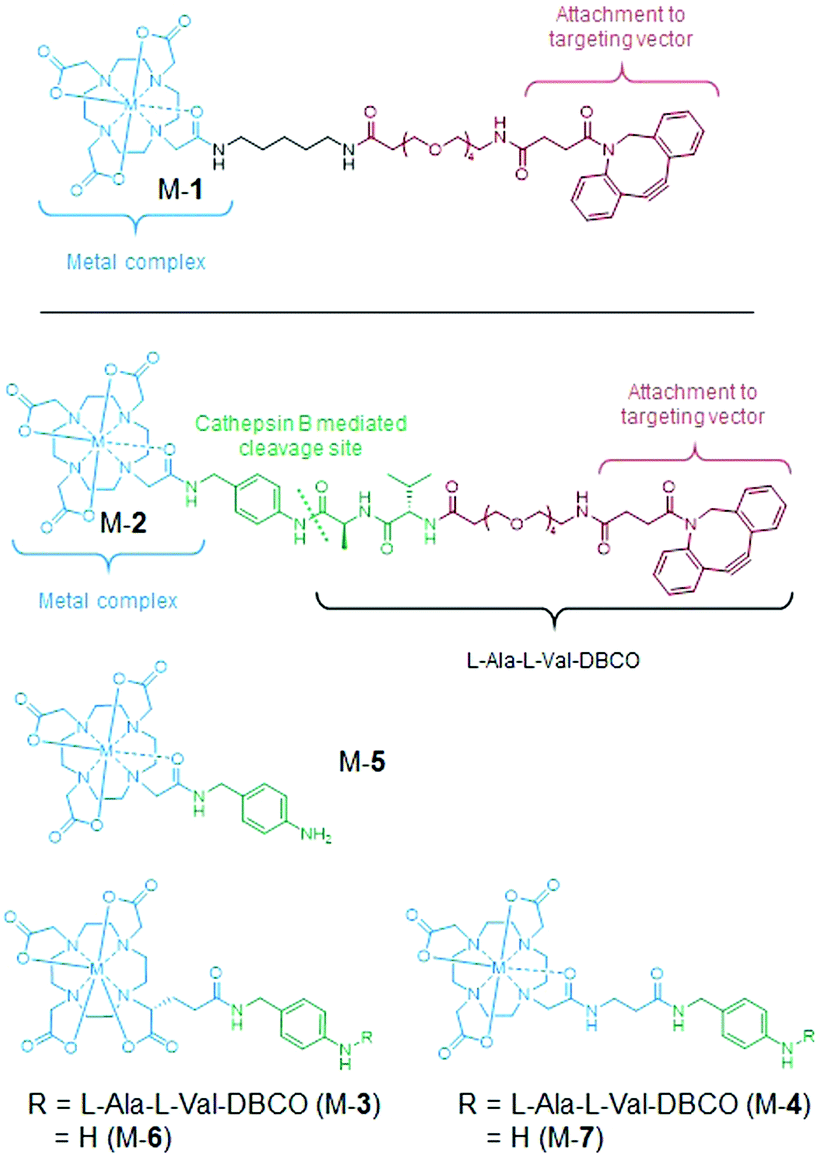
Trivalent metal complex geometry of the substrate governs cathepsin B enzymatic cleavage rate - Chemical Communications (RSC Publishing) DOI:10.1039/D0CC02862B

Proteomic Identification of Protease Cleavage Sites Characterizes Prime and Non-prime Specificity of Cysteine Cathepsins B, L, and S | Journal of Proteome Research
![PDF] Identification of Pathogenic T Cell Epitopes Near Cathepsin Cleavage Sites in Thyroglobulin | Semantic Scholar