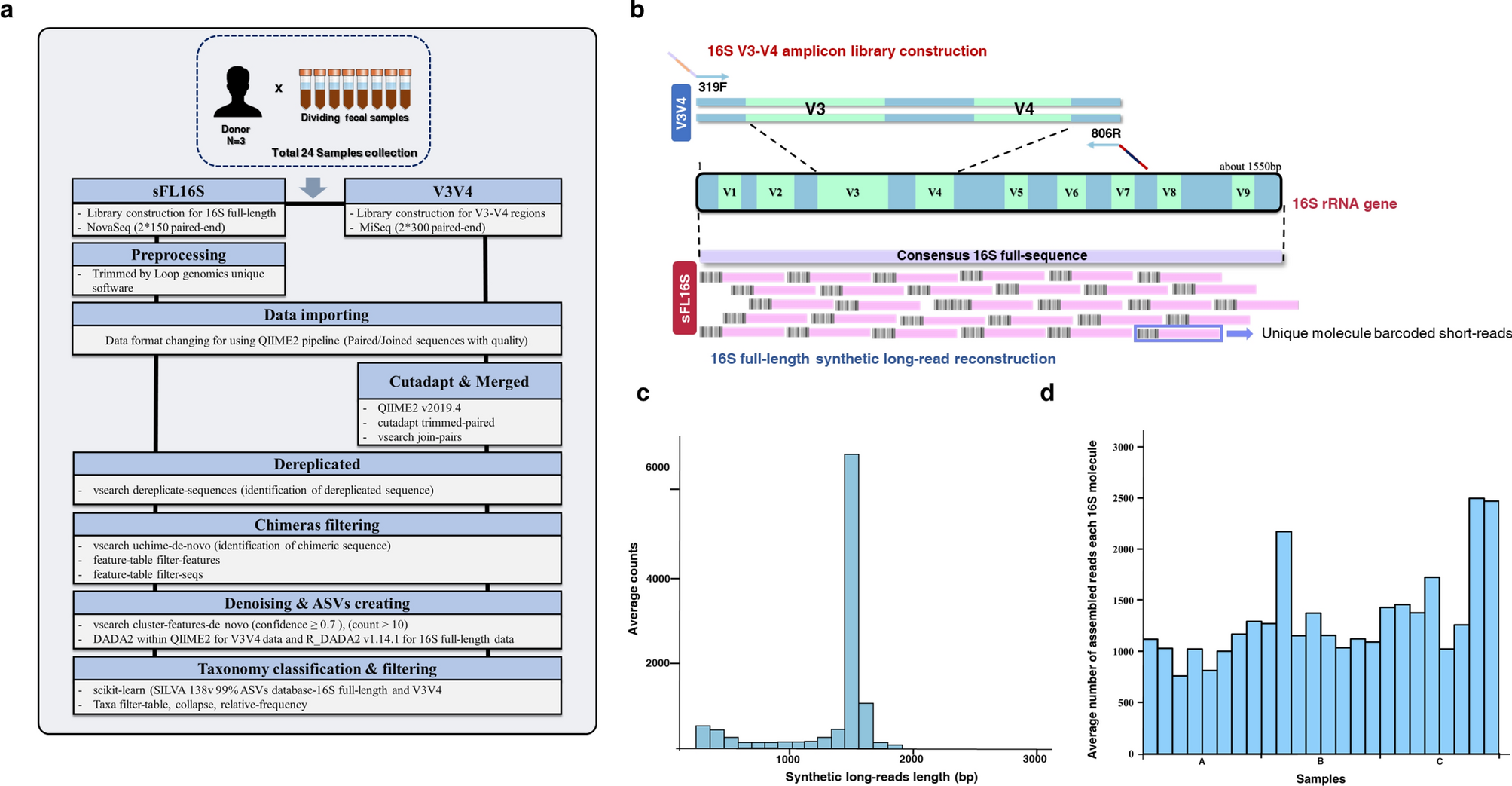
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports

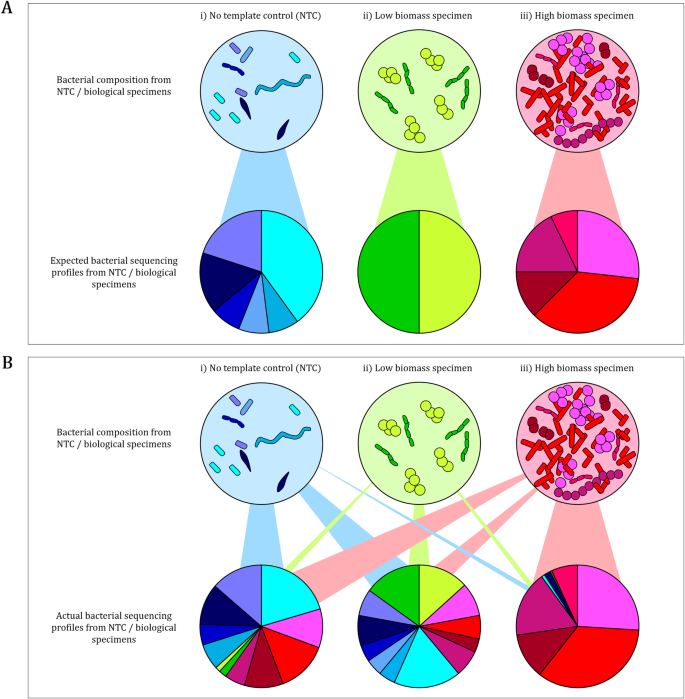
Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research - ScienceDirect
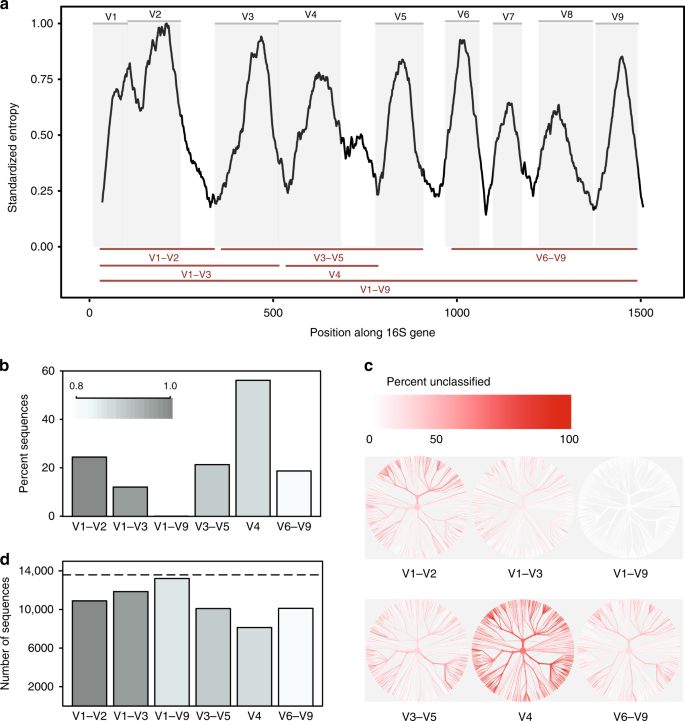
