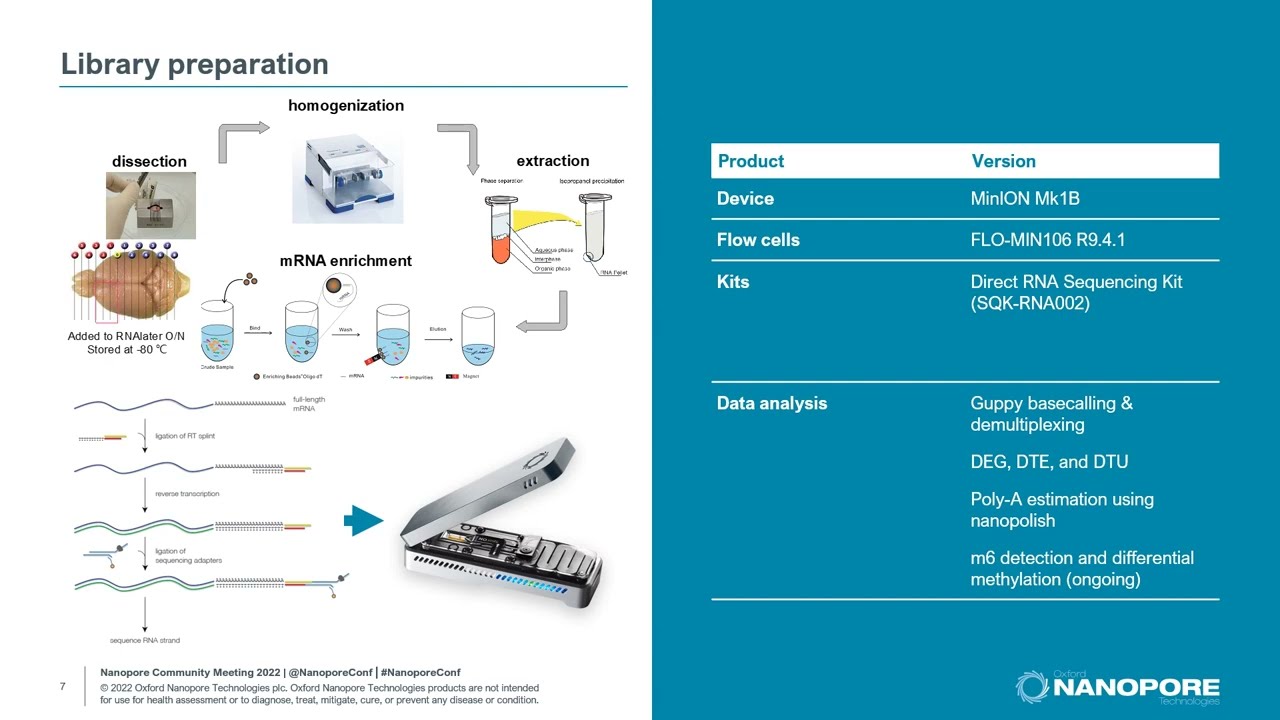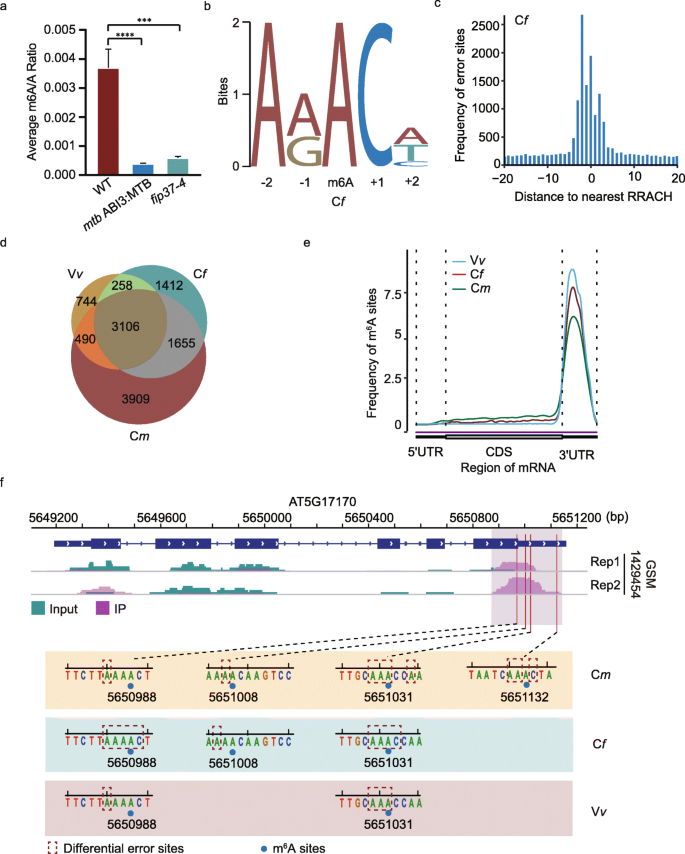
DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA | Genome Biology | Full Text

An overview of using direct RNA sequencing to detect RNA modifications.... | Download Scientific Diagram