PDF] Identification of Pathogenic T Cell Epitopes Near Cathepsin Cleavage Sites in Thyroglobulin | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/fea677ffbc2a1a80e96f0a385507b7254d33af11/2-TableI-1.png)
PDF] Identification of Pathogenic T Cell Epitopes Near Cathepsin Cleavage Sites in Thyroglobulin | Semantic Scholar

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal
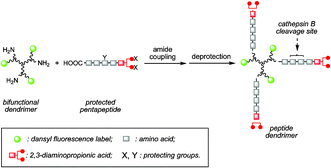
Fluorescent dendrimers with a peptide cathepsin B cleavage site for drug delivery applications - Chemical Communications (RSC Publishing)

Cathepsin B Dipeptidyl Carboxypeptidase and Endopeptidase Activities Demonstrated across a Broad pH Range | Biochemistry
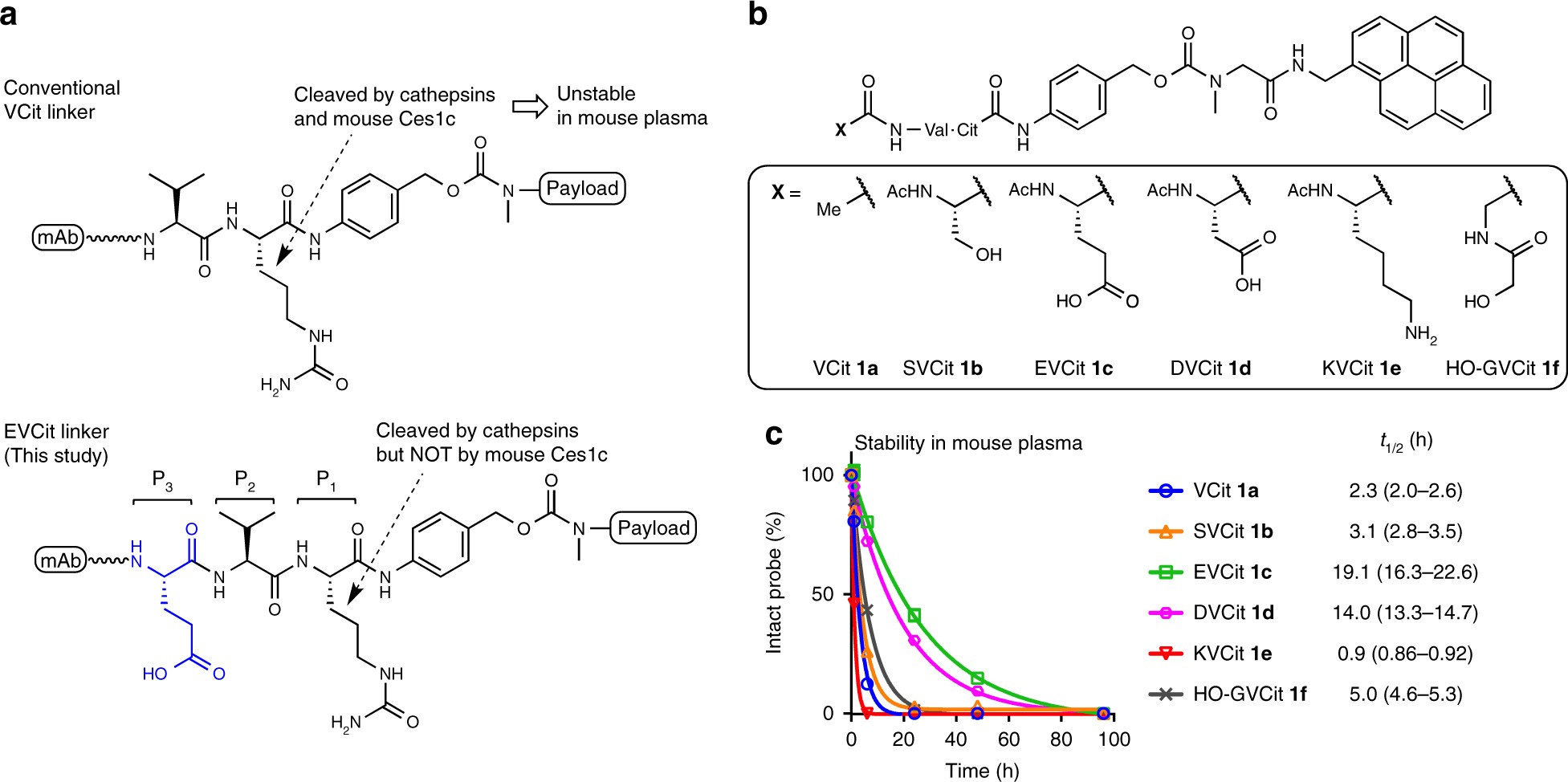
Glutamic acid–valine–citrulline linkers ensure stability and efficacy of antibody–drug conjugates in mice | Nature Communications

Selective Neutral pH Inhibitor of Cathepsin B Designed Based on Cleavage Preferences at Cytosolic and Lysosomal pH Conditions | ACS Chemical Biology

