Nucleosome Positioning on Large Tandem DNA Repeats of the '601' Sequence Engineered in Saccharomyces cerevisiae - ScienceDirect

Nucleosome Positioning on Large Tandem DNA Repeats of the '601' Sequence Engineered in Saccharomyces cerevisiae - ScienceDirect

Nucleosomal DNA constructs based on the Widom 601 sequence to study the... | Download Scientific Diagram

Asymmetric Unwrapping of Nucleosomes under Tension Directed by DNA Local Flexibility - ScienceDirect
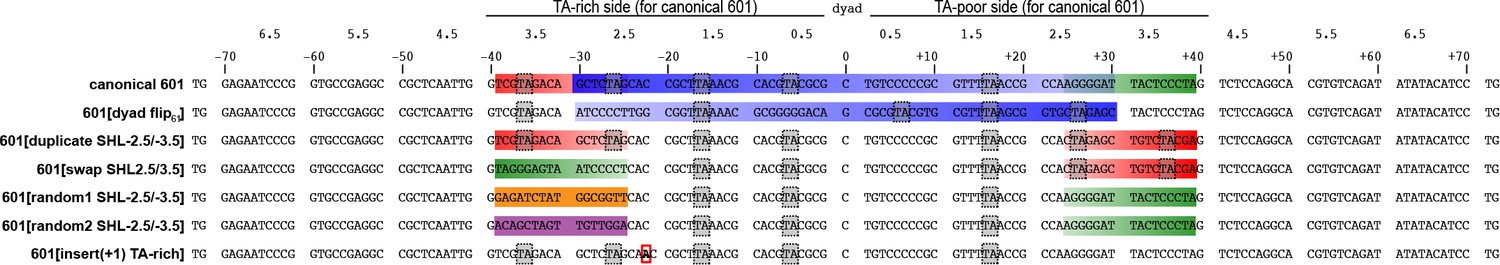

Figures and data in A twist defect mechanism for ATP-dependent translocation of nucleosomal DNA | eLife

Nucleosome positioning on large tandem DNA repeats of the '601' sequence engineered in Saccharomyces cerevisiae | bioRxiv

Nucleosomal DNA constructs based on the Widom 601 sequence to study the... | Download Scientific Diagram
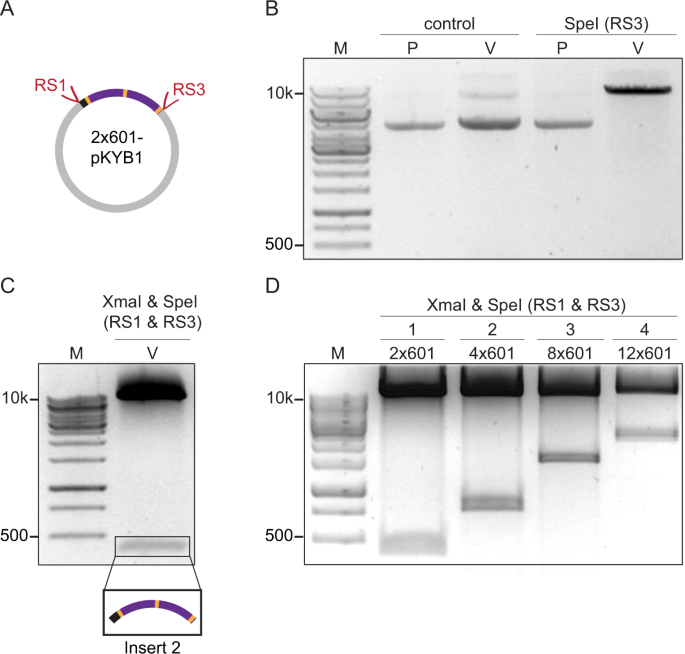
Constructing arrays of nucleosome positioning sequences using Gibson Assembly for single-molecule studies | Scientific Reports

