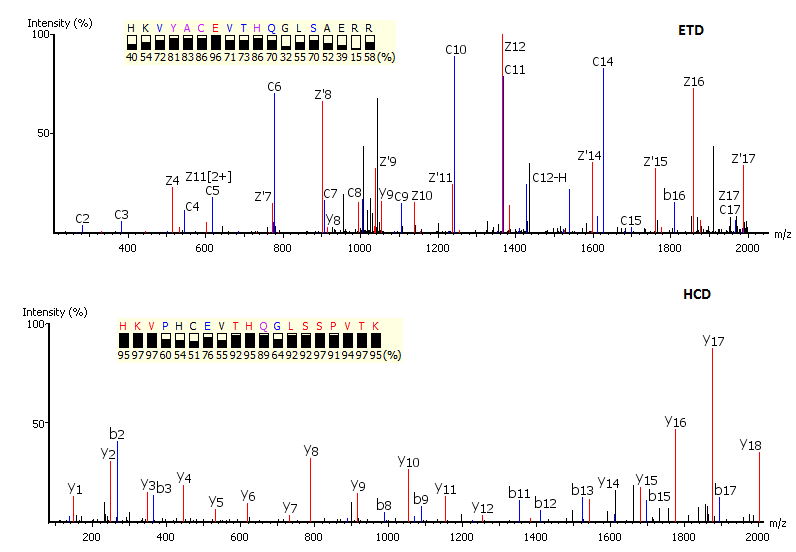
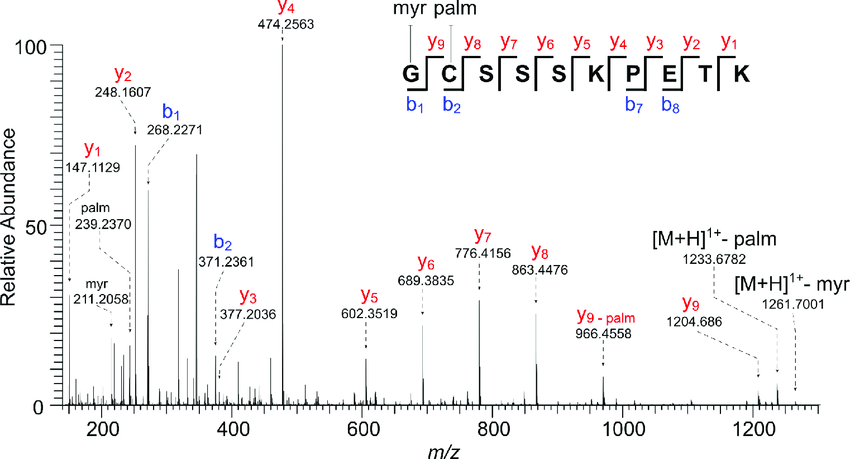
Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics - ScienceDirect
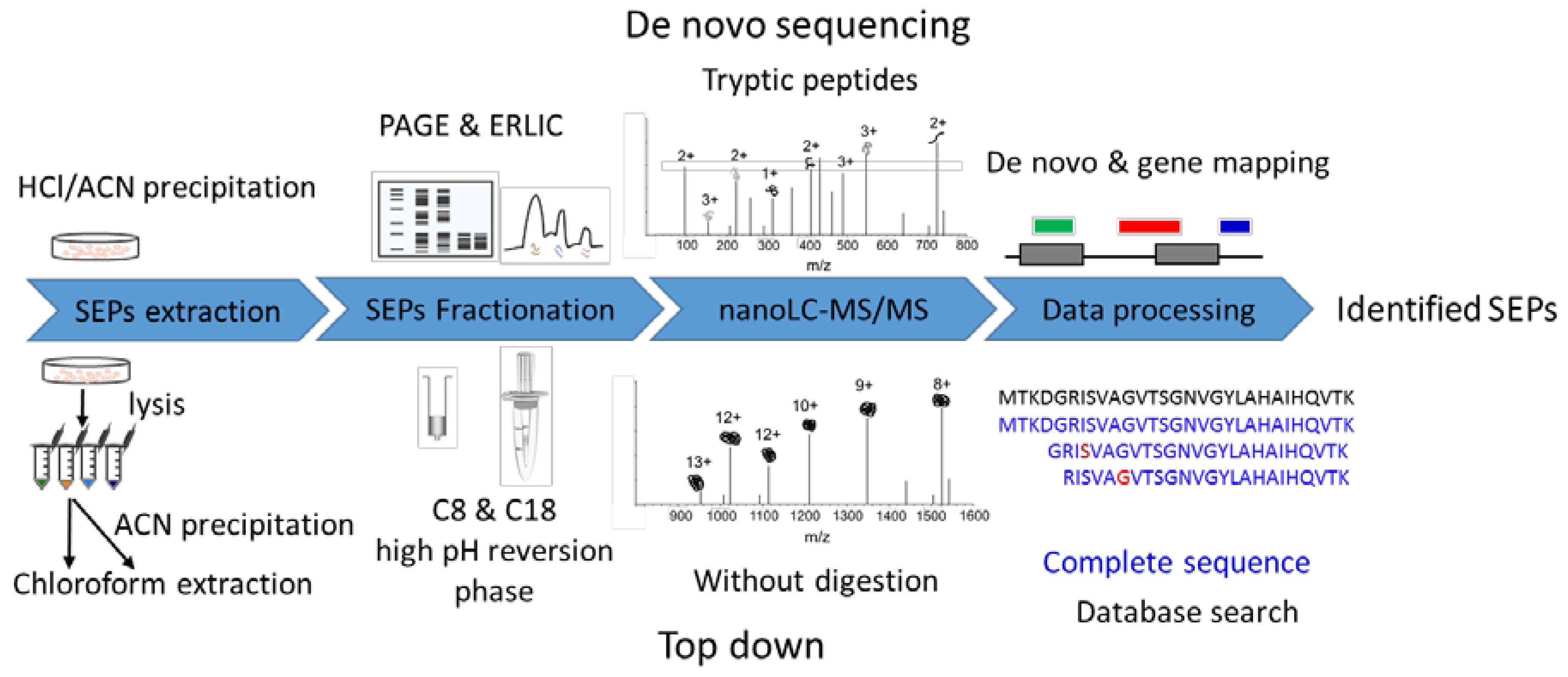
