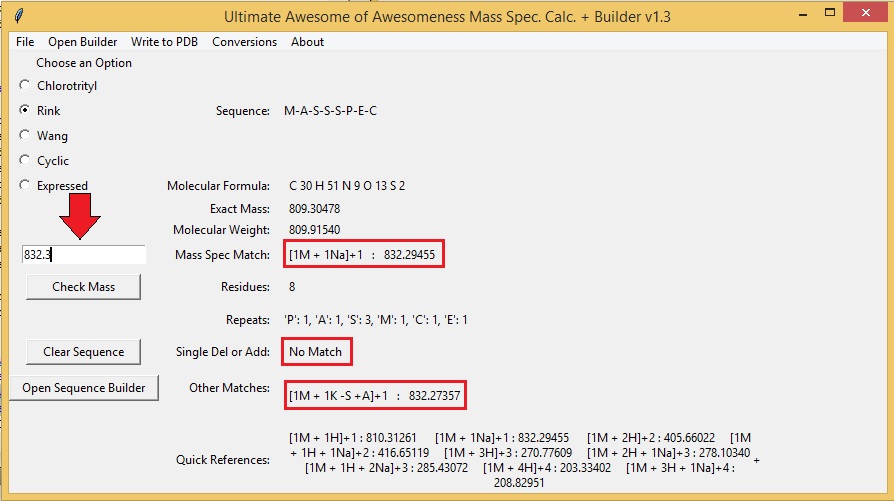
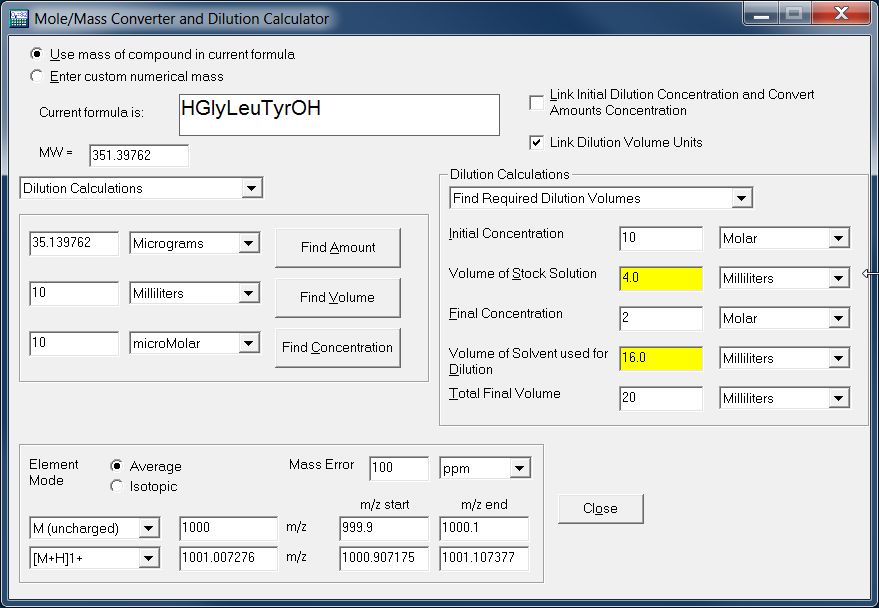
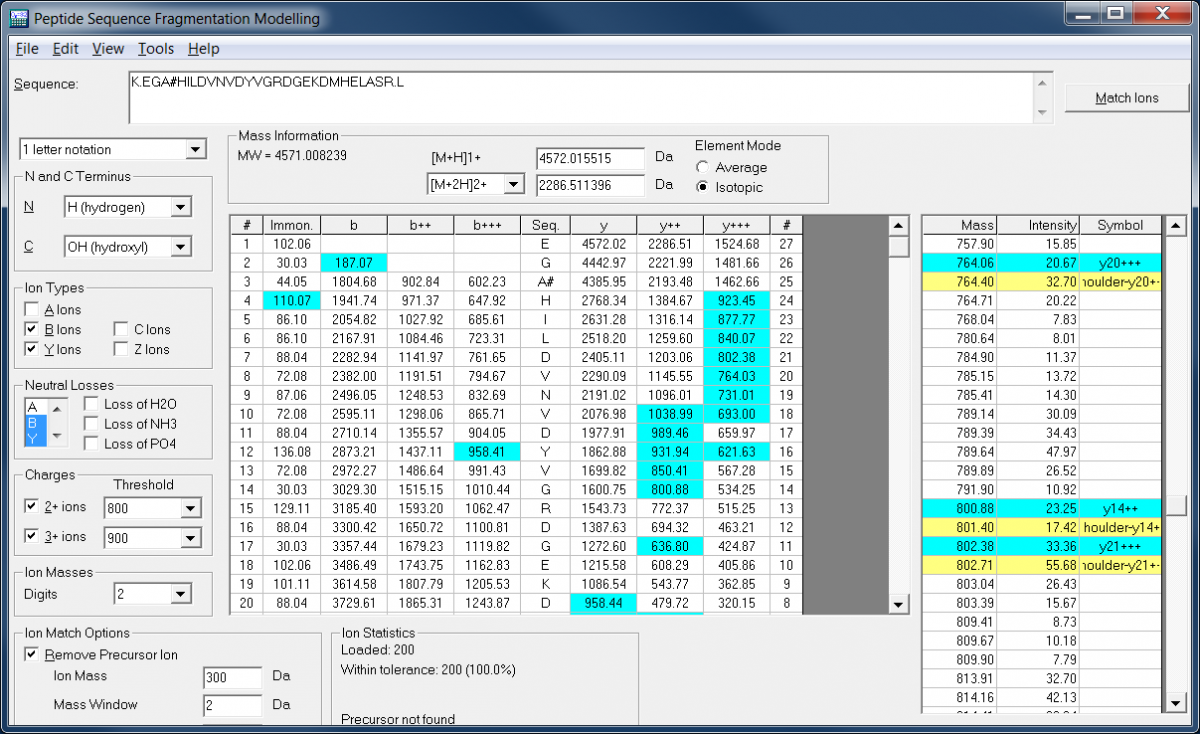
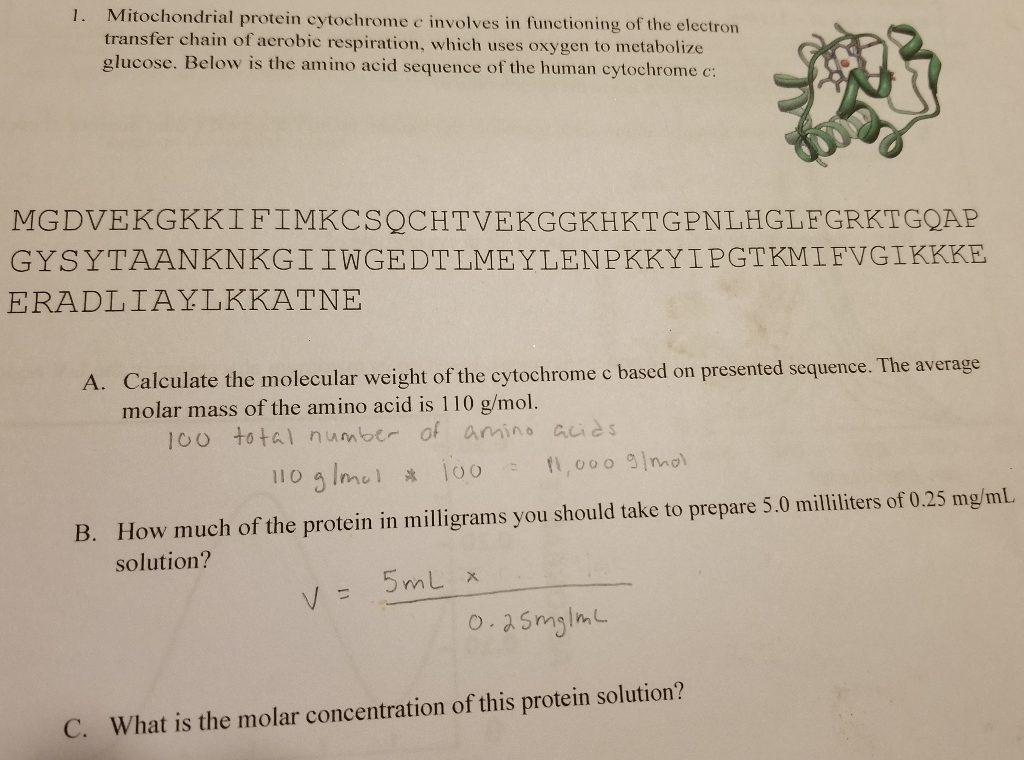SOLVED: Insert random 9-mer containing at least one of each