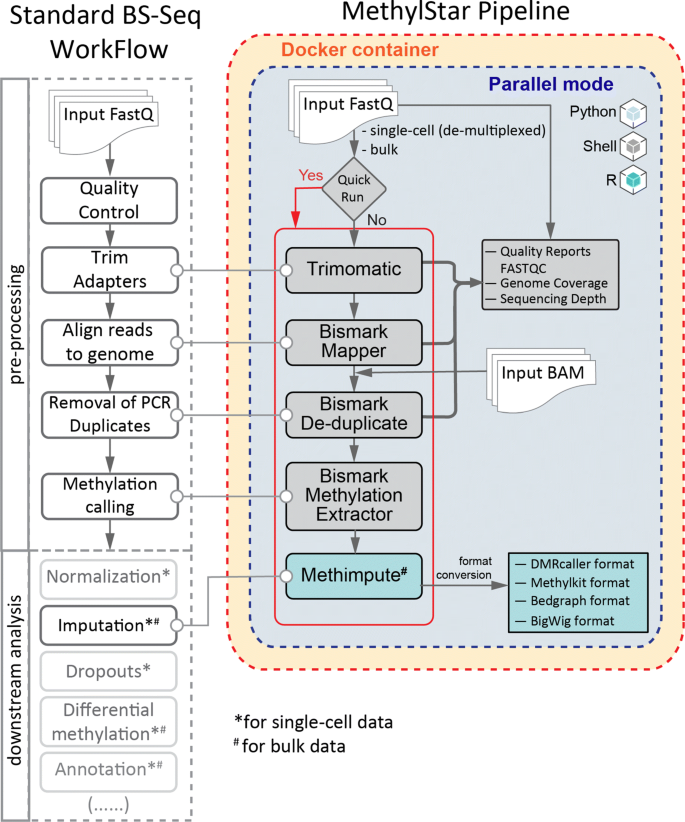
MethylStar: A fast and robust pre-processing pipeline for bulk or single-cell whole-genome bisulfite sequencing data | BMC Genomics | Full Text
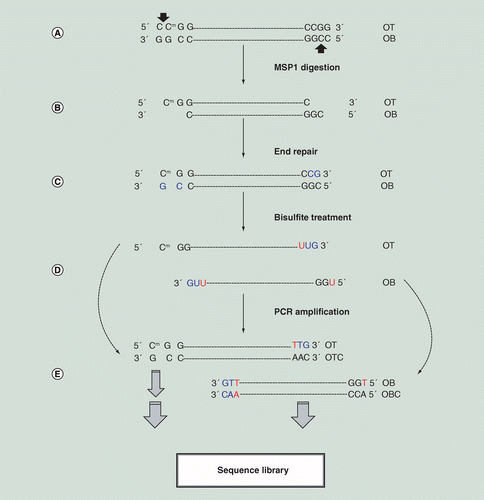
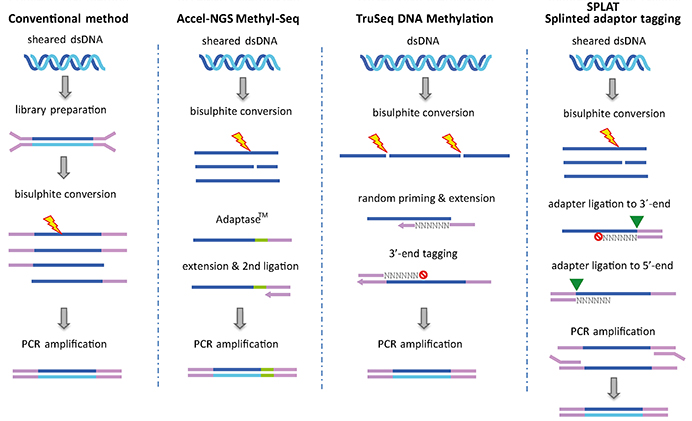
Base resolution methylome profiling: considerations in platform selection, data preprocessing and analysis | Epigenomics

Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing | Nature Communications

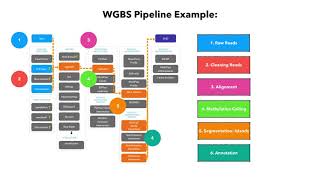
The workflow of analyzing DNA methylation using bisulfite sequencing data. | Download Scientific Diagram

Benchmarking DNA methylation analysis of 14 alignment algorithms for whole genome bisulfite sequencing in mammals - ScienceDirect
Methy-Pipe: An Integrated Bioinformatics Pipeline for Whole Genome Bisulfite Sequencing Data Analysis | PLOS ONE
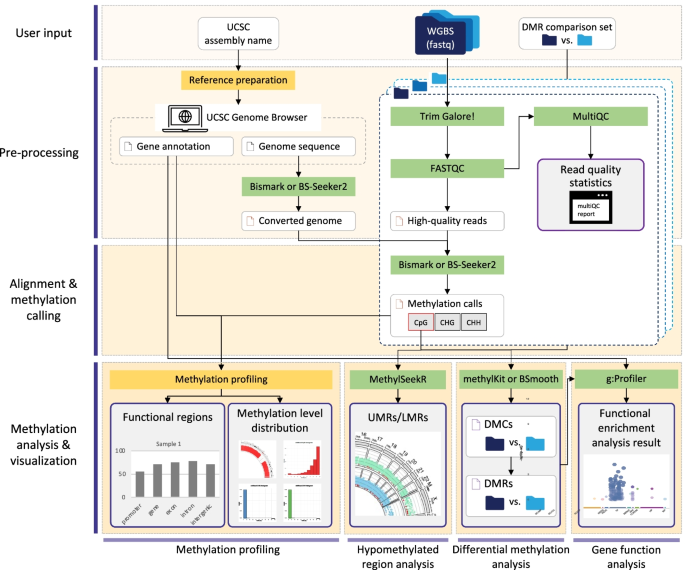
msPIPE: a pipeline for the analysis and visualization of whole-genome bisulfite sequencing data | BMC Bioinformatics | Full Text
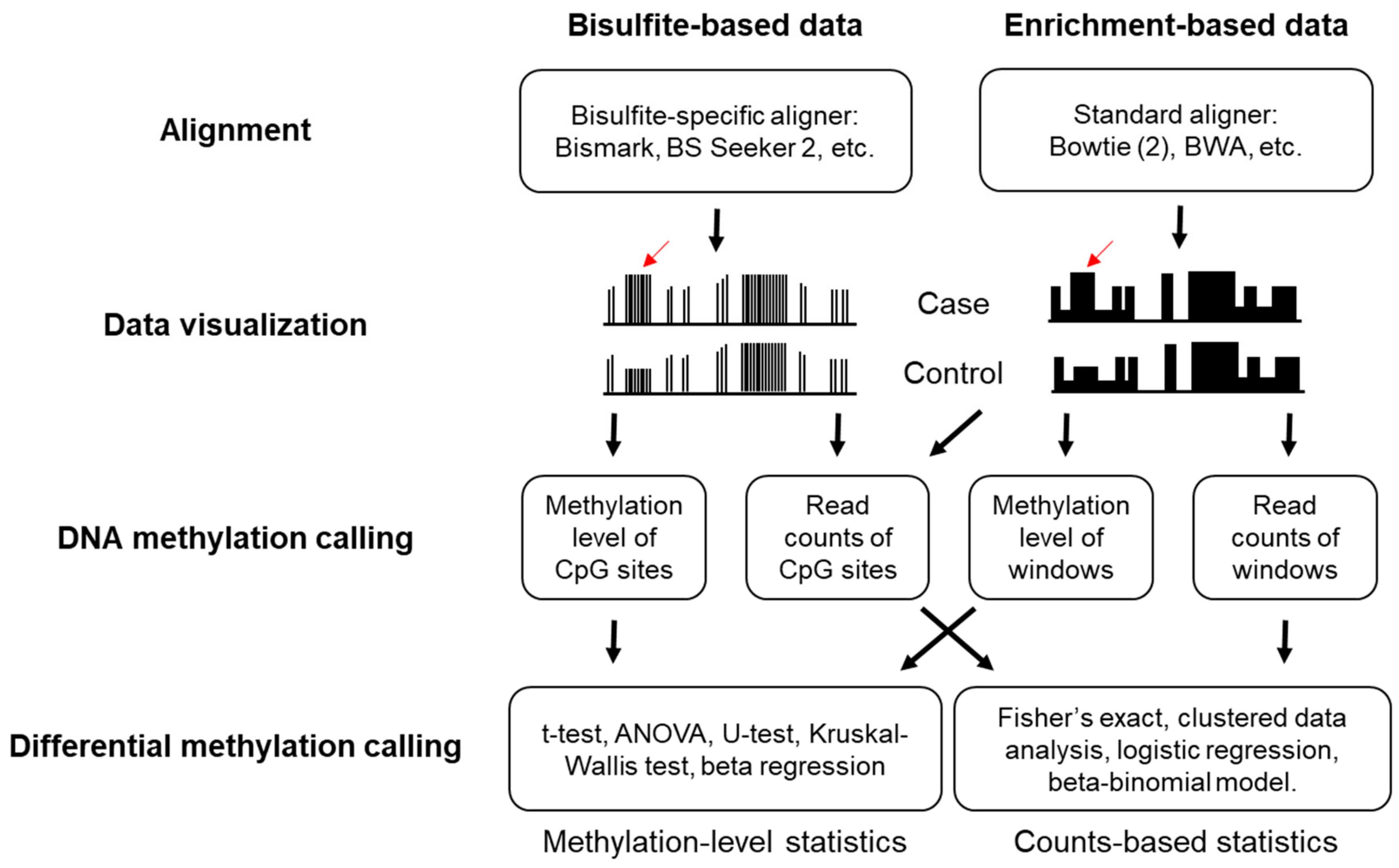
Cancers | Free Full-Text | Cell-Free DNA Methylation Profiling Analysis—Technologies and Bioinformatics

