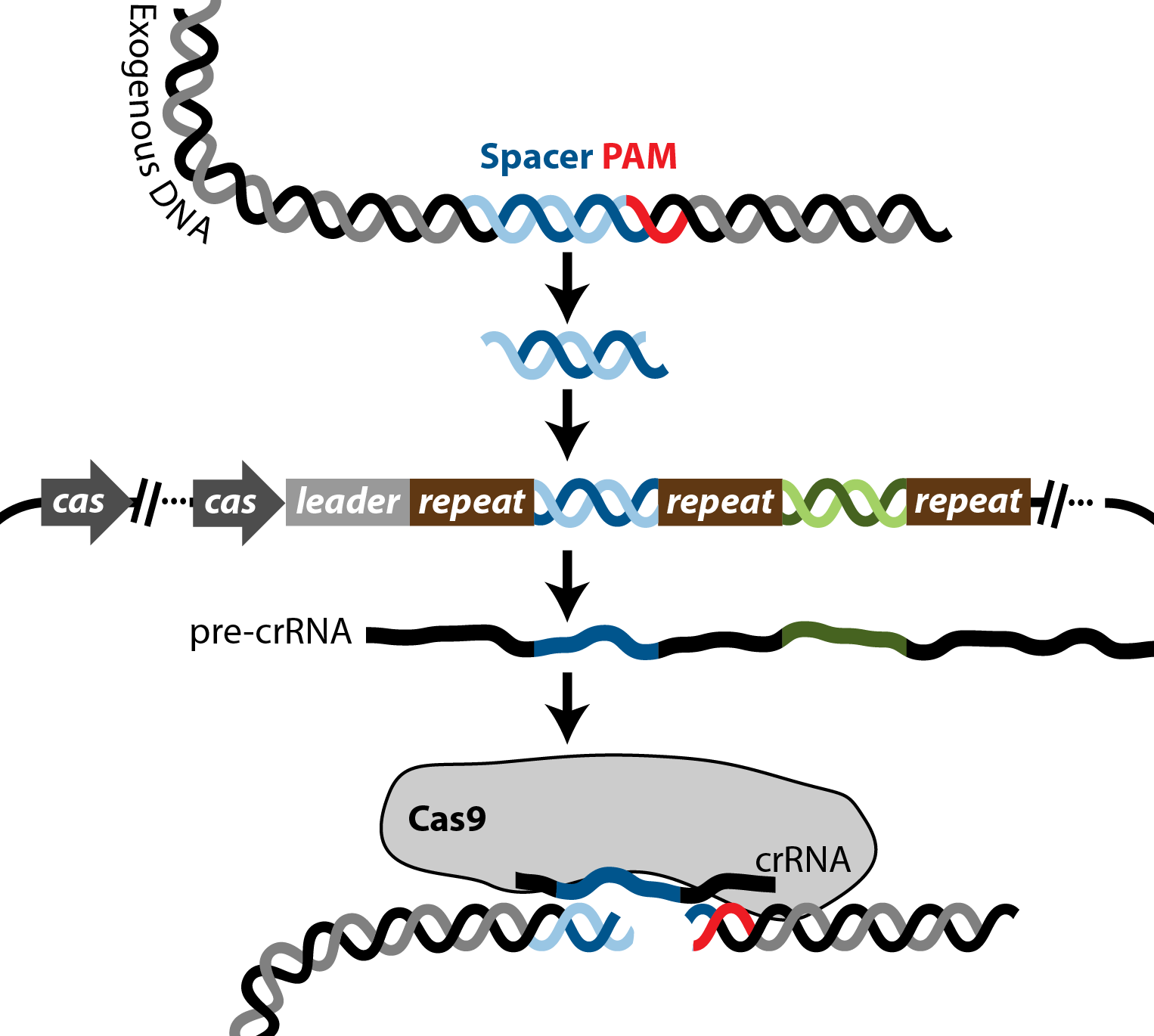
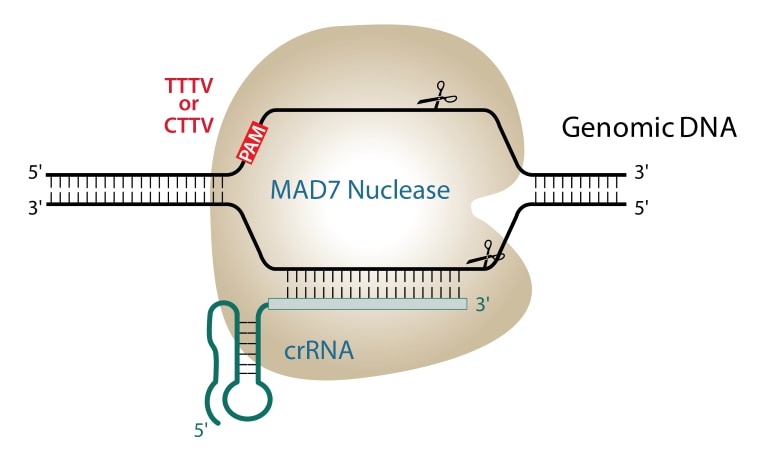
CRISPRseek: A Bioconductor Package to Identify Target-Specific Guide RNAs for CRISPR-Cas9 Genome-Editing Systems | PLOS ONE