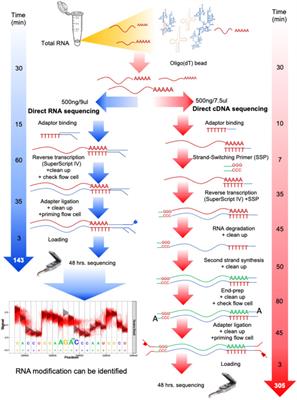
Frontiers | Native RNA or cDNA Sequencing for Transcriptomic Analysis: A Case Study on Saccharomyces cerevisiae
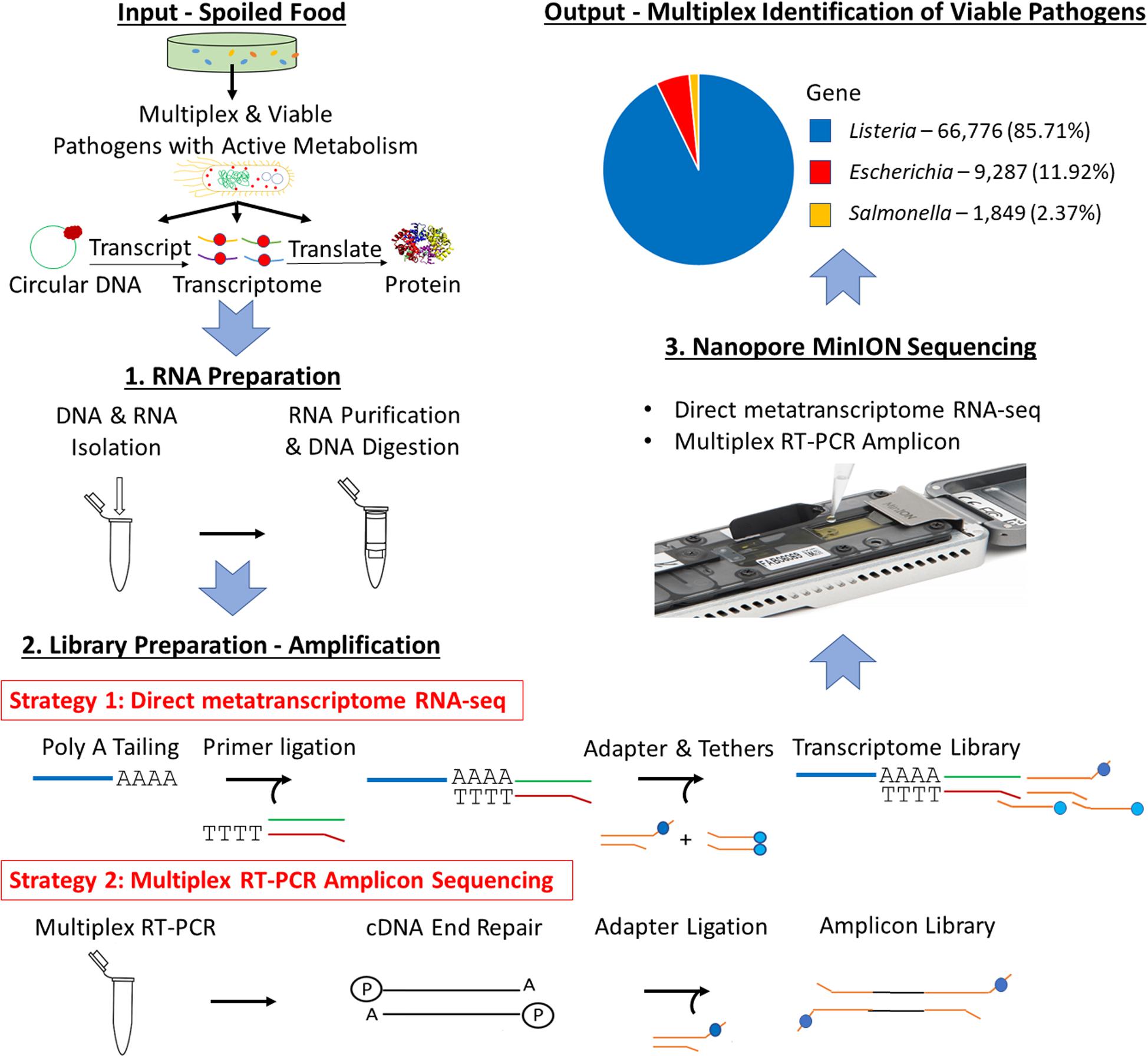
