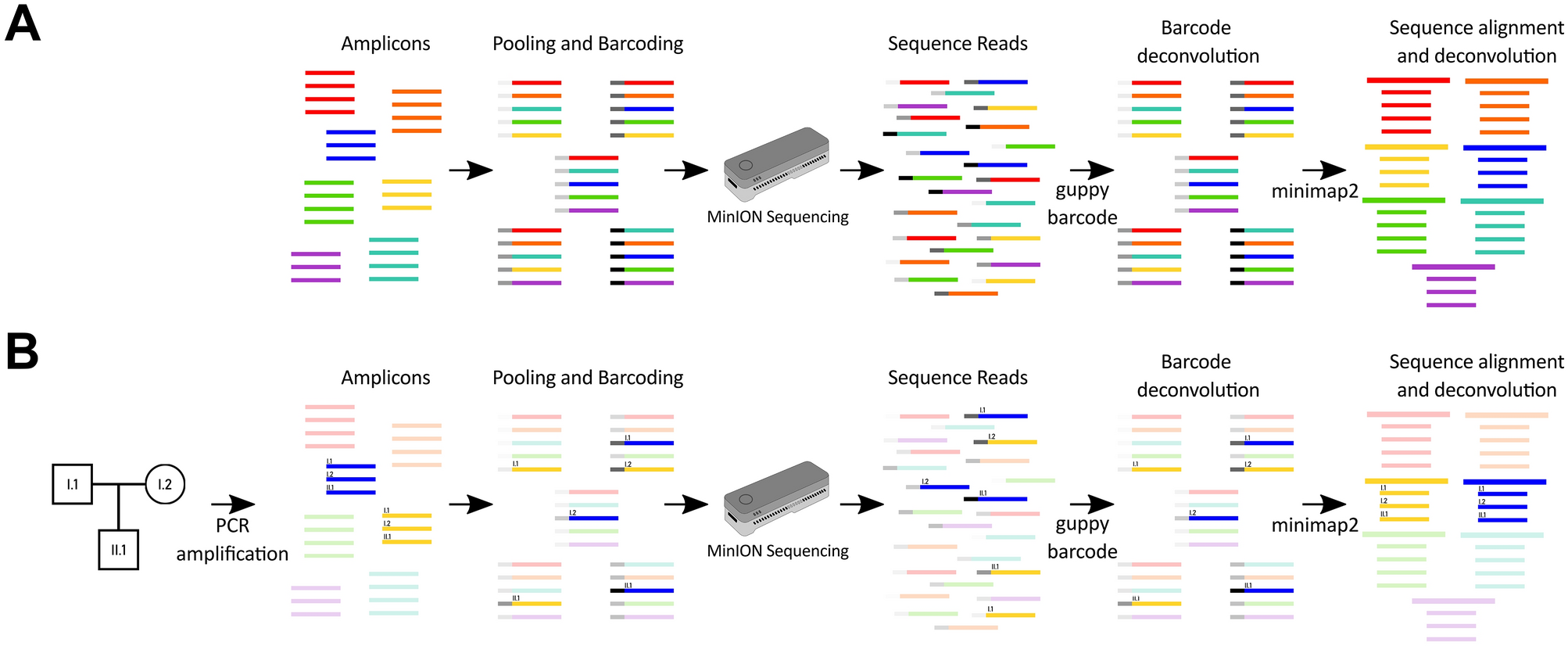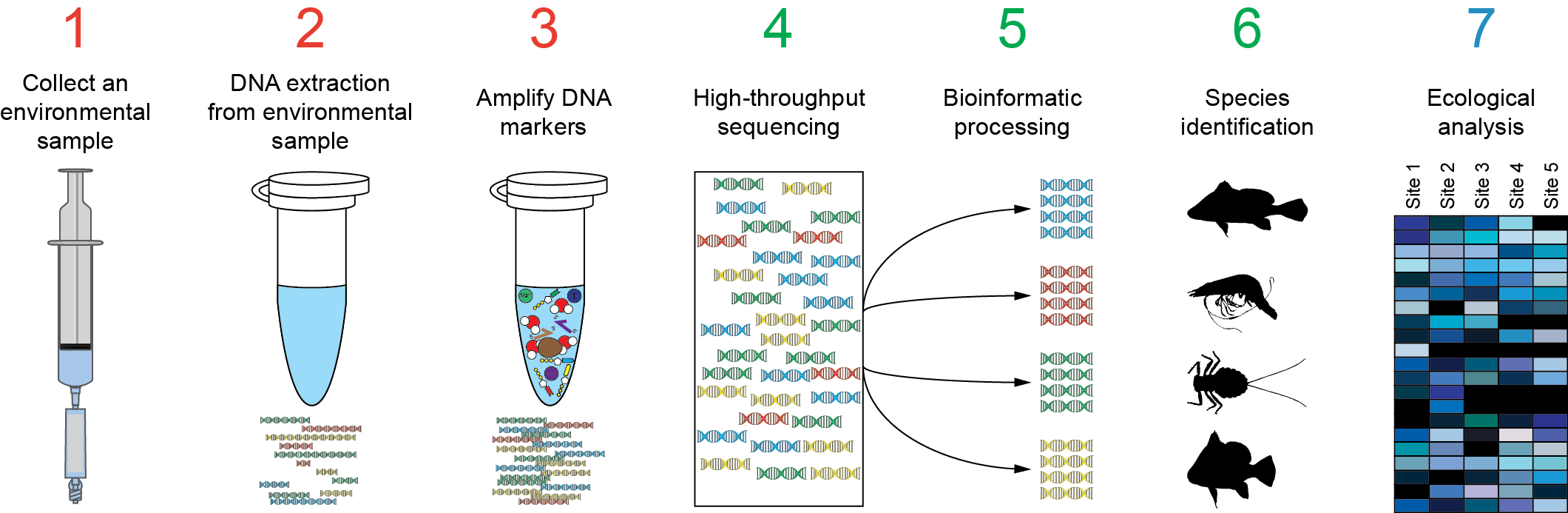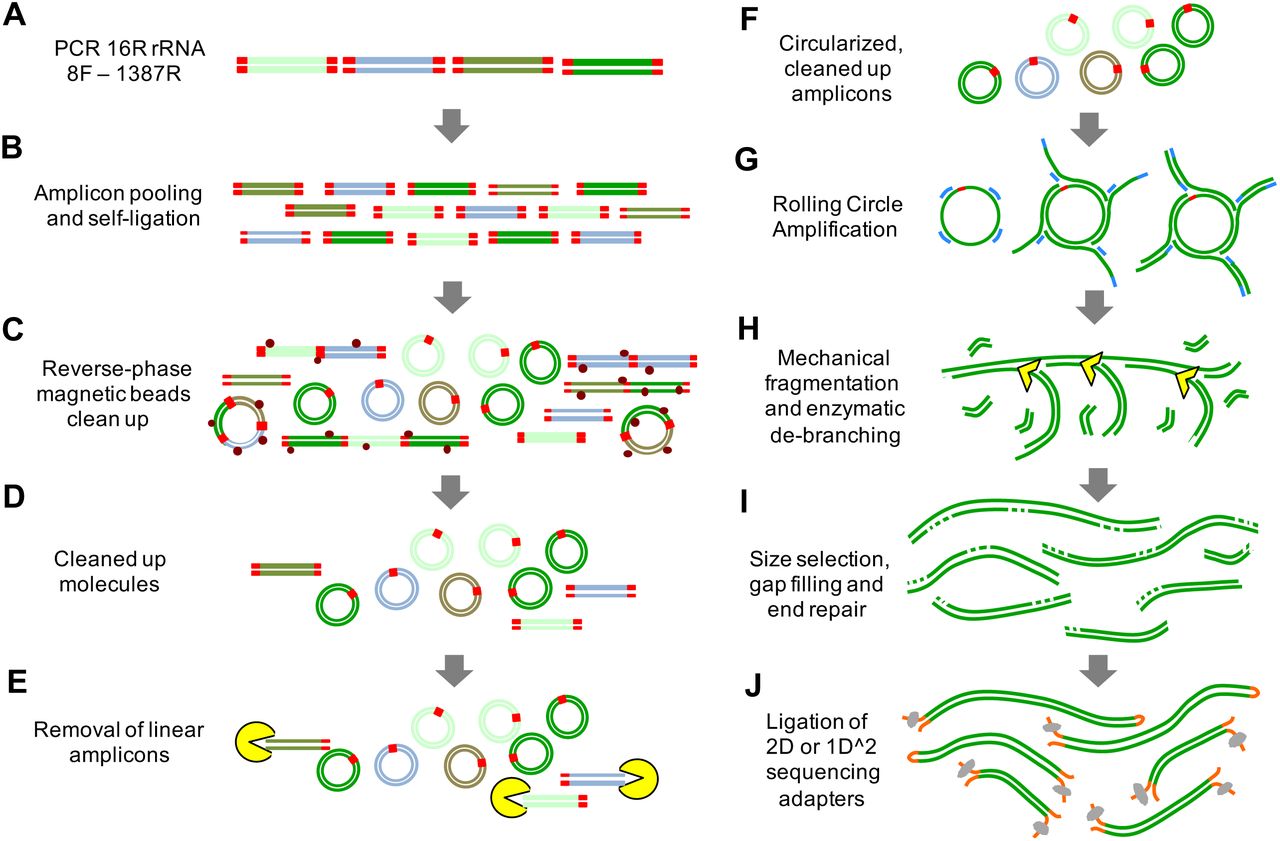
NanoAmpli-Seq: A workflow for amplicon sequencing from mixed microbial communities on the nanopore sequencing platform | bioRxiv
