SBF-SEM and FIB-SEM analysis of Erg11-APEX2 in S. cerevisiae. (A, B,... | Download Scientific Diagram

Three-Dimensional Visualization of APEX2-Tagged Erg11 in Saccharomyces cerevisiae Using Focused Ion Beam Scanning Electron Microscopy | mSphere
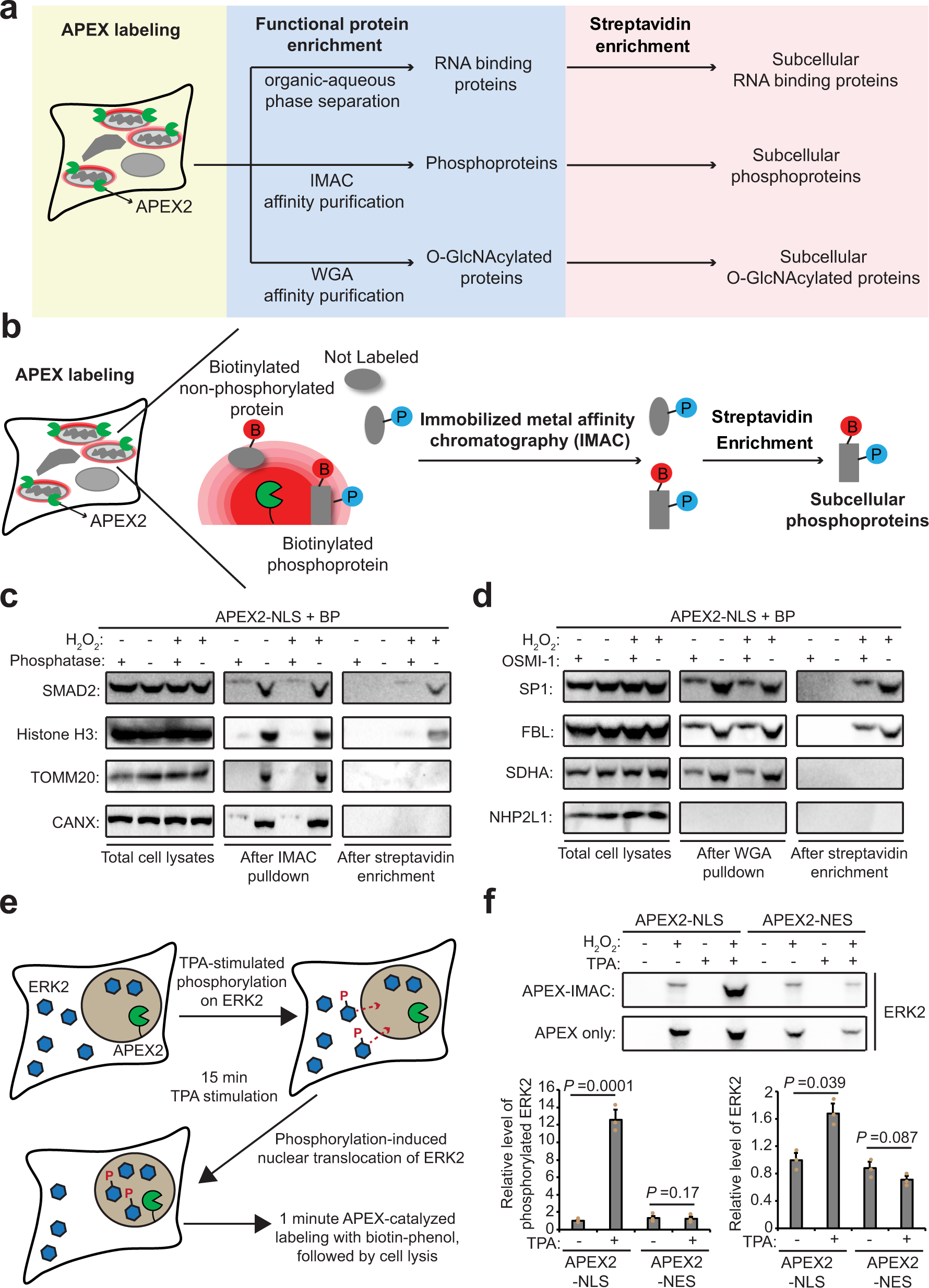
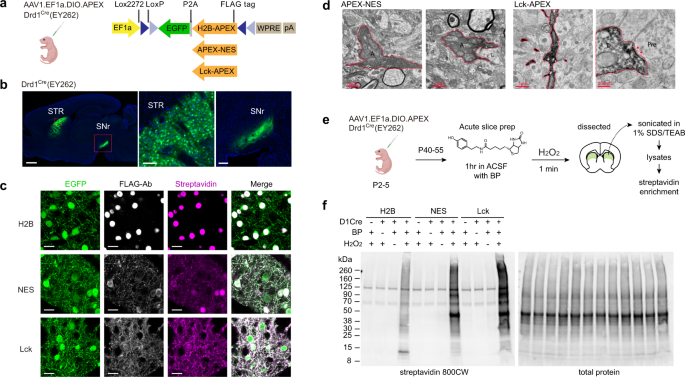