Computational predictions of cysteine cathepsin‐mediated fibrinogen proteolysis - Ferrall‐Fairbanks - 2018 - Protein Science - Wiley Online Library

The two cathepsin B-like proteases of Arabidopsis thaliana are closely related enzymes with discrete endopeptidase and carboxydipeptidase activities
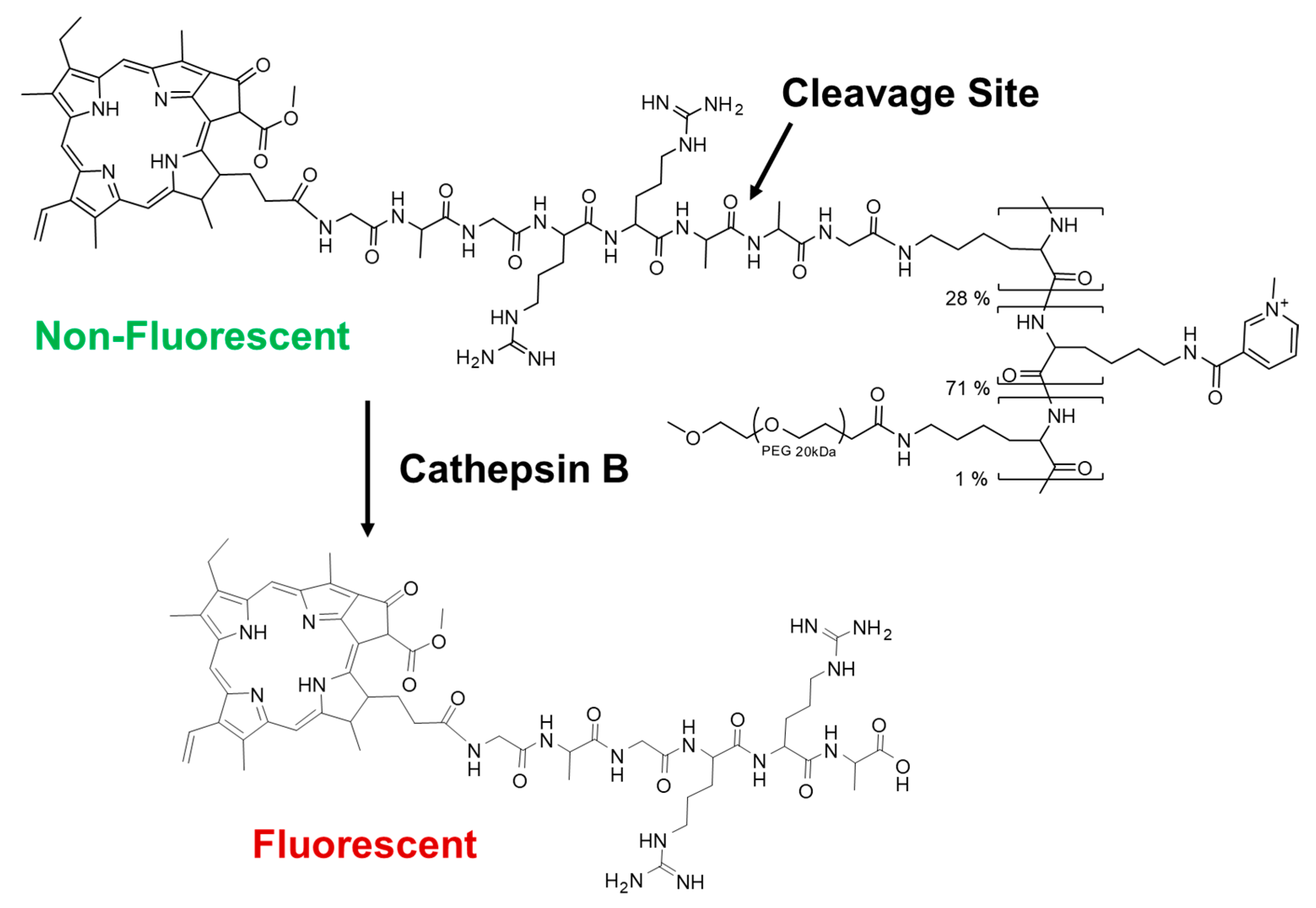
Pharmaceuticals | Free Full-Text | Cathepsin B-Cleavable Polymeric Photosensitizer Prodrug for Selective Photodynamic Therapy: In Vitro Studies