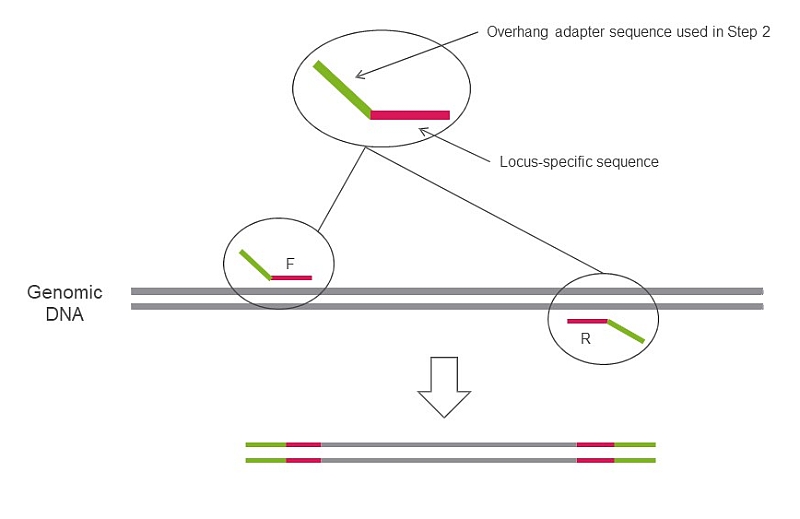
![PDF] Accurate multiplexing and filtering for high-throughput amplicon- sequencing | Semantic Scholar PDF] Accurate multiplexing and filtering for high-throughput