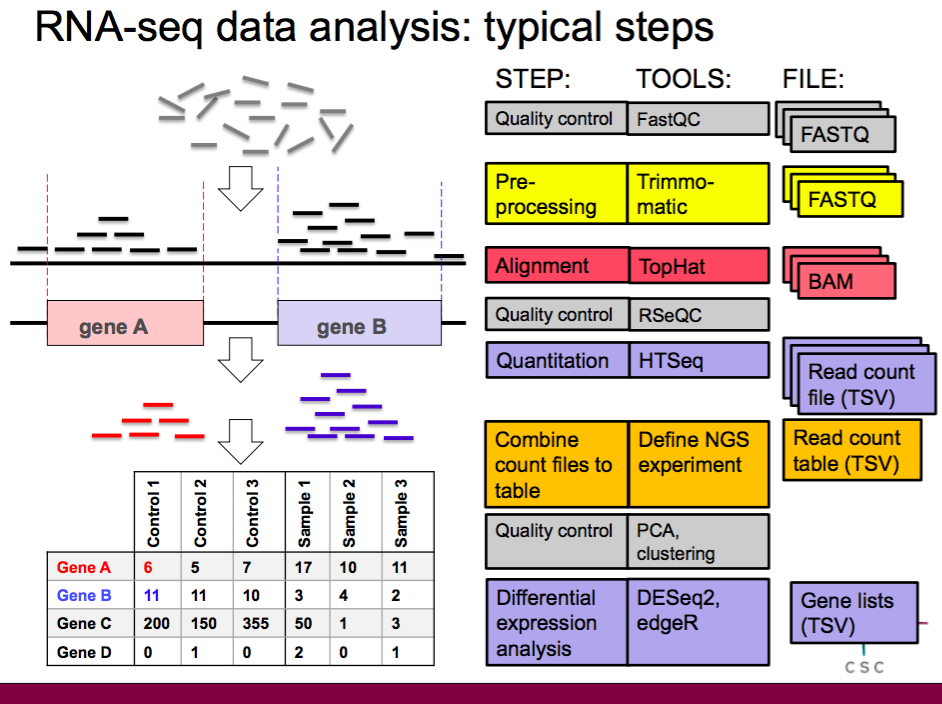
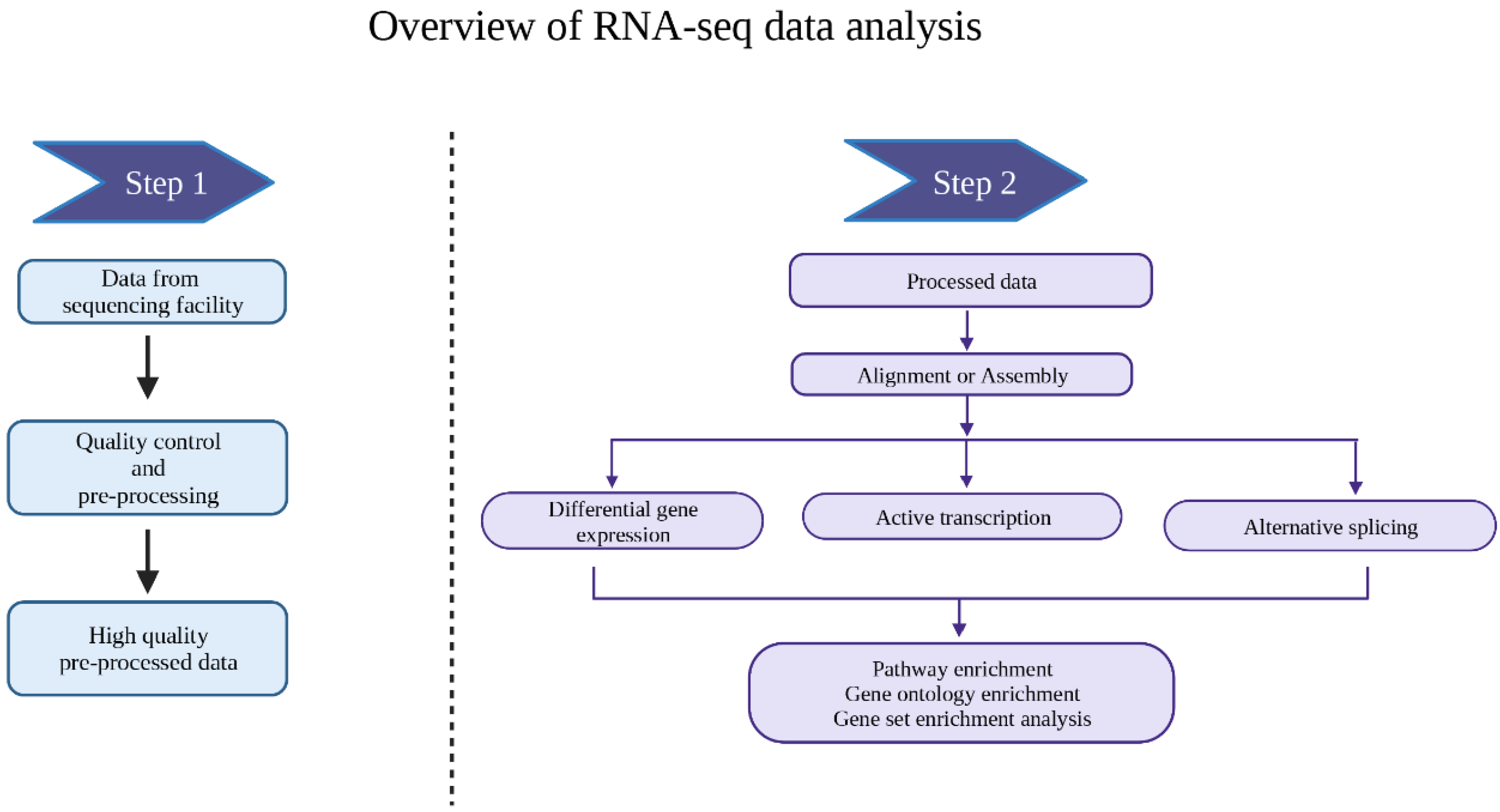
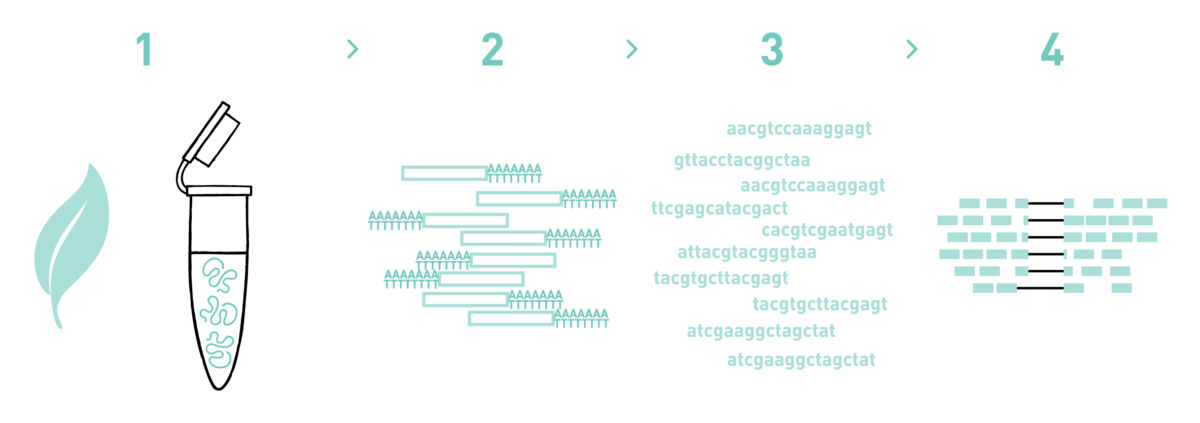
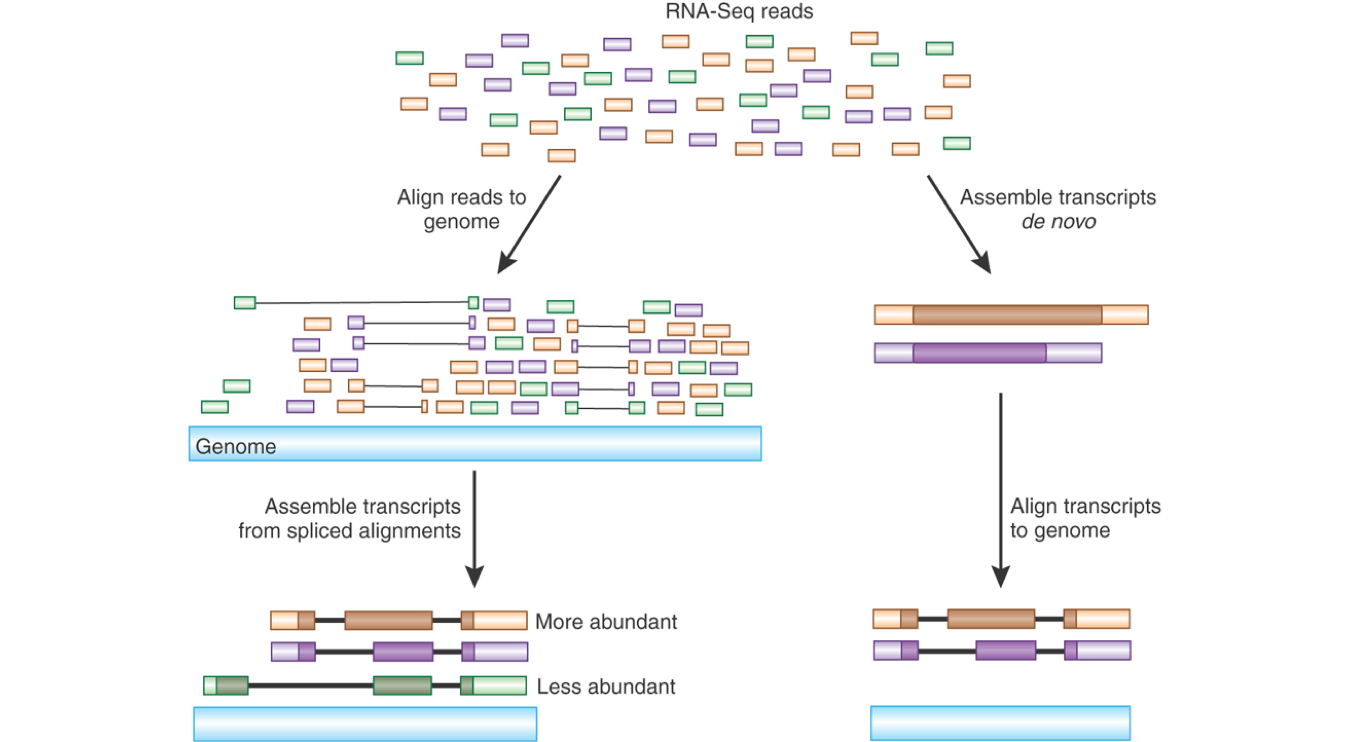
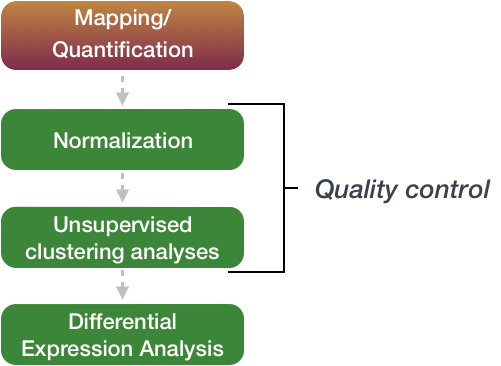
Differential RNA-seq of Vibrio cholerae identifies the VqmR small RNA as a regulator of biofilm formation | PNAS