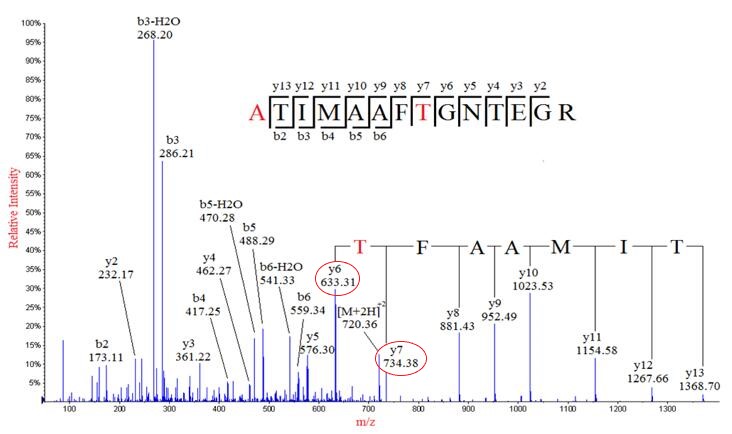
IJMS | Free Full-Text | Improved Identification of Small Open Reading Frames Encoded Peptides by Top-Down Proteomic Approaches and De Novo Sequencing

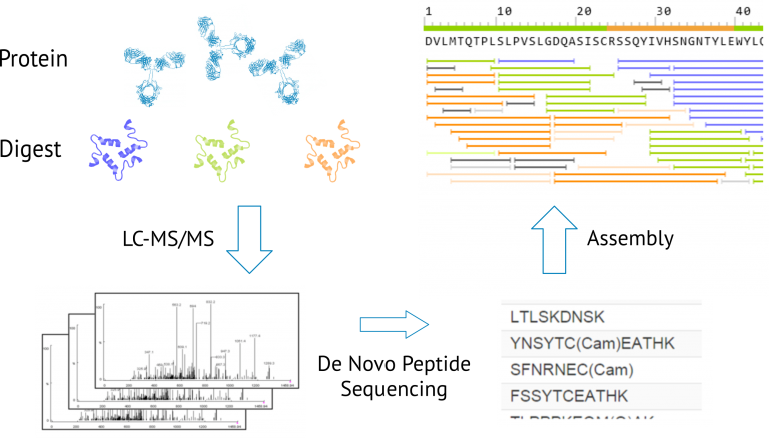
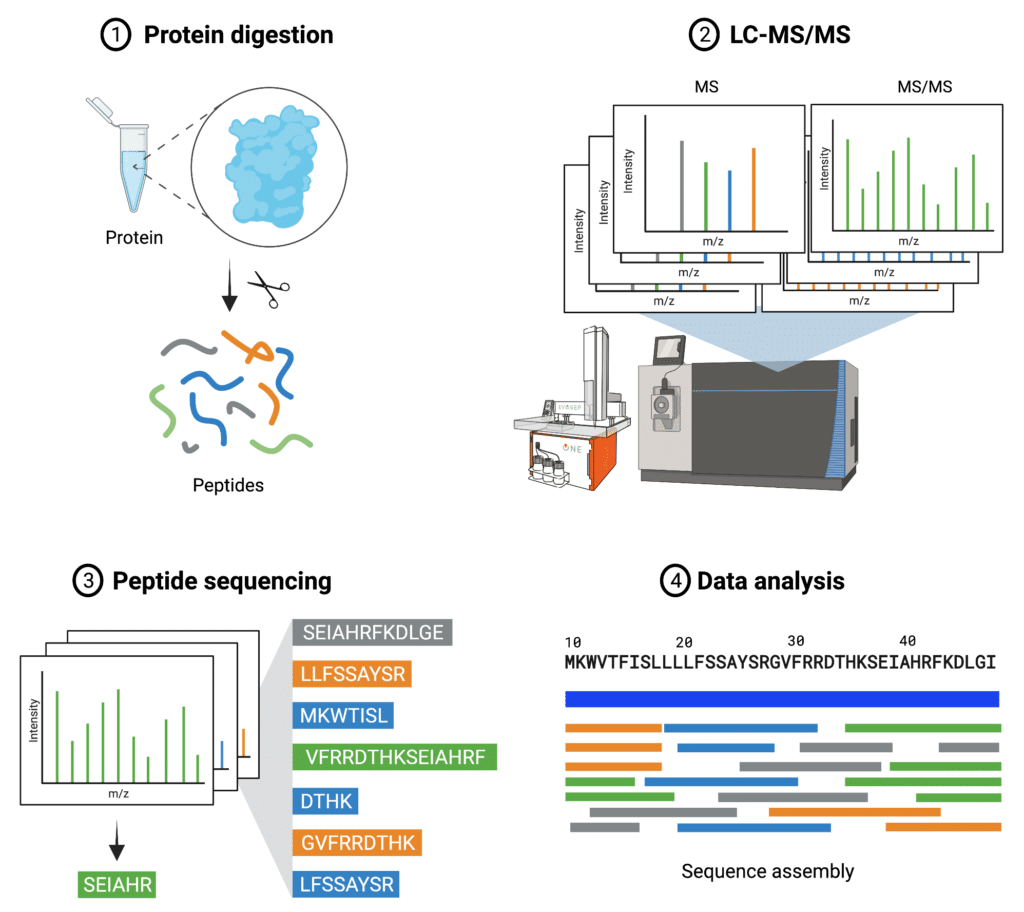
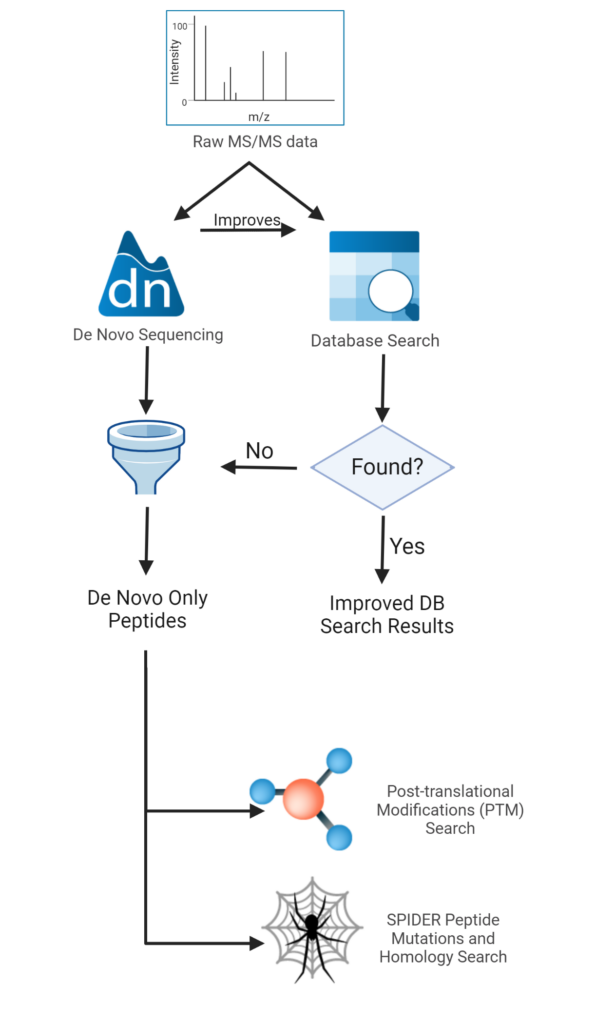
Pipeline involved in Peptide Sequencing using Tandem Mass Spectrometry. | Download Scientific Diagram