amplicon- sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/997fc050713694d23e59913543fb9fb3f3ce4b56/4-Figure1-1.png)
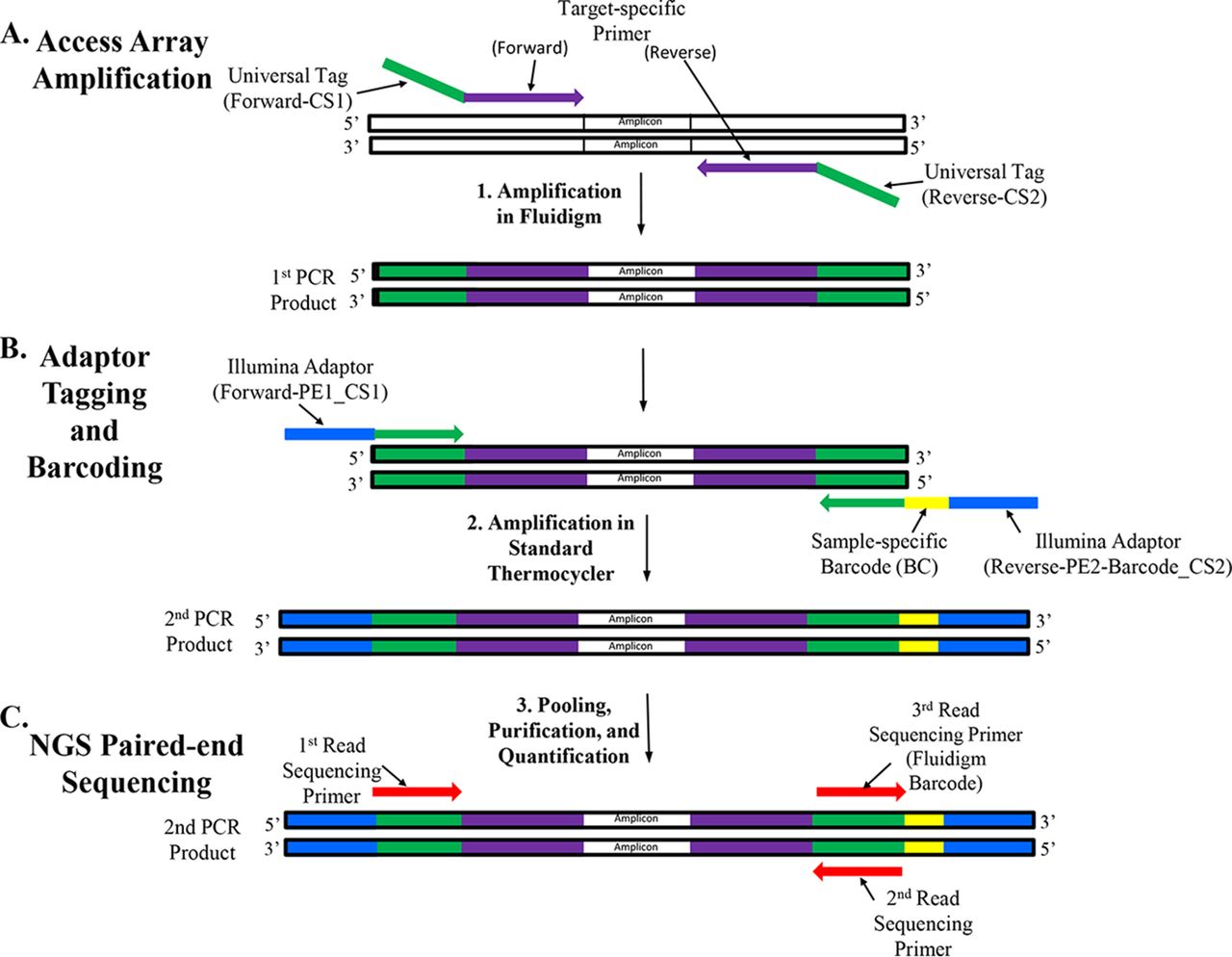
PDF] Accurate multiplexing and filtering for high-throughput amplicon- sequencing | Semantic Scholar