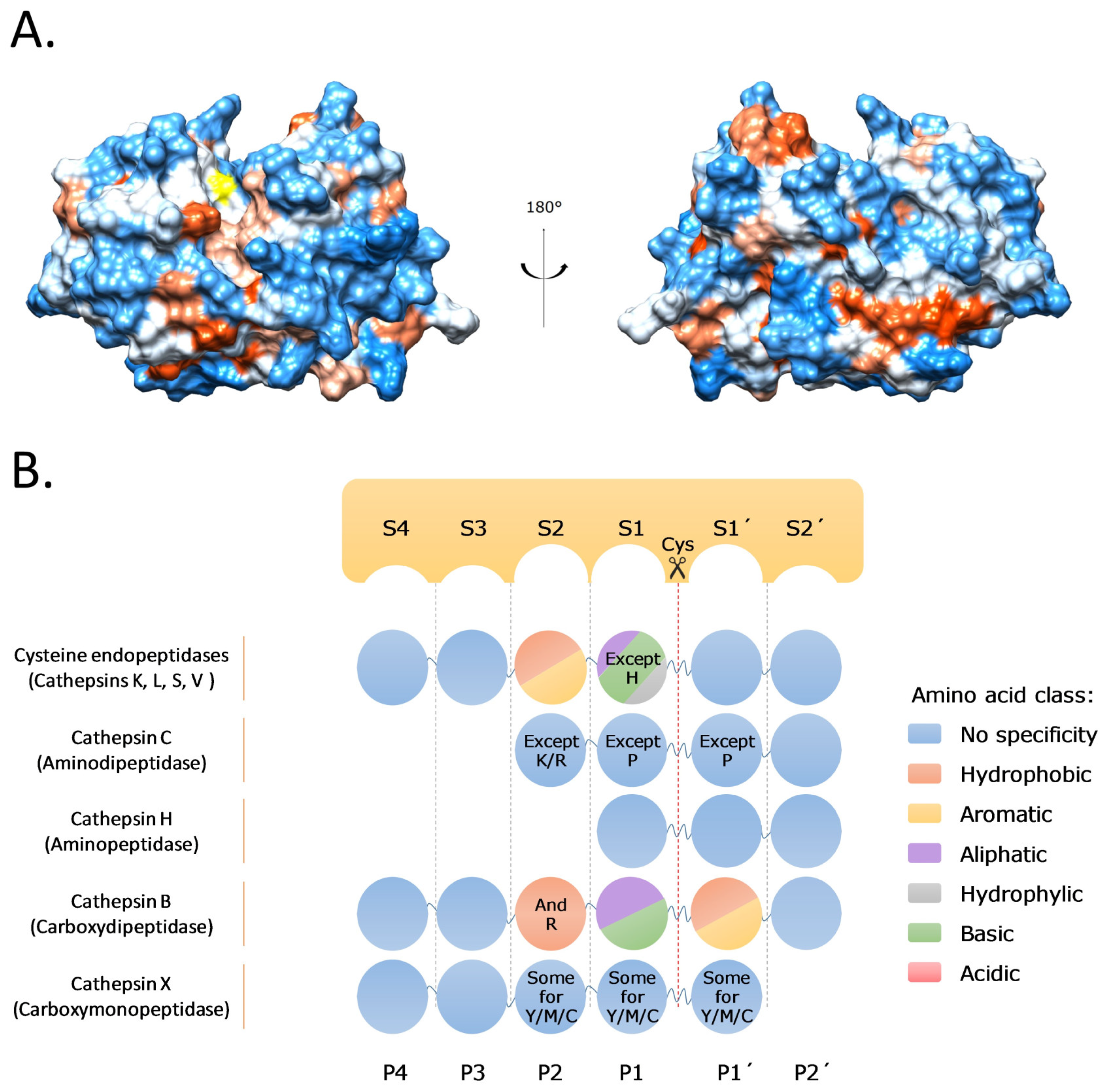
Cells | Free Full-Text | Cysteine Cathepsins and Their Extracellular Roles: Shaping the Microenvironment

Trivalent metal complex geometry of the substrate governs cathepsin B enzymatic cleavage rate - Chemical Communications (RSC Publishing)

TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells | Life Science Alliance
![PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/694d662801bcbb00cd4c5638a9a6d7ac44b518cf/4-Figure2-1.png)
PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar

Computational predictions of cysteine cathepsin‐mediated fibrinogen proteolysis - Ferrall‐Fairbanks - 2018 - Protein Science - Wiley Online Library
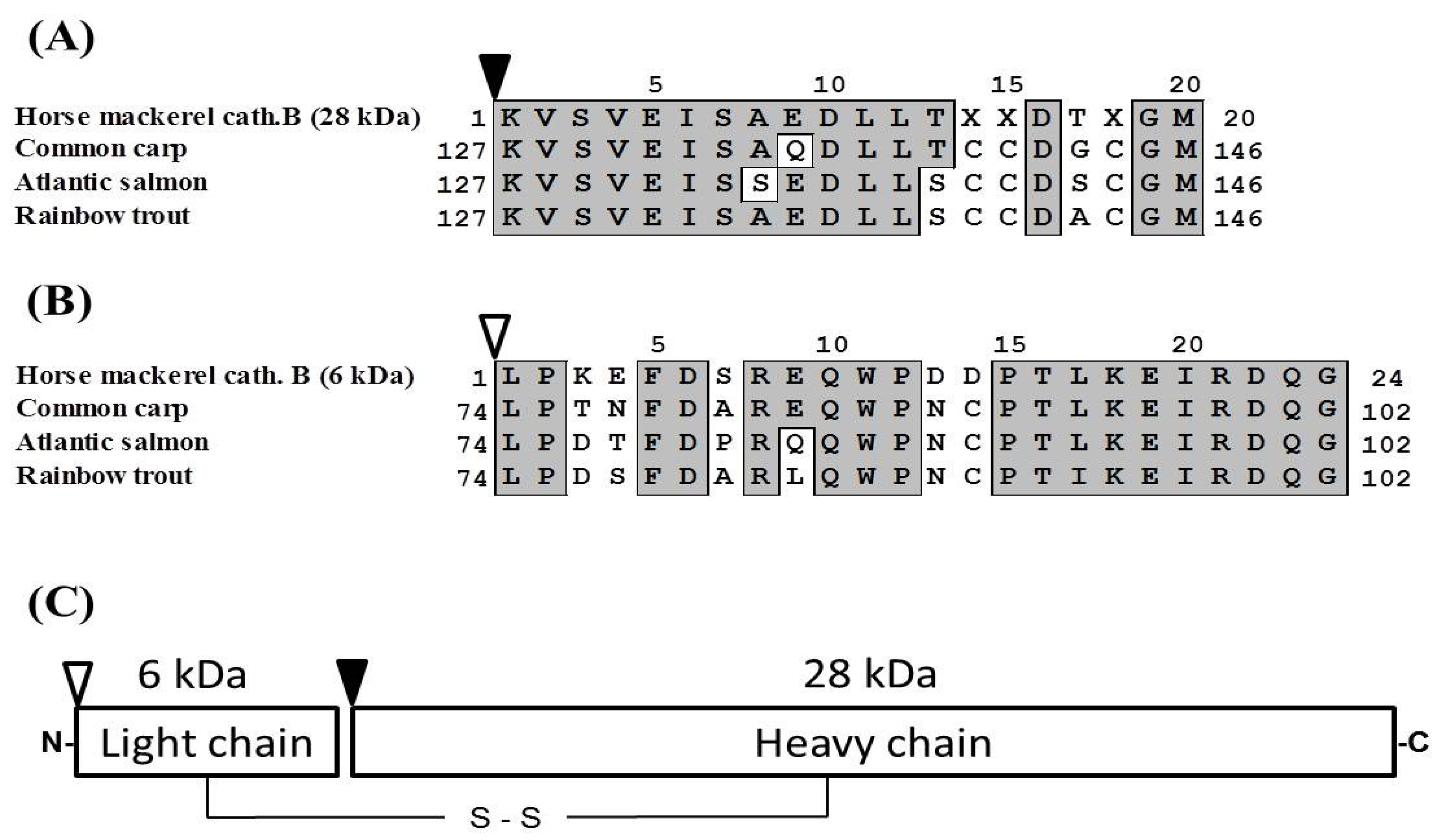
Marine Drugs | Free Full-Text | Purification and Characterization of Cathepsin B from the Muscle of Horse Mackerel Trachurus japonicus

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal