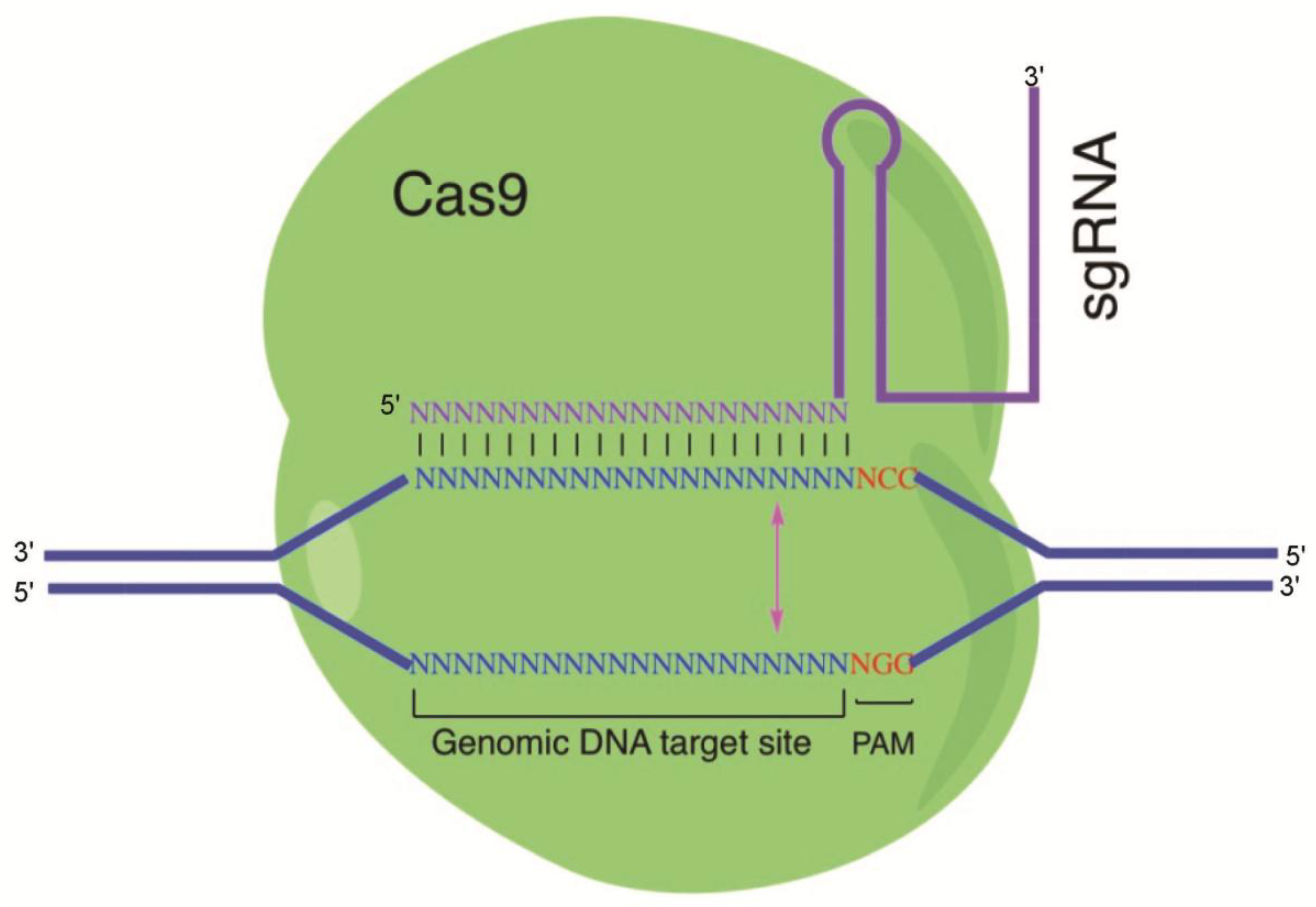
Viruses | Free Full-Text | Potential Application of the CRISPR/Cas9 System against Herpesvirus Infections