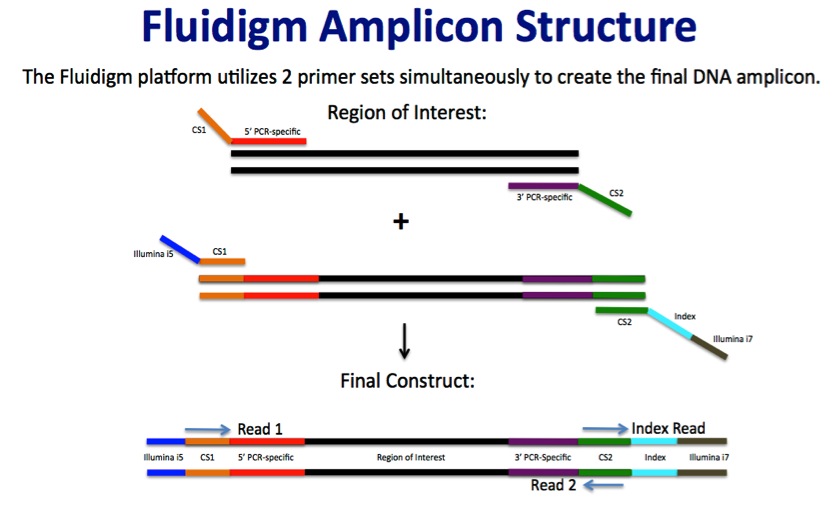
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems

Uživatel Keith Slotkin na Twitteru: „Please vote for my lab's entry into the @BioRender figure competition: Ultra-deep DNA methylation patterns