Spatiotemporally-resolved mapping of RNA binding proteins via functional proximity labeling reveals a mitochondrial mRNA anchor promoting stress recovery | Nature Communications
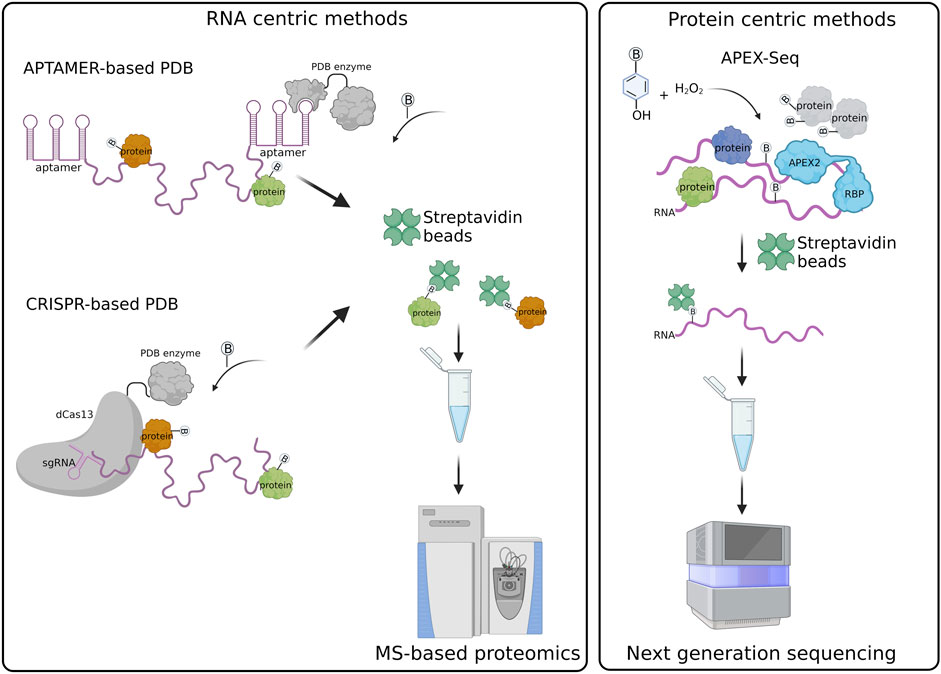
Frontiers | Proximity-dependent biotinylation technologies for mapping RNA-protein interactions in live cells