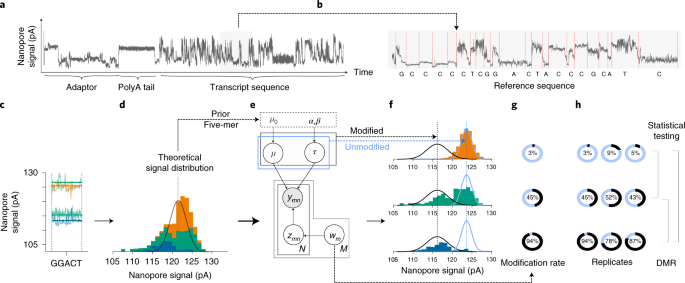
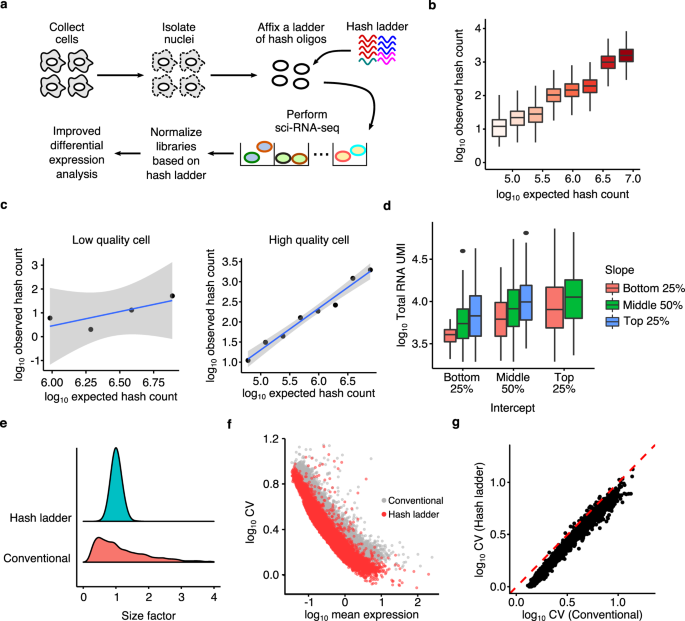
Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore | Nature Biotechnology

Biosensors | Free Full-Text | A Comparison of Methods for RNA-Seq Differential Expression Analysis and a New Empirical Bayes Approach