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications
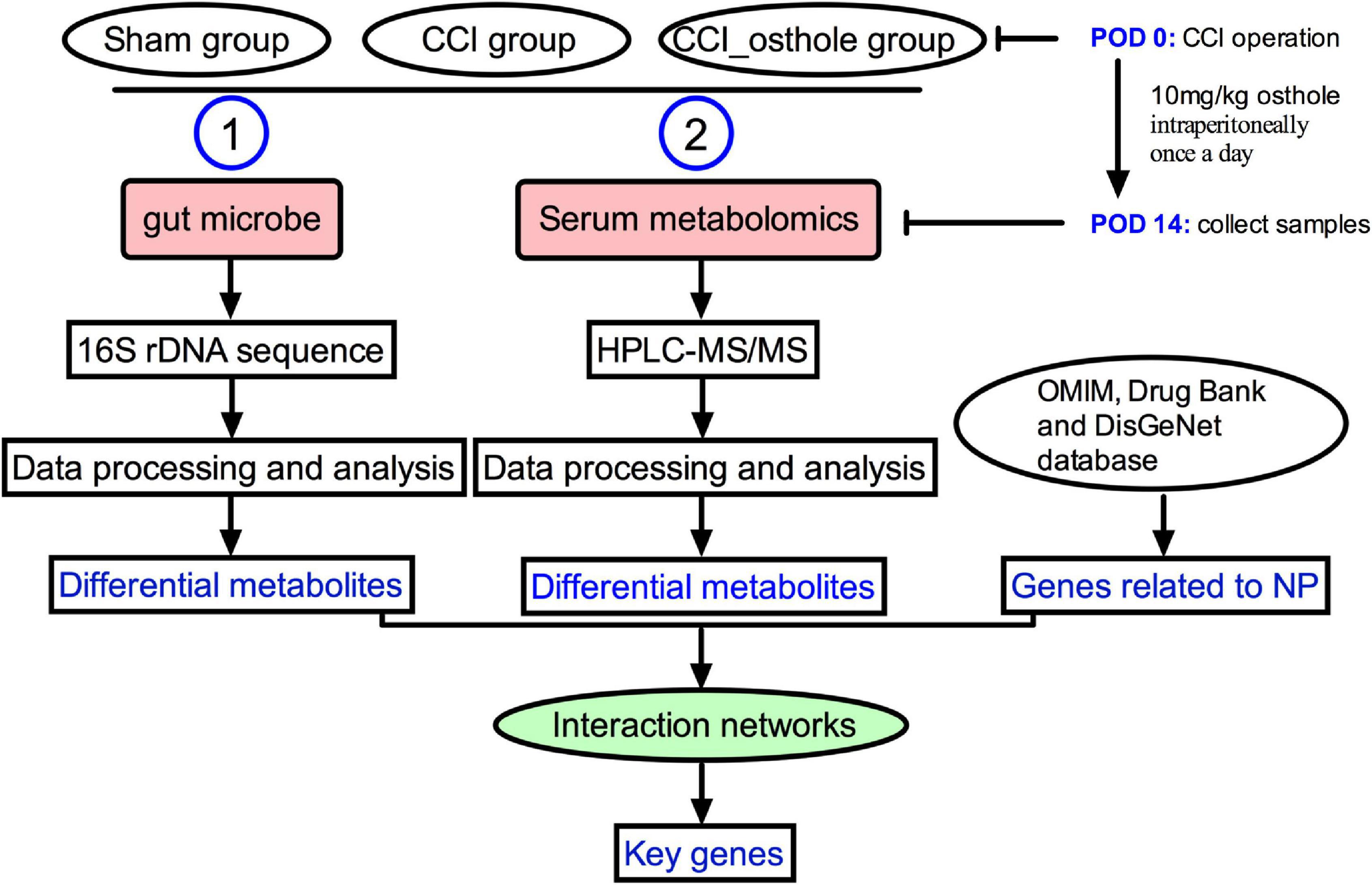
Frontiers | Integrated 16S rRNA Gene Sequencing and Metabolomics Analysis to Investigate the Important Role of Osthole on Gut Microbiota and Serum Metabolites in Neuropathic Pain Mice