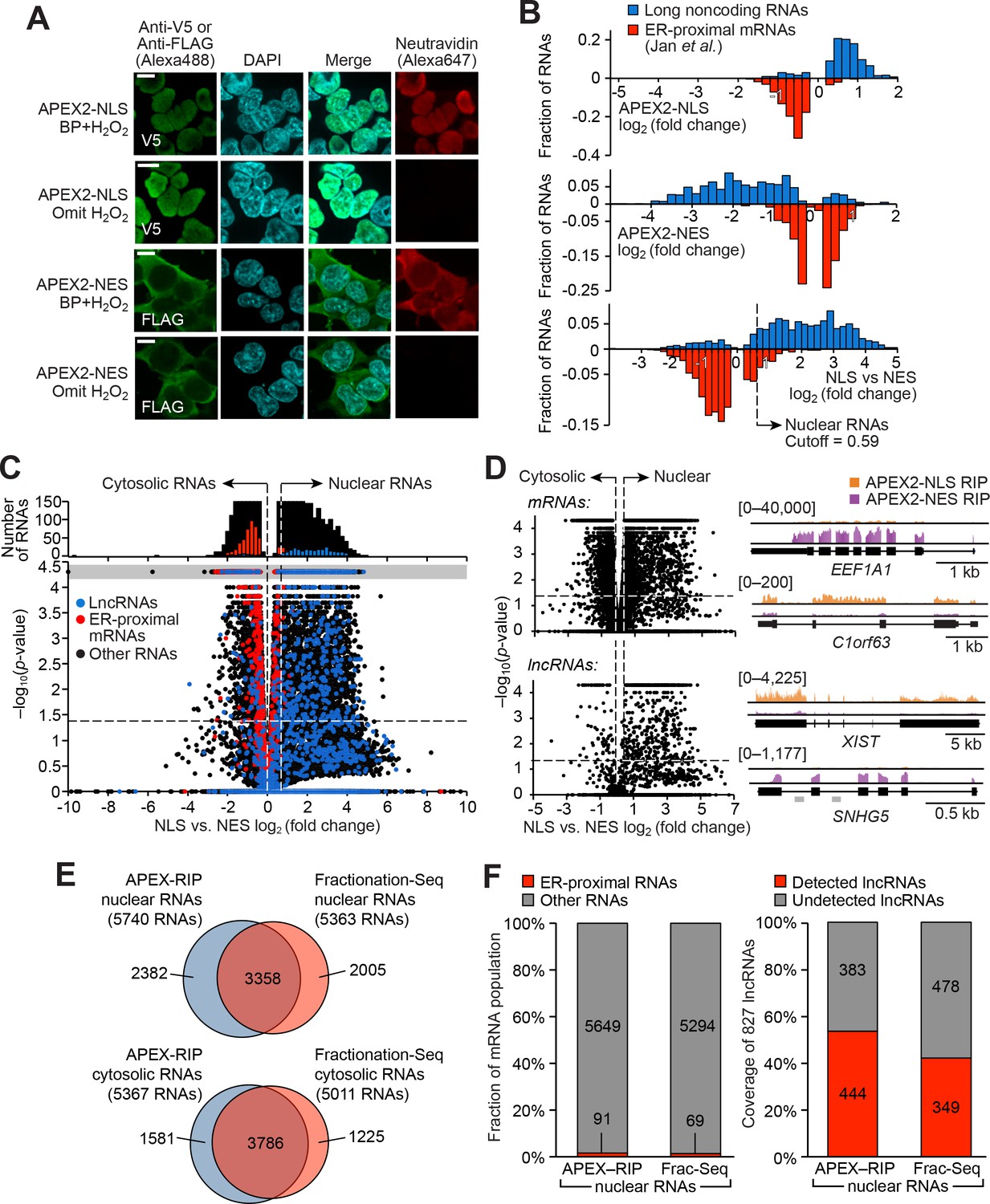
Figures and data in Live-cell mapping of organelle-associated RNAs via proximity biotinylation combined with protein-RNA crosslinking | eLife
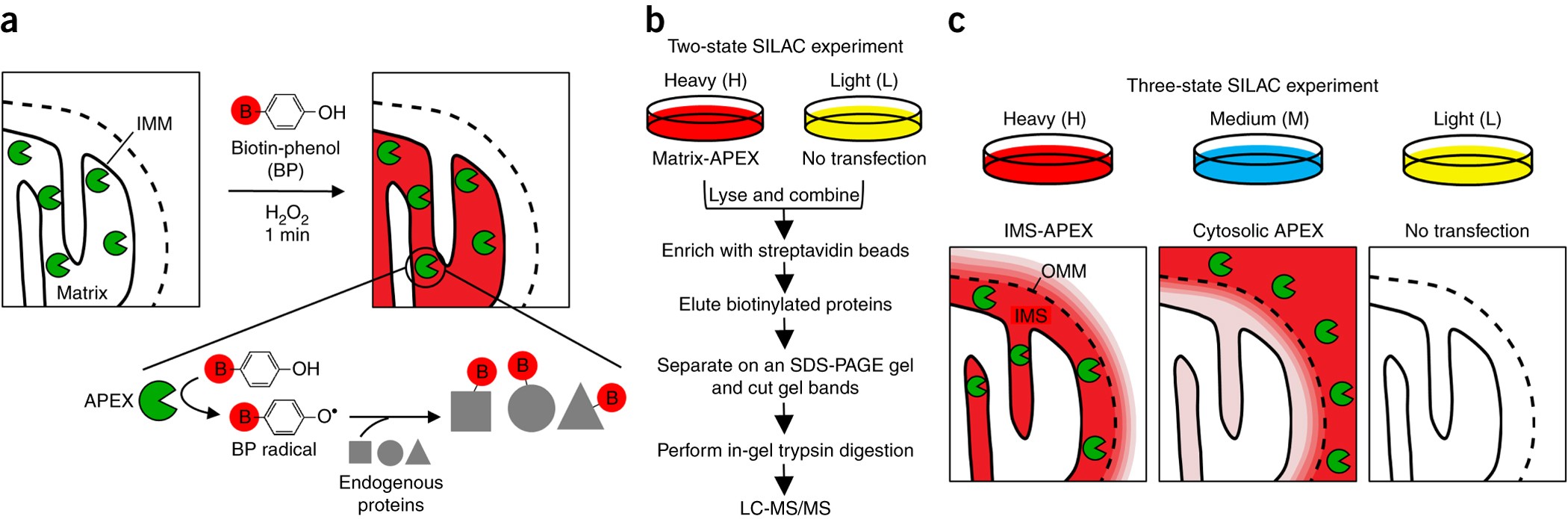
Spatially resolved proteomic mapping in living cells with the engineered peroxidase APEX2 | Nature Protocols