Validate CRISPR-Edited Cells using Imaging and Western Blot Detection on a Microplate Reader | Molecular Devices

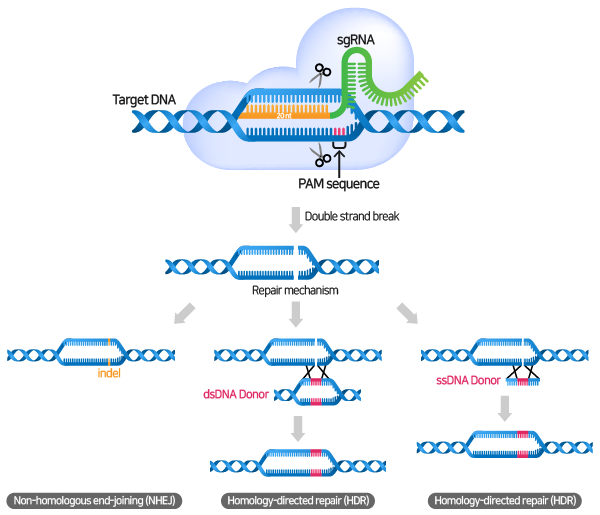
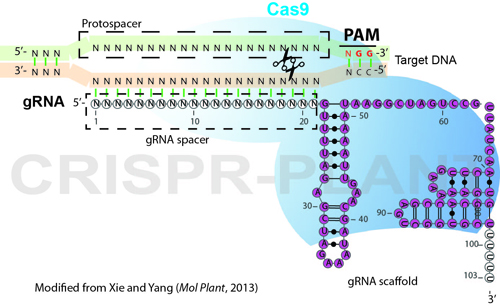