via amplicon sequencing. By postdoc Andrea McCue. https://t.co/CiKXWX0Cug https://t.co/PywCf3EY2Y“ / Twitter
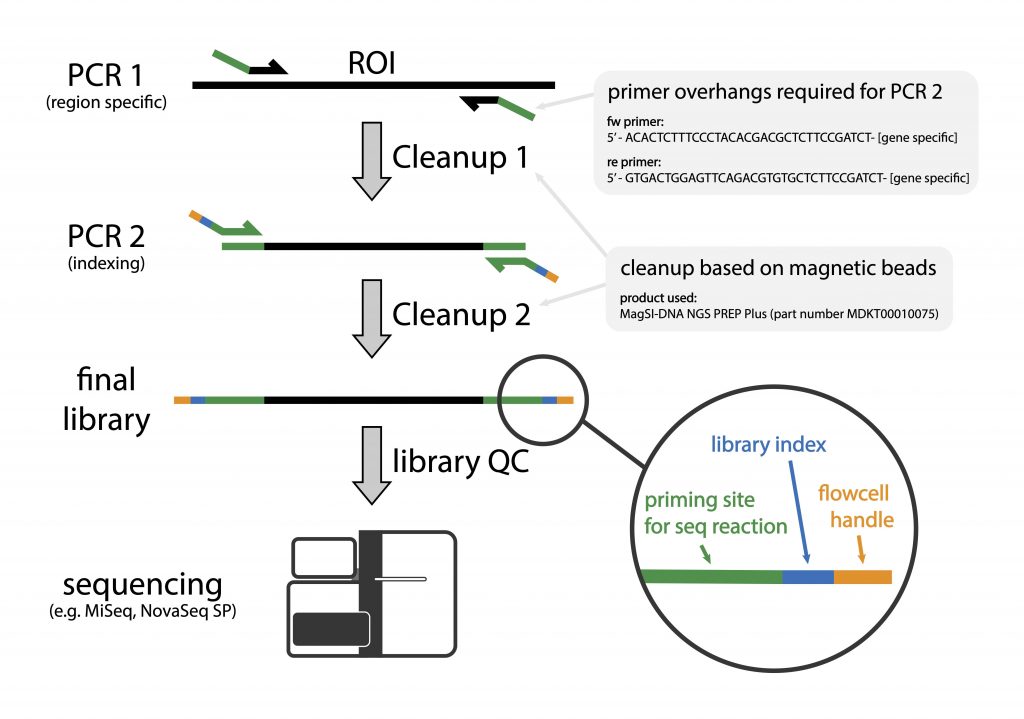
Highly multiplexed gene amplicon sequencing | Joint Microbiome Facility of the Medical