In vivo discovery of RNA proximal proteins in human cells via proximity-dependent biotinylation | bioRxiv
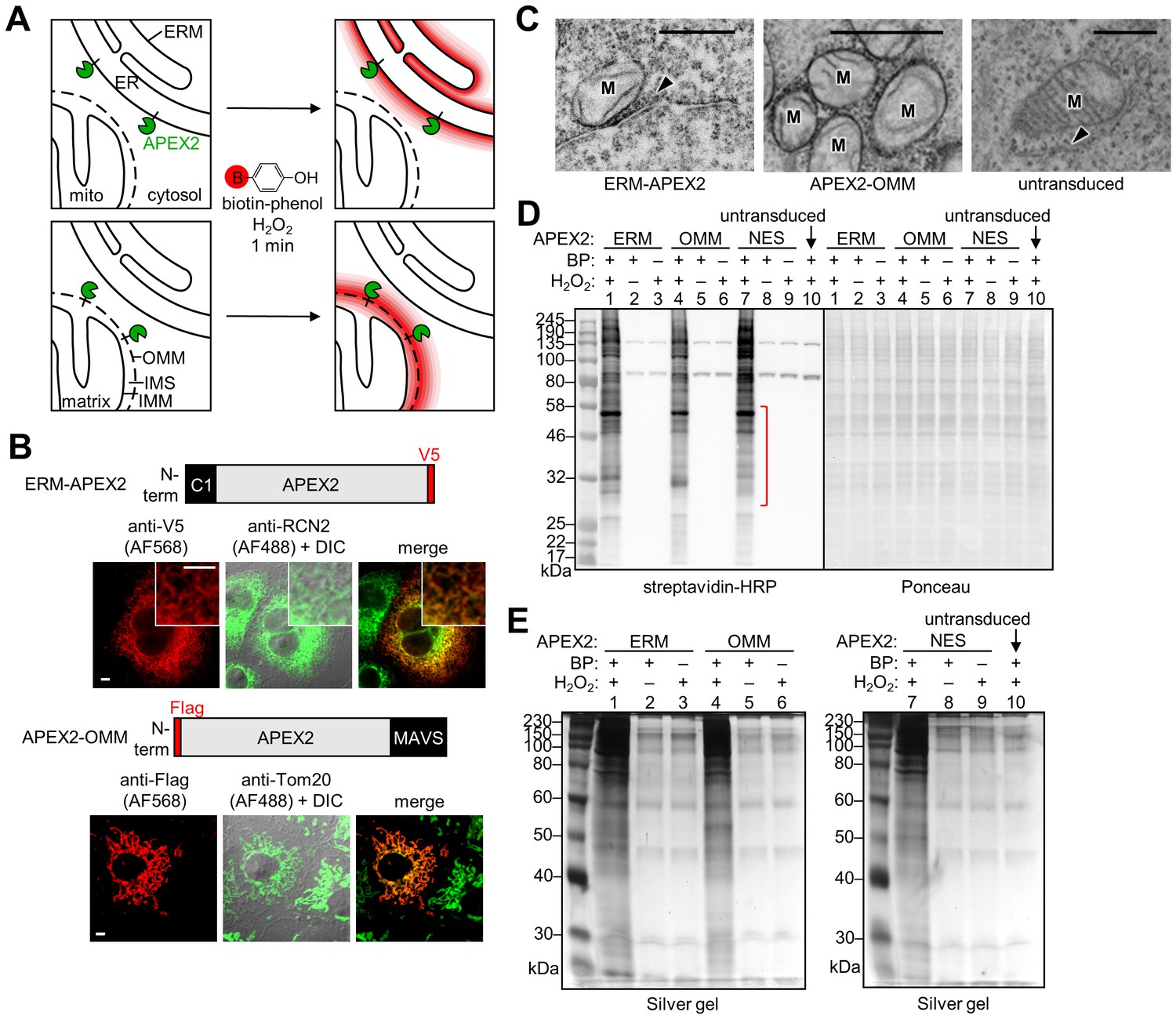
Proteomic mapping of cytosol-facing outer mitochondrial and ER membranes in living human cells by proximity biotinylation | eLife