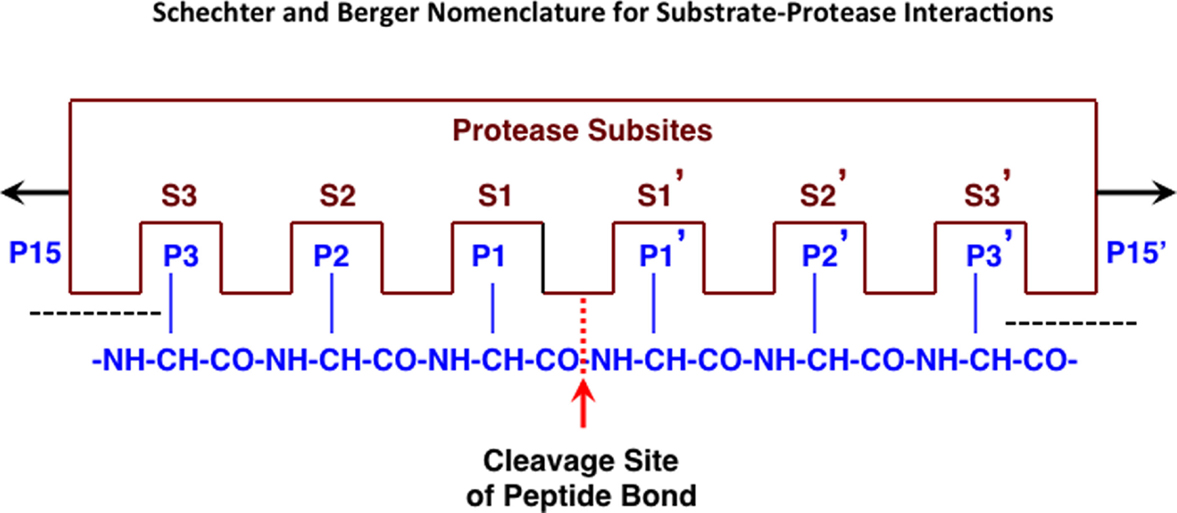
Frontiers | Cathepsin B is a New Drug Target for Traumatic Brain Injury Therapeutics: Evidence for E64d as a Promising Lead Drug Candidate
Workflow to analyze cathepsin B substrate cleavage site preferences for... | Download Scientific Diagram

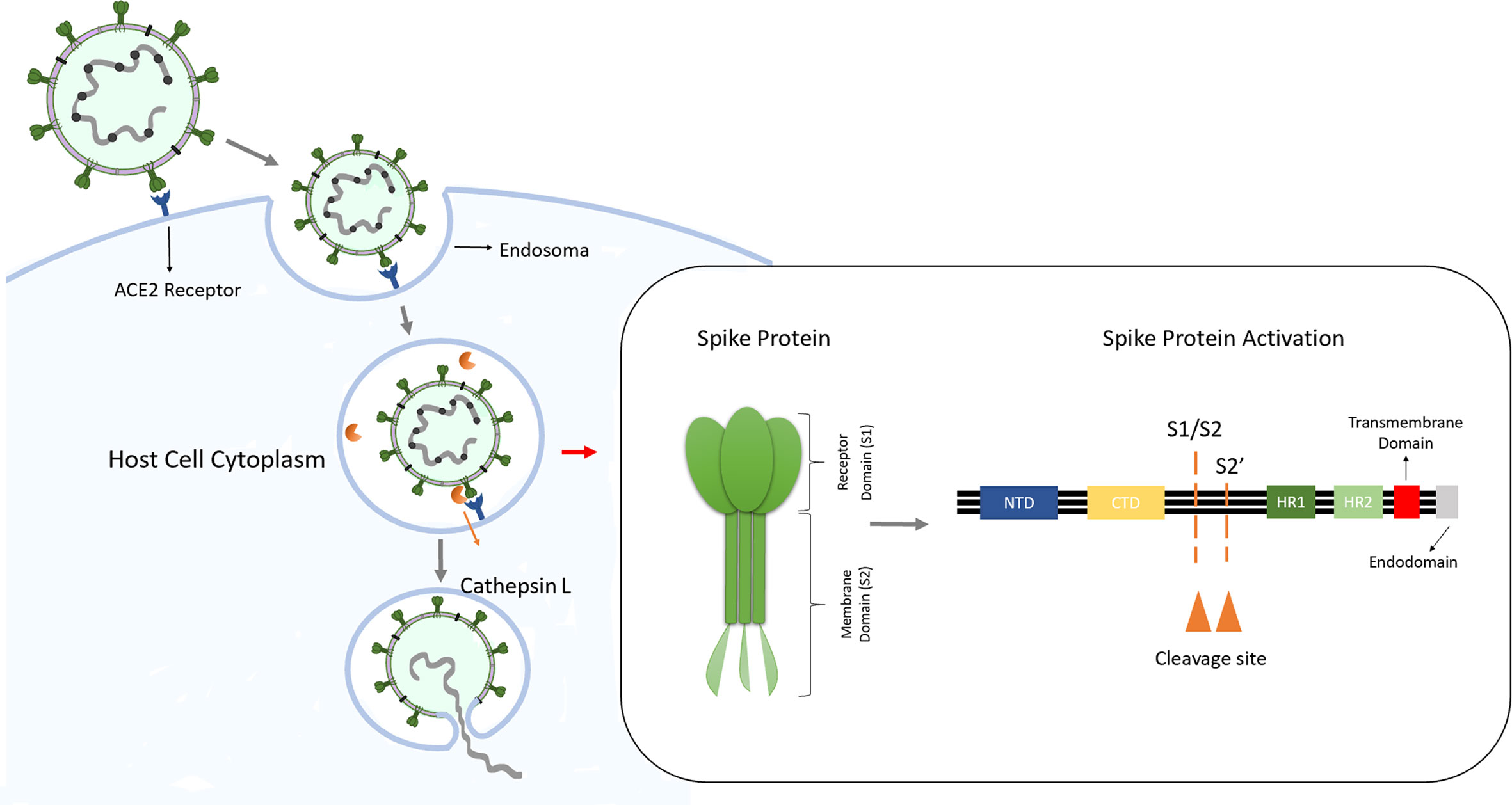
Novel cleavage sites identified in SARS-CoV-2 spike protein reveal mechanism for cathepsin L-facilitated viral infection and treatment strategies | Cell Discovery