Genome context and expression of brrF. A) Coverage from differential... | Download Scientific Diagram
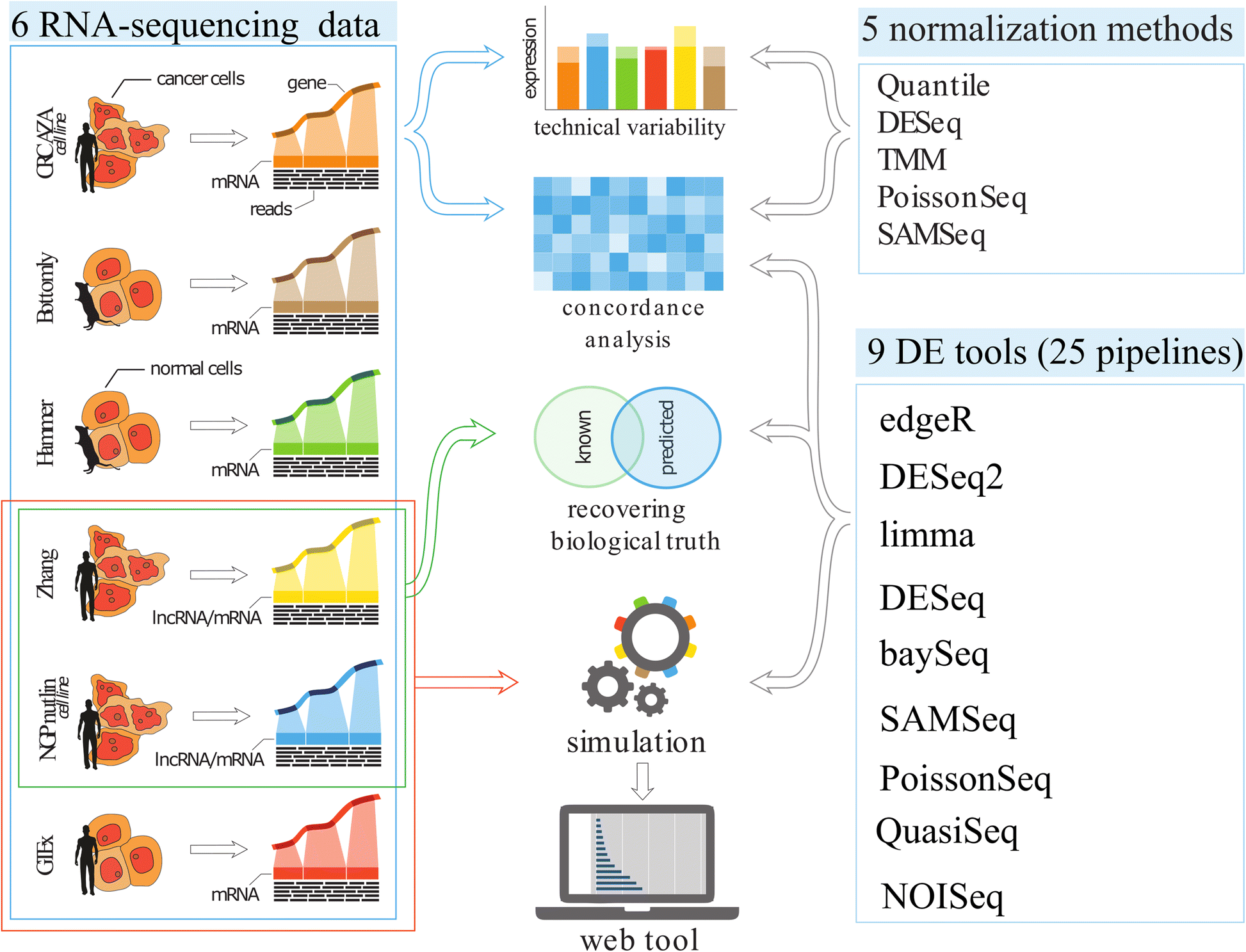
Differential gene expression analysis tools exhibit substandard performance for long non-coding RNA-sequencing data | Genome Biology | Full Text