Frontiers | Direct Metatranscriptome RNA-seq and Multiplex RT-PCR Amplicon Sequencing on Nanopore MinION – Promising Strategies for Multiplex Identification of Viable Pathogens in Food

Oxford Nanopore on Twitter: "Direct RNA sequencing kits being shipped to some early users. Busy week. https://t.co/7yDOYrmLb3 https://t.co/1bnY5E79o1" / Twitter
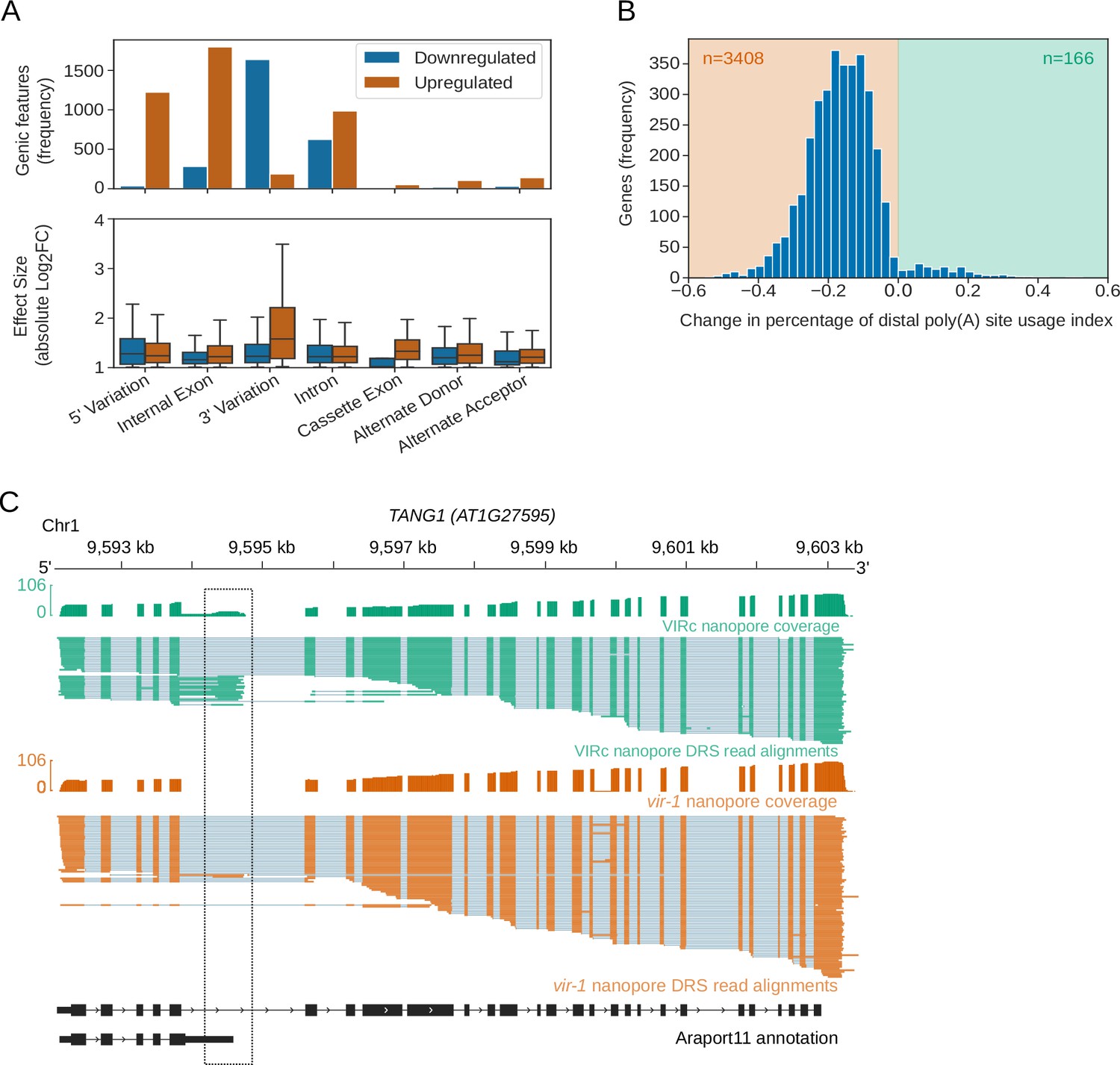
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife
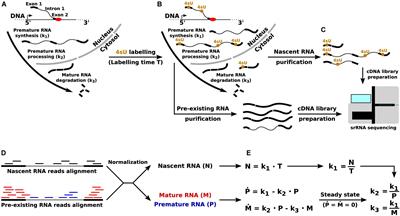
Frontiers | Direct RNA Sequencing for the Study of