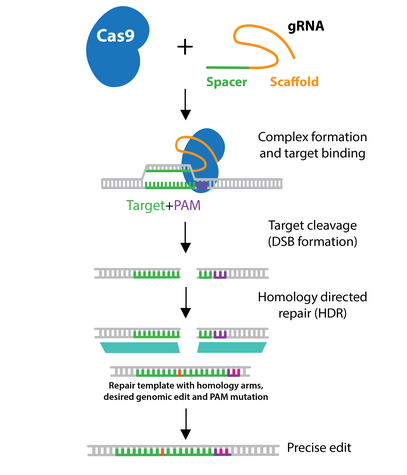
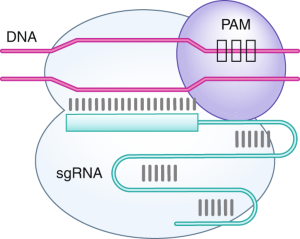
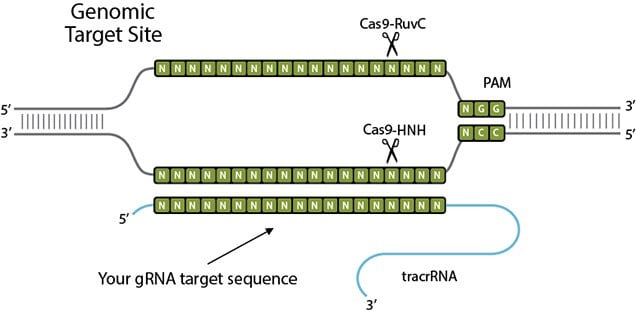
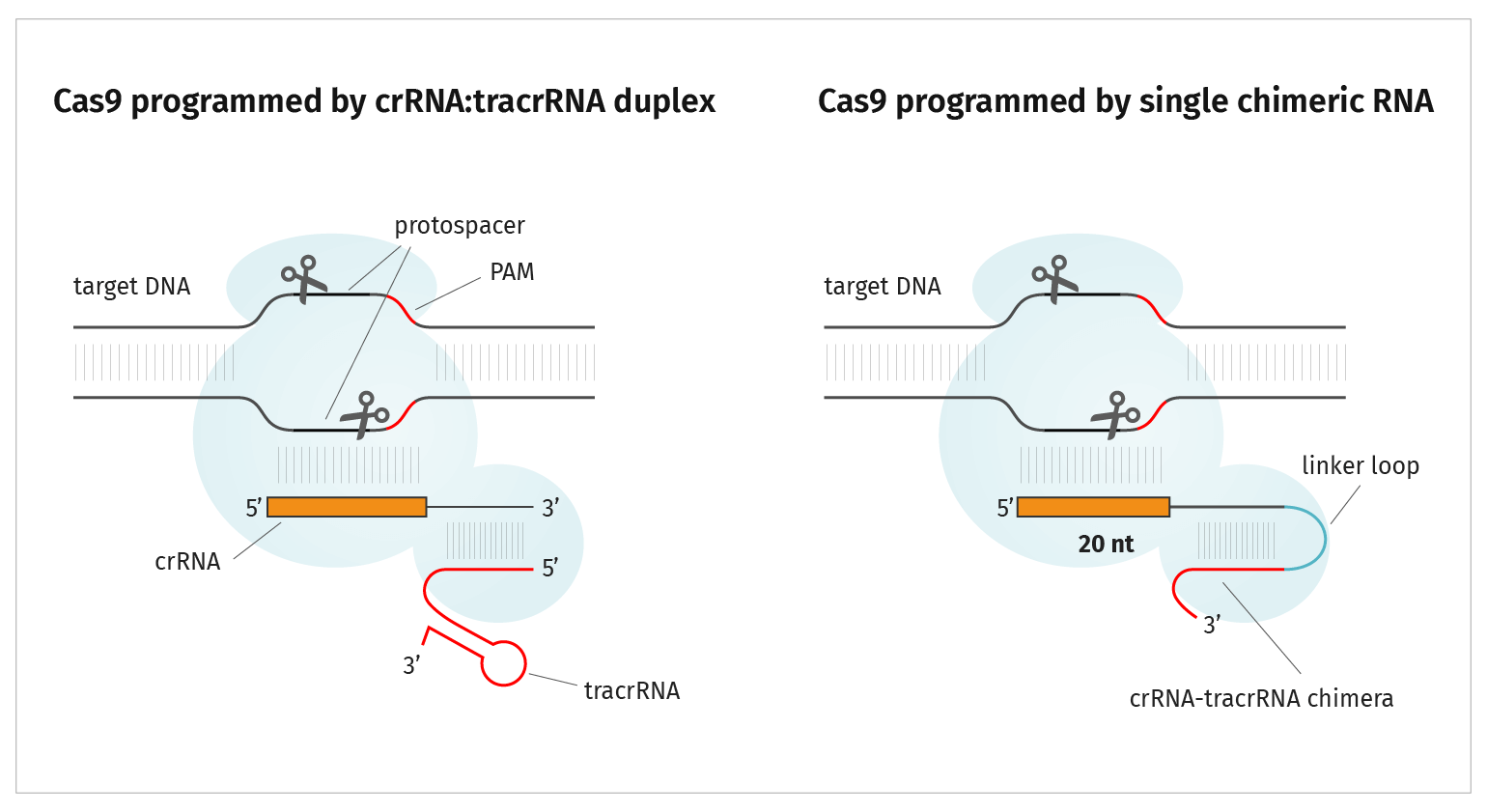
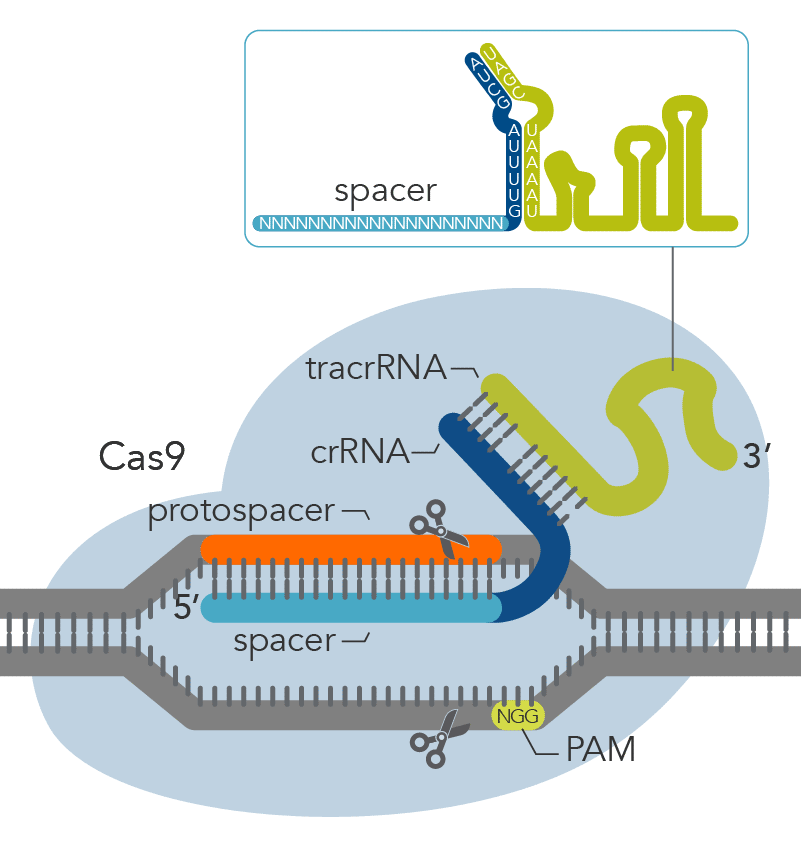