Proximity RNA Labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules - ScienceDirect

IJMS | Free Full-Text | Differential CFTR-Interactome Proximity Labeling Procedures Identify Enrichment in Multiple SLC Transporters

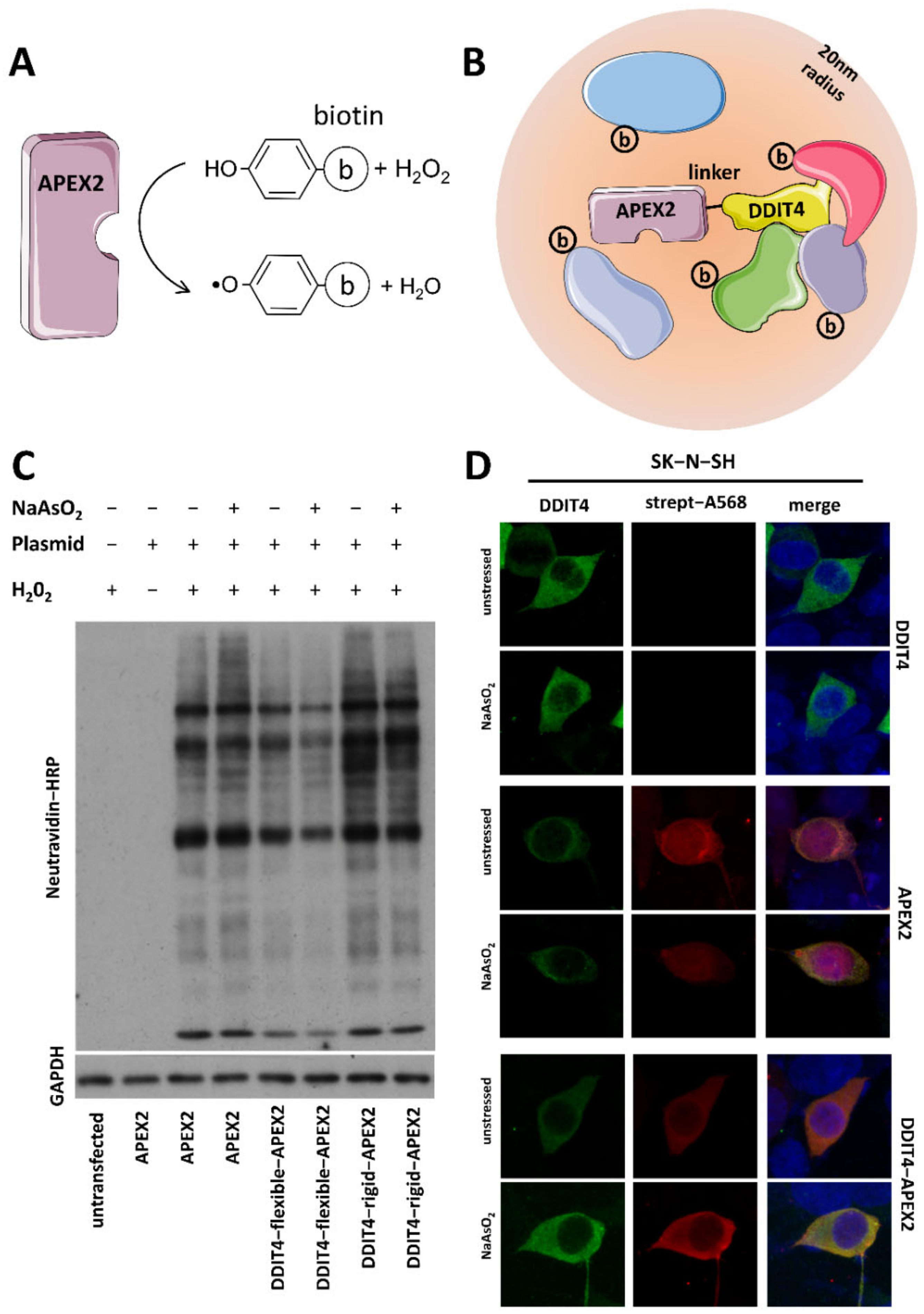
Targeting APEX2 to human telomerase RNA with CRISPR-Cas13. (A) Design... | Download Scientific Diagram