Synthesis, Processing, and Degradation of Modified Transcripts
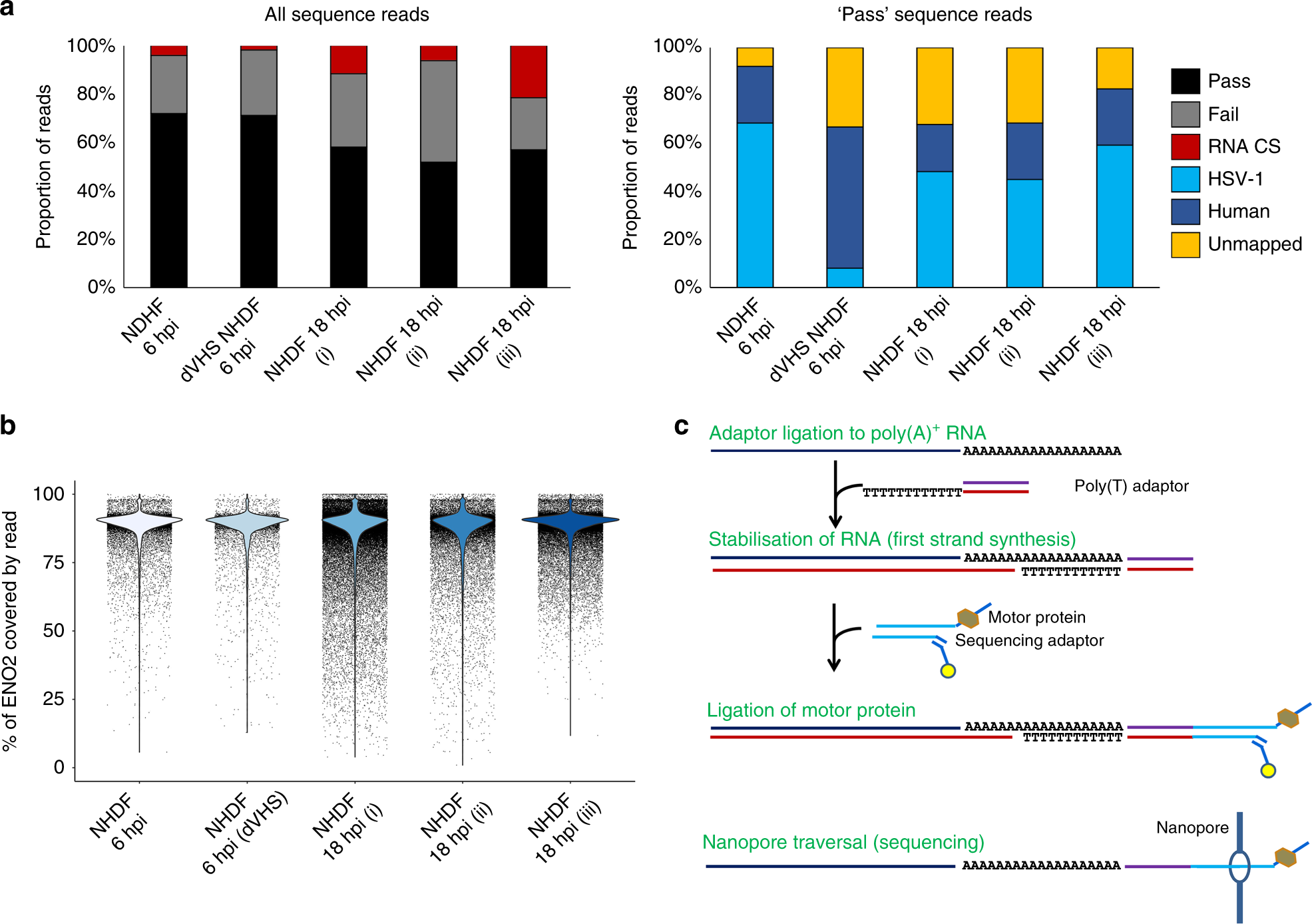
Direct RNA sequencing on nanopore arrays redefines the transcriptional complexity of a viral pathogen | Nature Communications
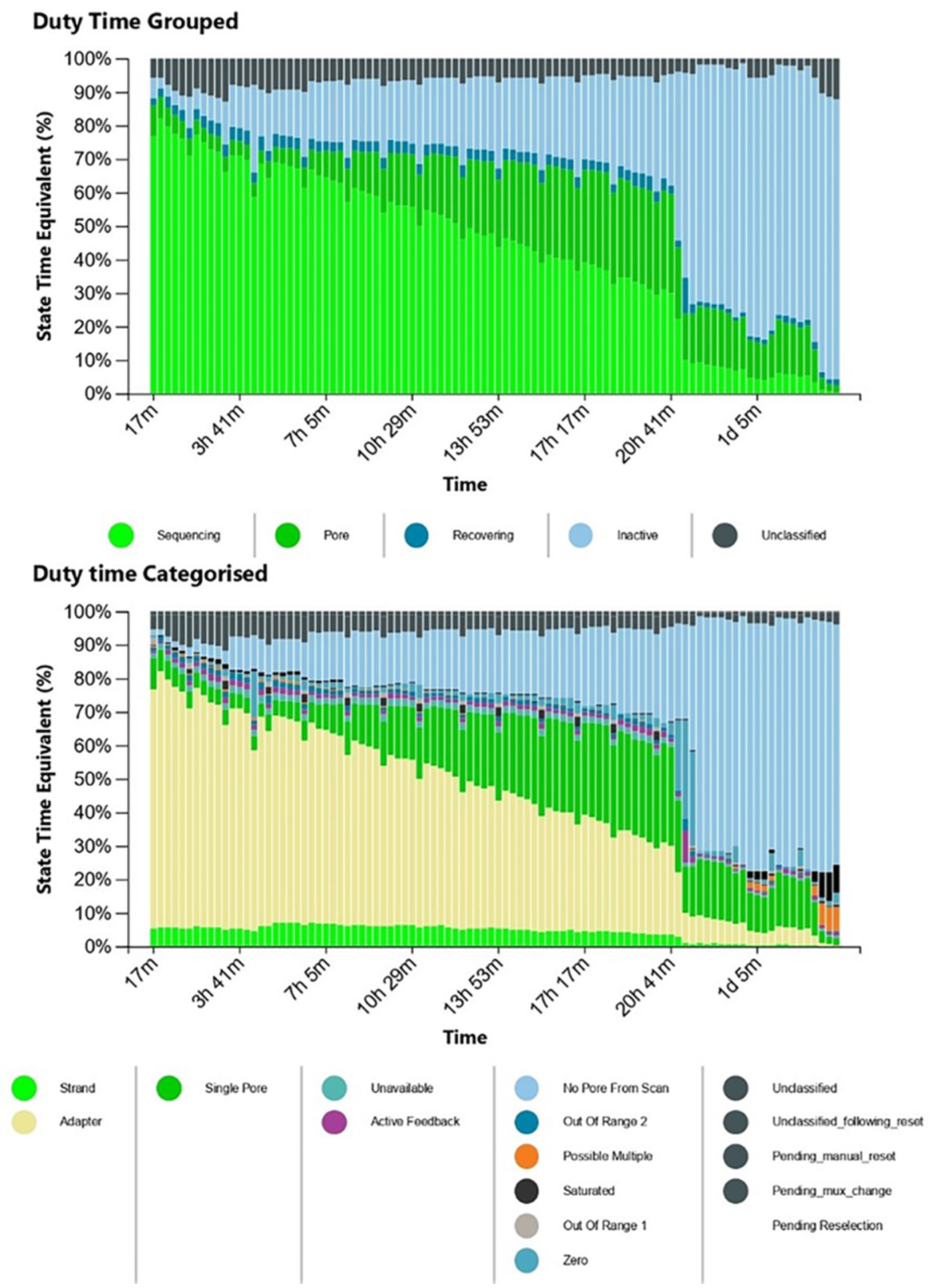