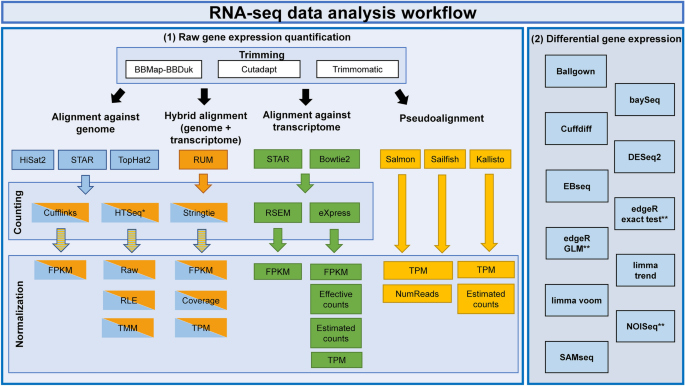
Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis | Scientific Reports
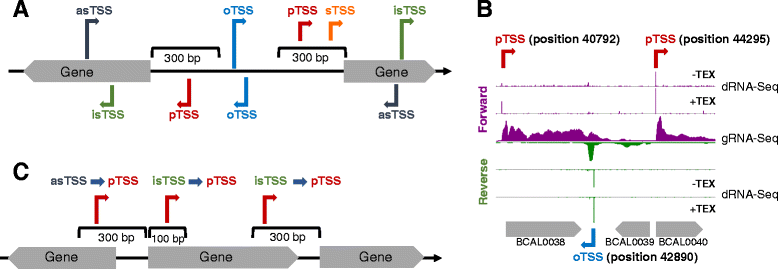
Genome-wide transcription start site profiling in biofilm-grown Burkholderia cenocepacia J2315 | BMC Genomics | Full Text