University of Vienna and the University of Vienna
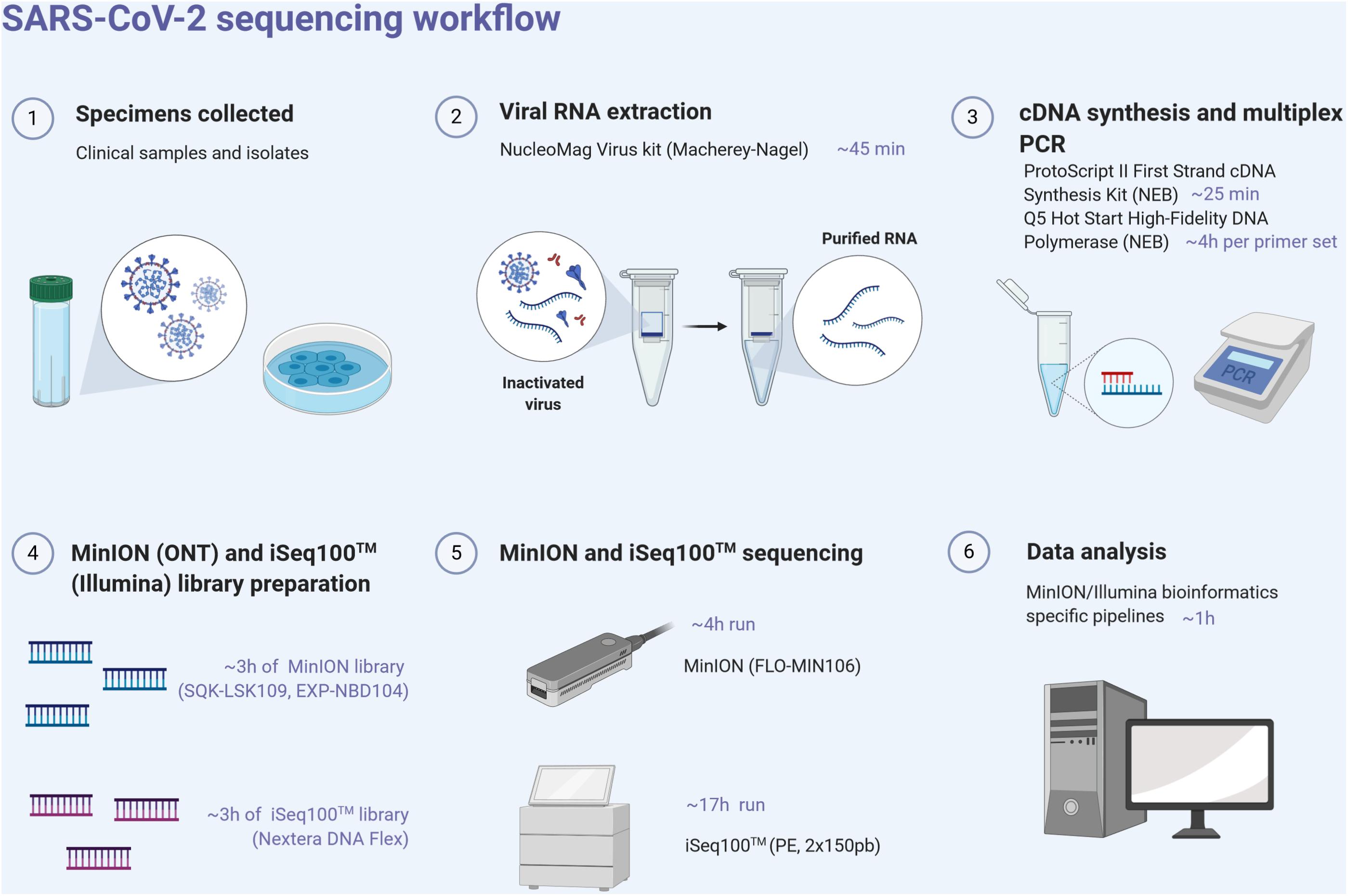
Frontiers | Rapid Genomic Characterization of SARS-CoV-2 by Direct Amplicon-Based Sequencing Through Comparison of MinION and Illumina iSeq100TM System