Life | Free Full-Text | Direct RNA Nanopore Sequencing of SARS-CoV-2 Extracted from Critical Material from Swabs
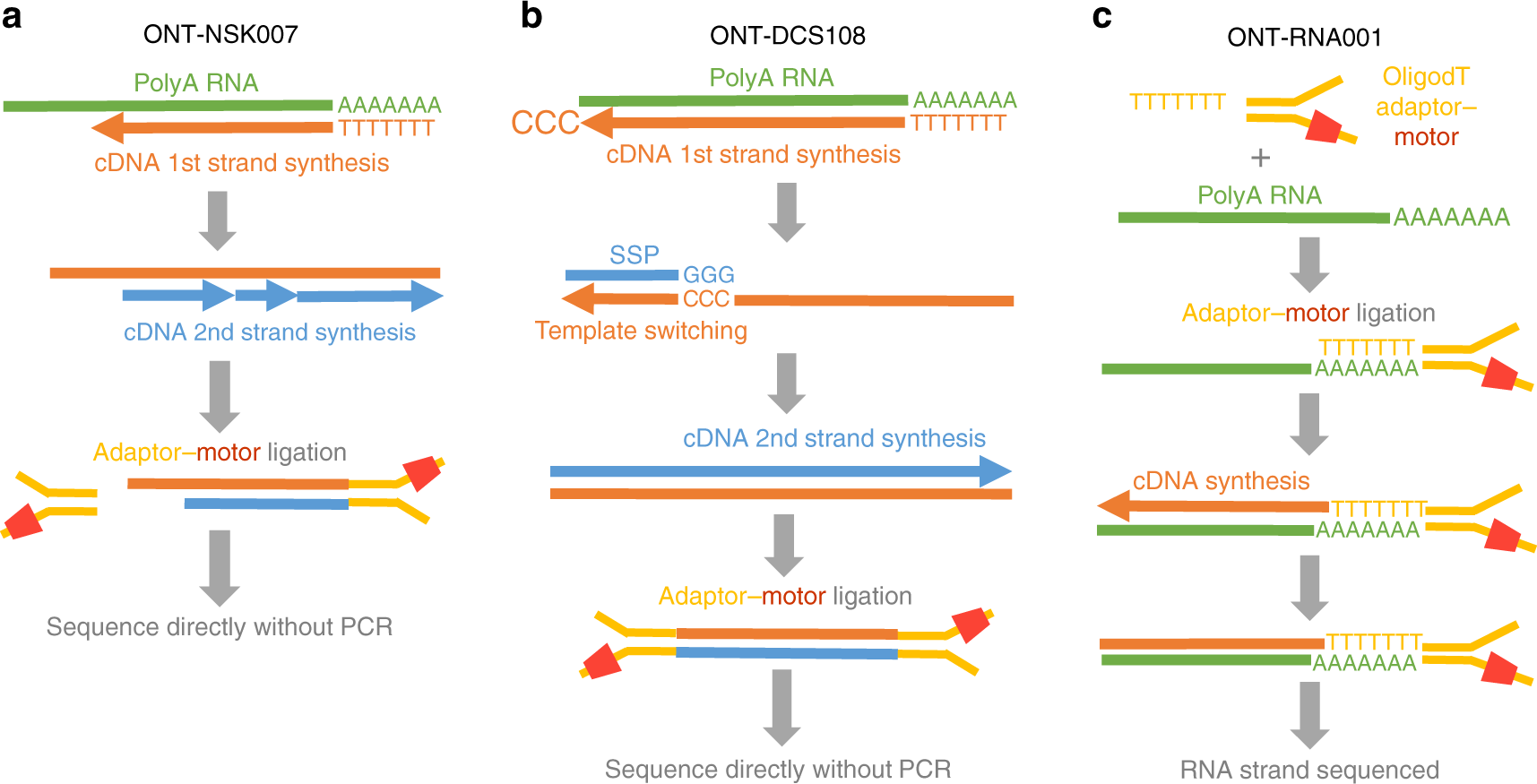
A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes | Nature Communications