Webinar: Epigenetics Part I – Bisulfite Sequence Analysis and Adenosine to Inosine Modifications - YouTube
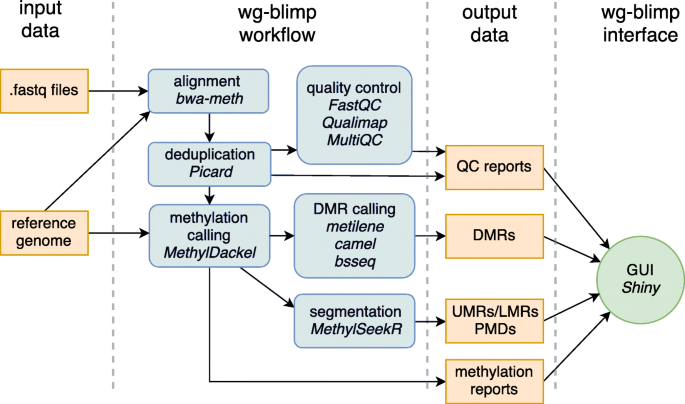
wg-blimp: an end-to-end analysis pipeline for whole genome bisulfite sequencing data | BMC Bioinformatics | Full Text

MethylScore, a pipeline for accurate and context-aware identification of differentially methylated regions from population-scale plant whole-genome bisulfite sequencing data | Quantitative Plant Biology | Cambridge Core

GemBS – high through-put processing pipeline for DNA methylation data from Whole Genome Bisulfite Sequencing (WGBS) | bioRxiv


![PDF] MethylSig: a whole genome DNA methylation analysis pipeline | Semantic Scholar PDF] MethylSig: a whole genome DNA methylation analysis pipeline | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/ced5bb274877737921b0e71c311e9043d480021c/3-Figure1-1.png)

![PDF] Bicycle: a bioinformatics pipeline to analyze bisulfite sequencing data | Semantic Scholar PDF] Bicycle: a bioinformatics pipeline to analyze bisulfite sequencing data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3fce655ad9994e0081aaf428ba74adf45682a11a/2-Figure1-1.png)