CRISPR-Cas9 mechanism. a A single guide RNA (sgRNA) is a fusion between... | Download Scientific Diagram
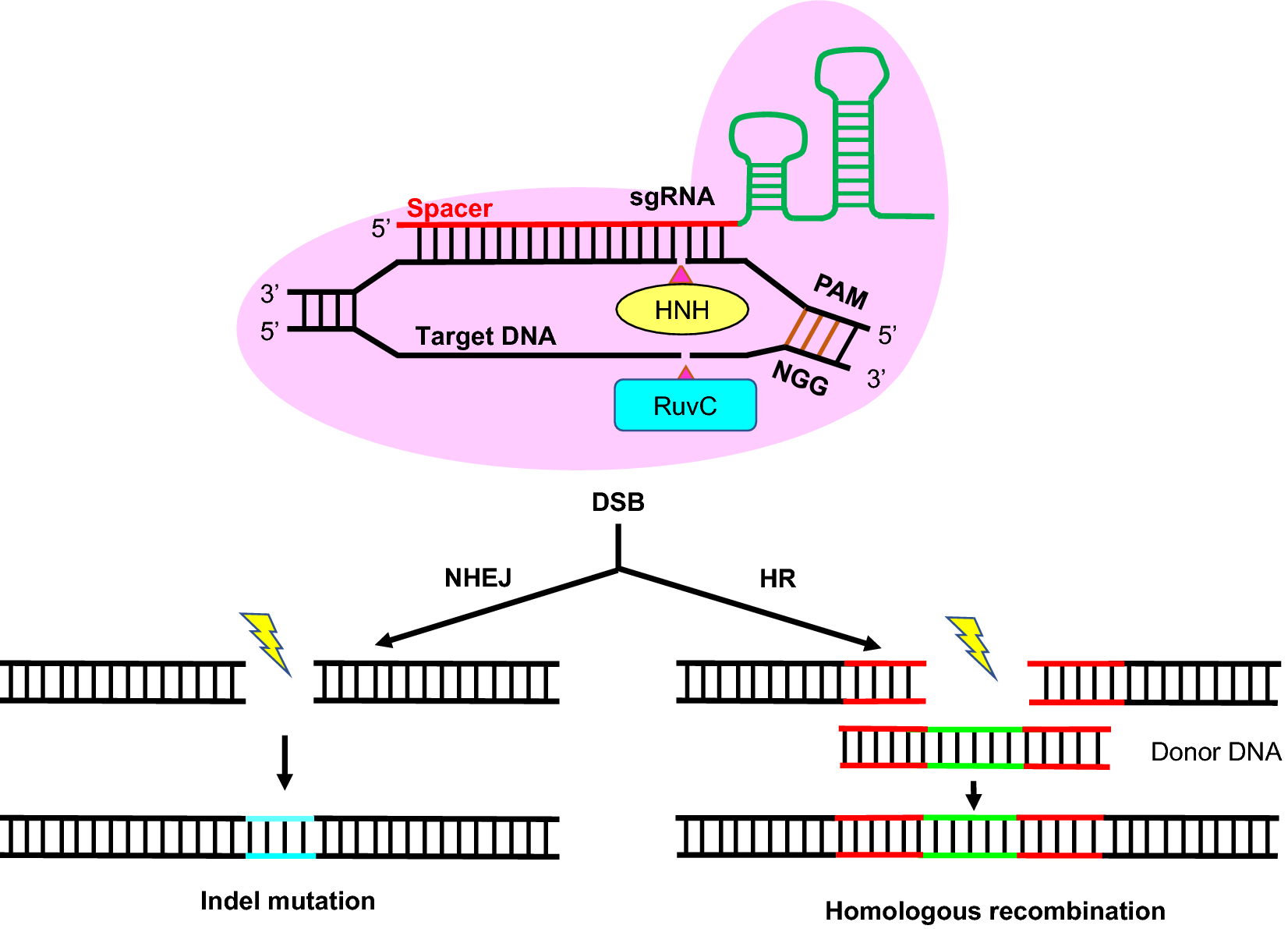
CRISPR-mediated genome editing in non-conventional yeasts for biotechnological applications | Microbial Cell Factories | Full Text
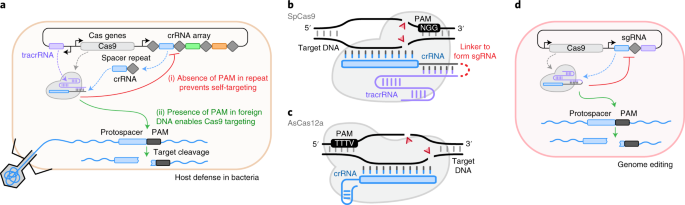







.png?sfvrsn=58c63b07_0)