nucleobase in the following GCCUGUUGG-(N9)-CCAACAAU sequence: () Calculate the molecular weight, percent GC content and molar absorptivity for the oligomer designed: You (

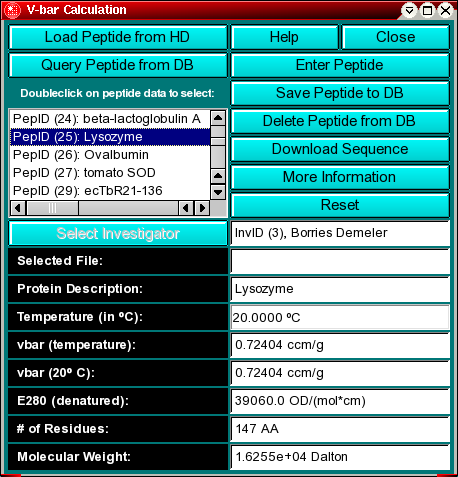
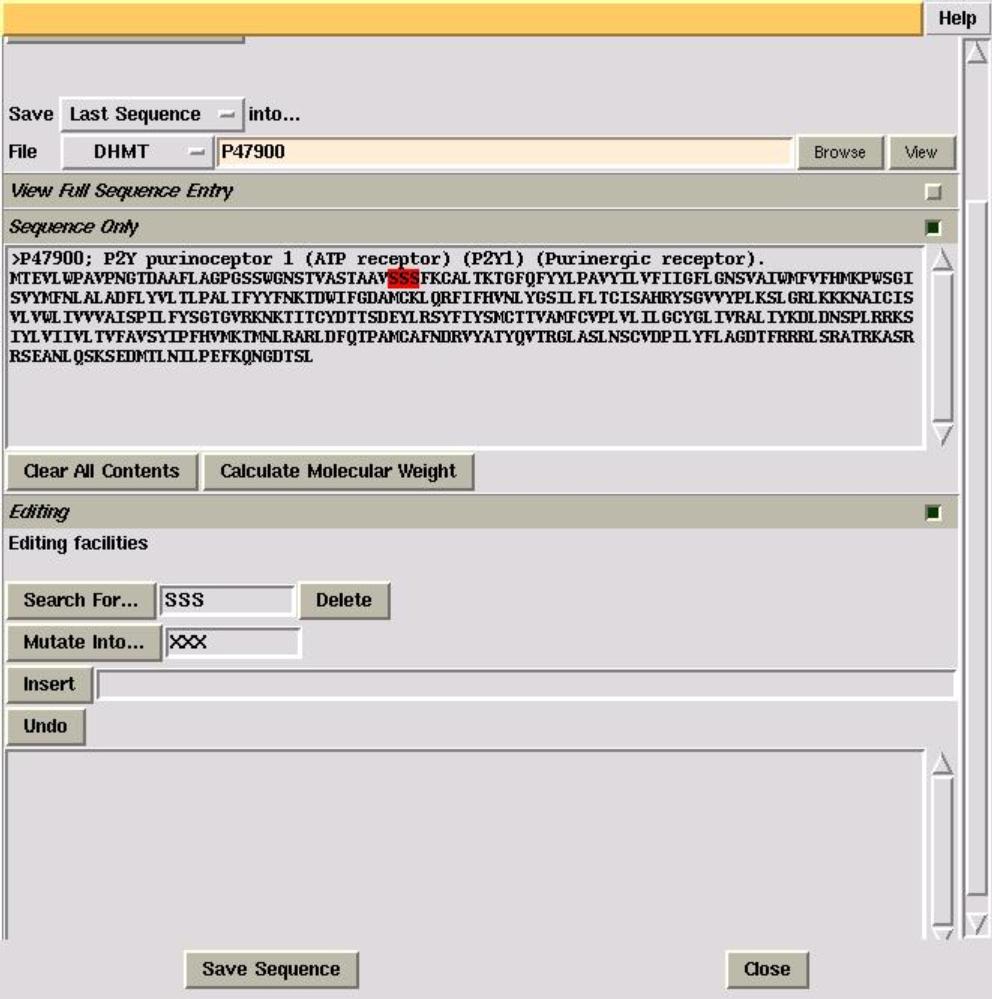
SOLVED: The OTCΔ protein is a mutant of OTC that has a deletion of 85 amino acids from Leucine 63 to Aspartic Acid 147 (85 amino acids total). Using the OTC sequence

Flowchart summarizing the mass spectral