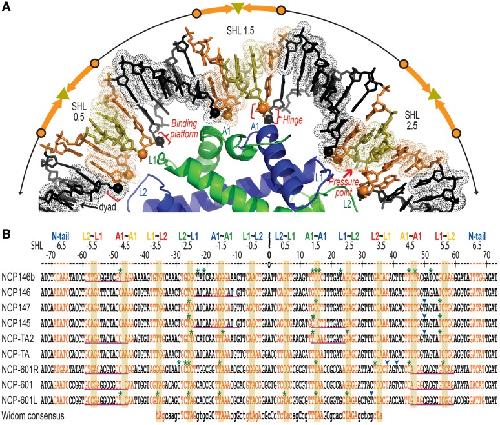
Crystal Structures of Nucleosome Core Particles Containing the '601' Strong Positioning Sequence - ScienceDirect
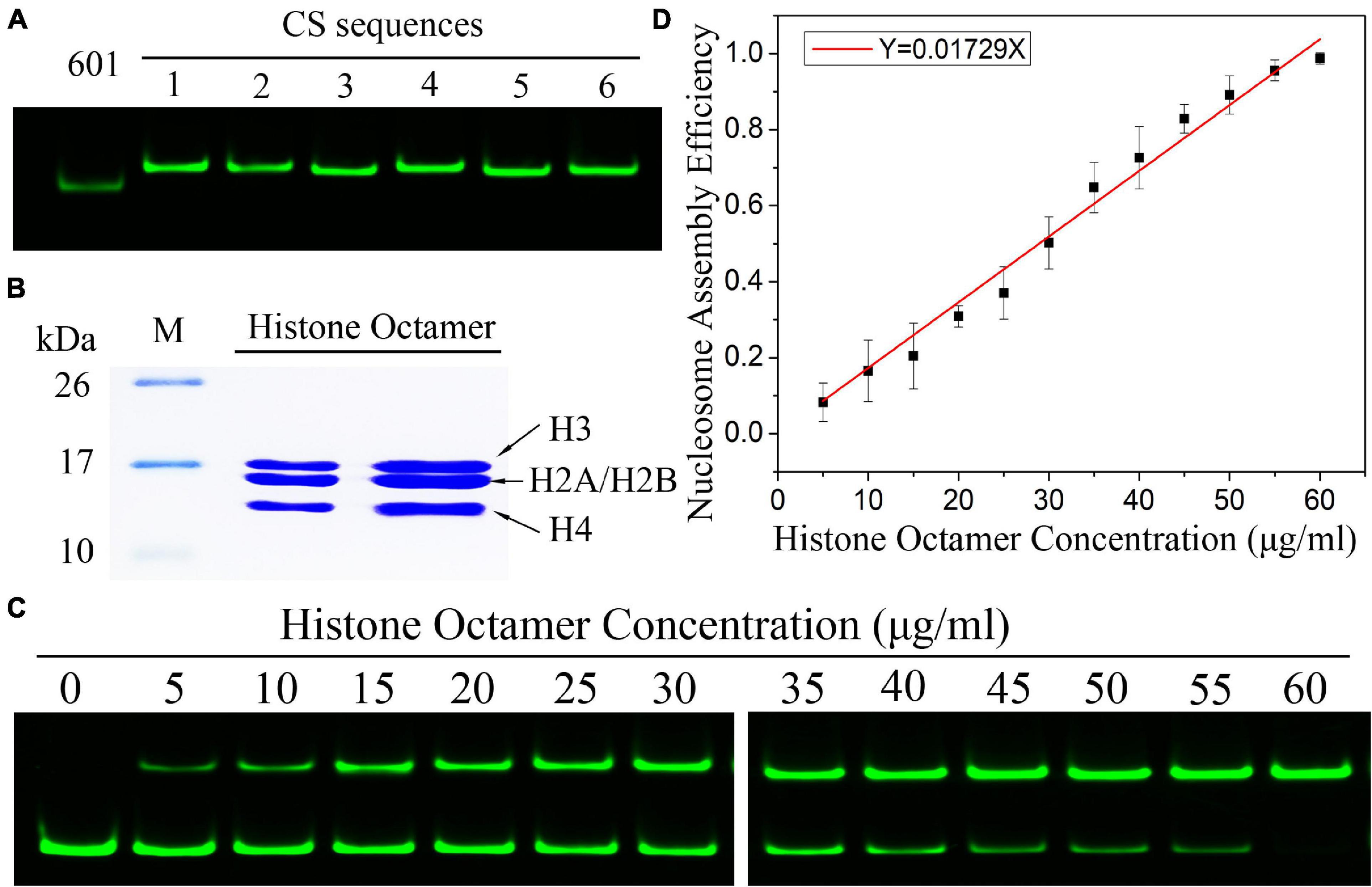
Frontiers | Nucleosome Assembly and Disassembly in vitro Are Governed by Chemical Kinetic Principles
Nucleosome positioning on large tandem DNA repeats of the '601' sequence engineered in Saccharomyces cerevisiae
Table S1. DNA sequences used to position nucleosomes, related to Figure 1 The 601 nucleosome positioning sequence (Lowary and Wi

Nucleosome Assembly 601 Sequence DNA, Biotinylated for chromatin assembly and nucleosome reconstitution, useful for pull-down and binding assays

Constructing arrays of nucleosome positioning sequences using Gibson Assembly for single-molecule studies | Scientific Reports

Linker histone H1 and H3K56 acetylation are antagonistic regulators of nucleosome dynamics | Nature Communications