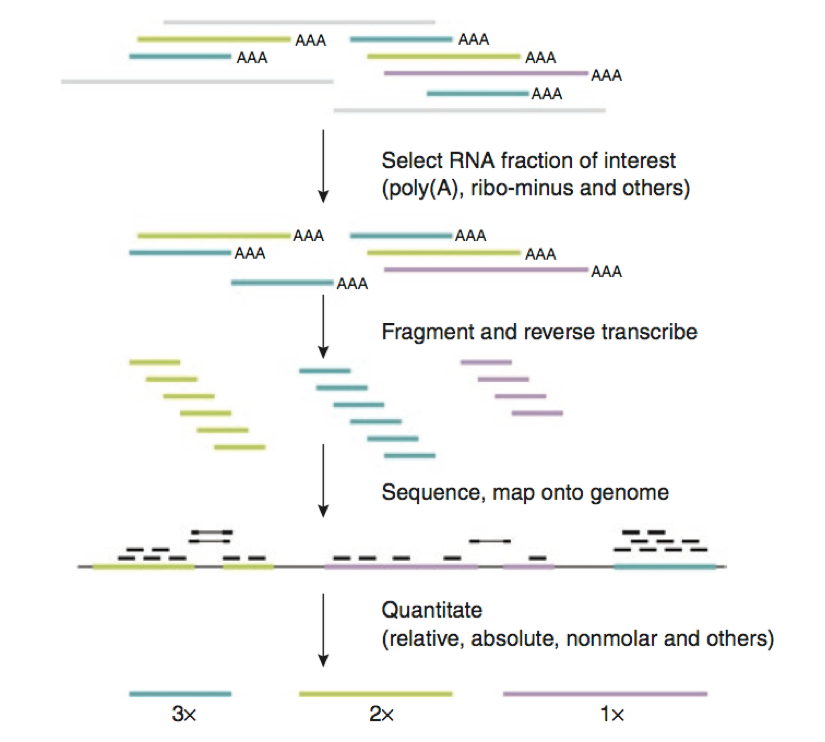
![PDF] RNA‐seq: Applications and Best Practices | Semantic Scholar PDF] RNA‐seq: Applications and Best Practices | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/54136f301fe8ae8ec0d5a67098c8587d728dfb23/11-Figure2-1.png)