peak assignment algorithm used... | Download Scientific Diagram
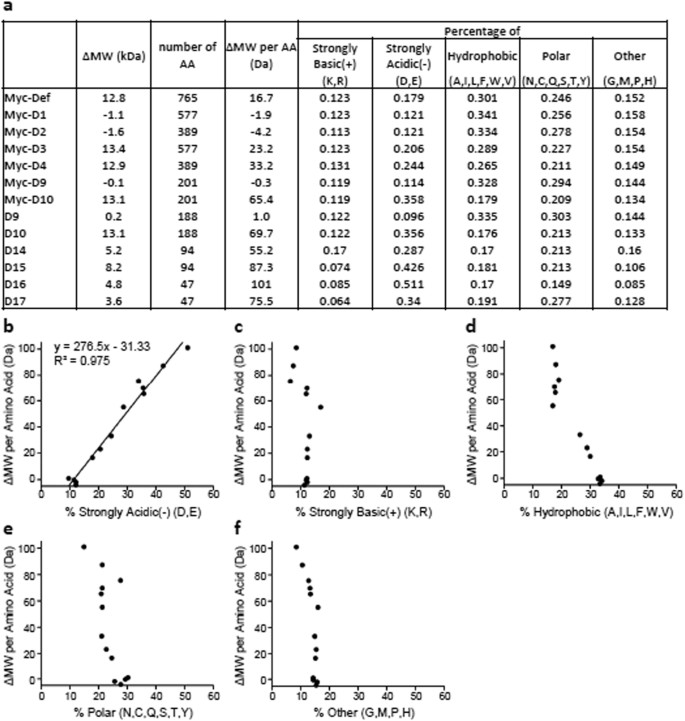
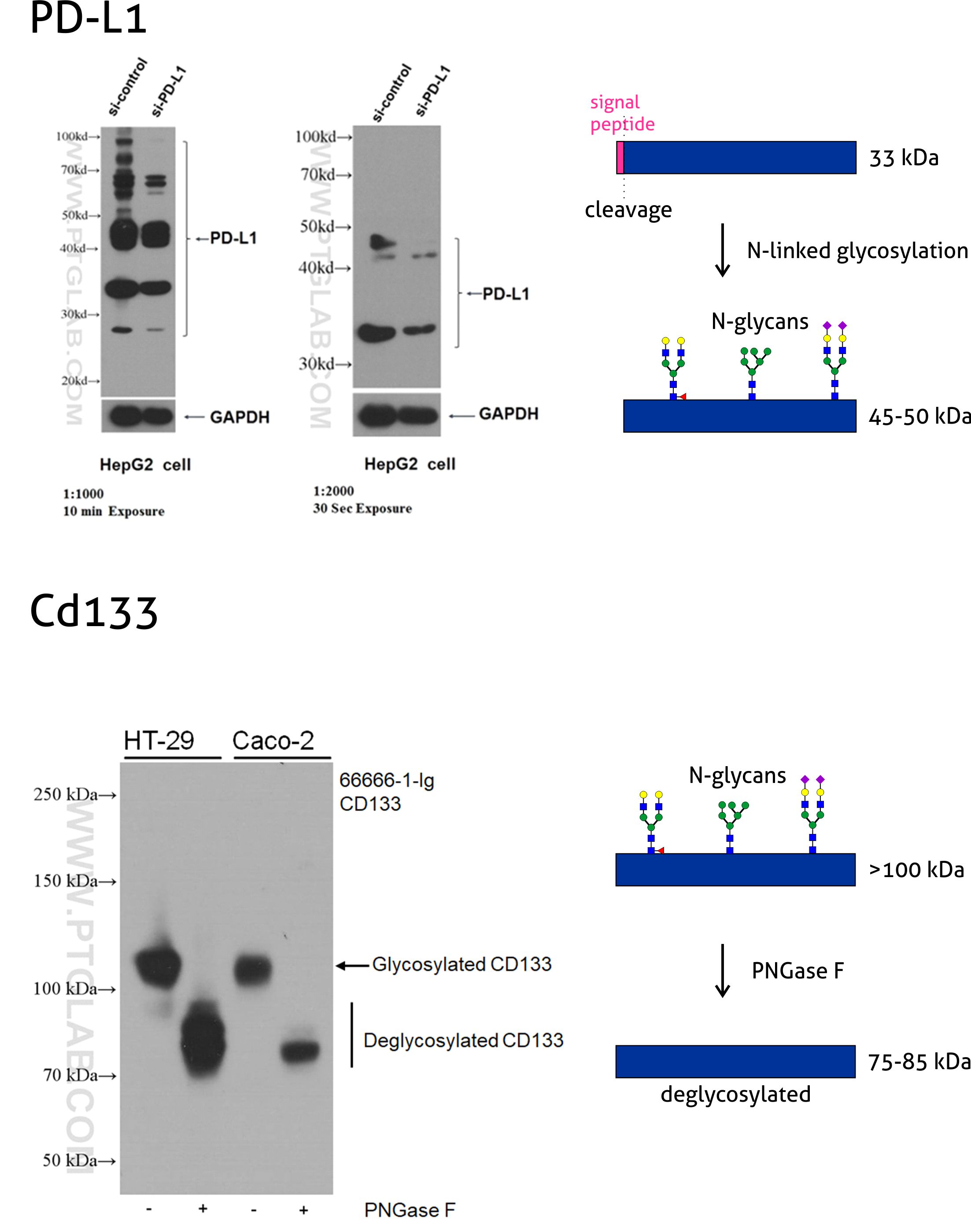
An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide | Scientific Reports