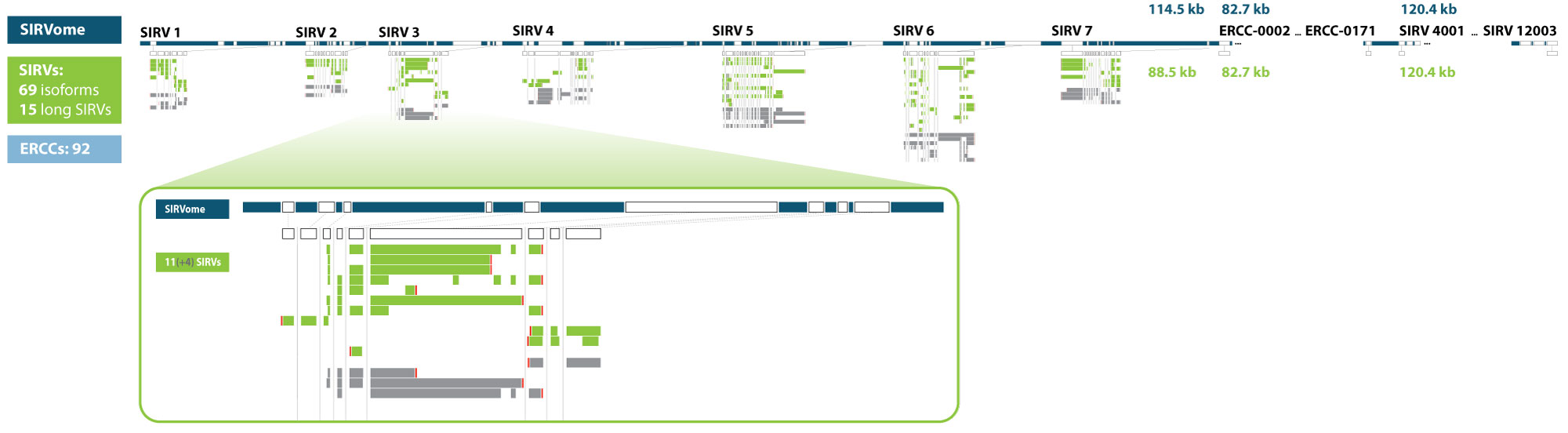
Strand-specific preparation of full-length PCR-based and PCR-free cDNA libraries by strand-switching

Benchmarking of the Oxford Nanopore MinION sequencing for quantitative and qualitative assessment of cDNA populations | RNA-Seq Blog

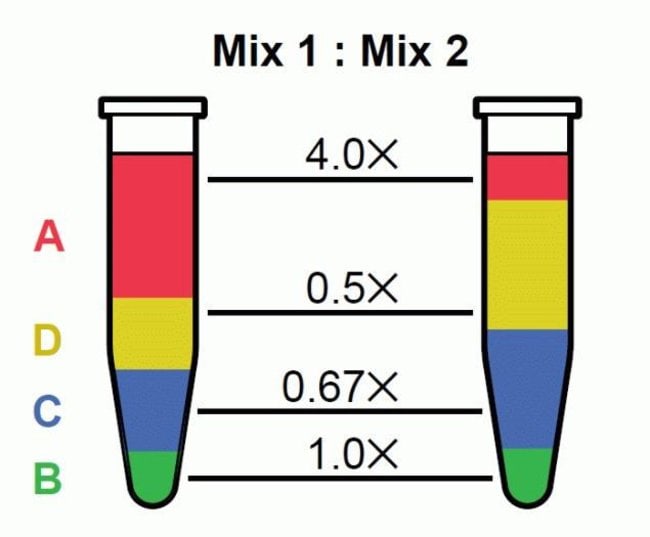
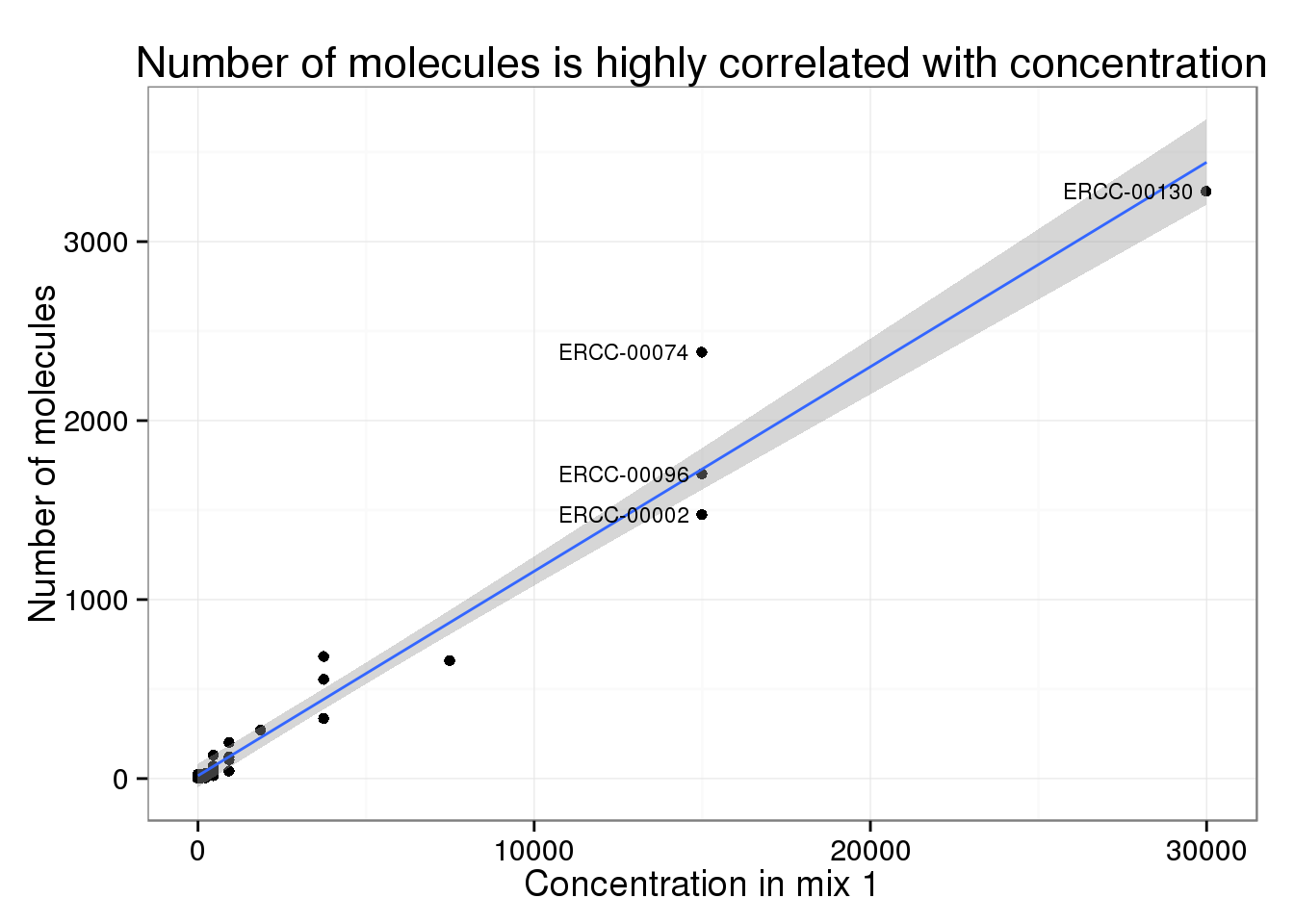
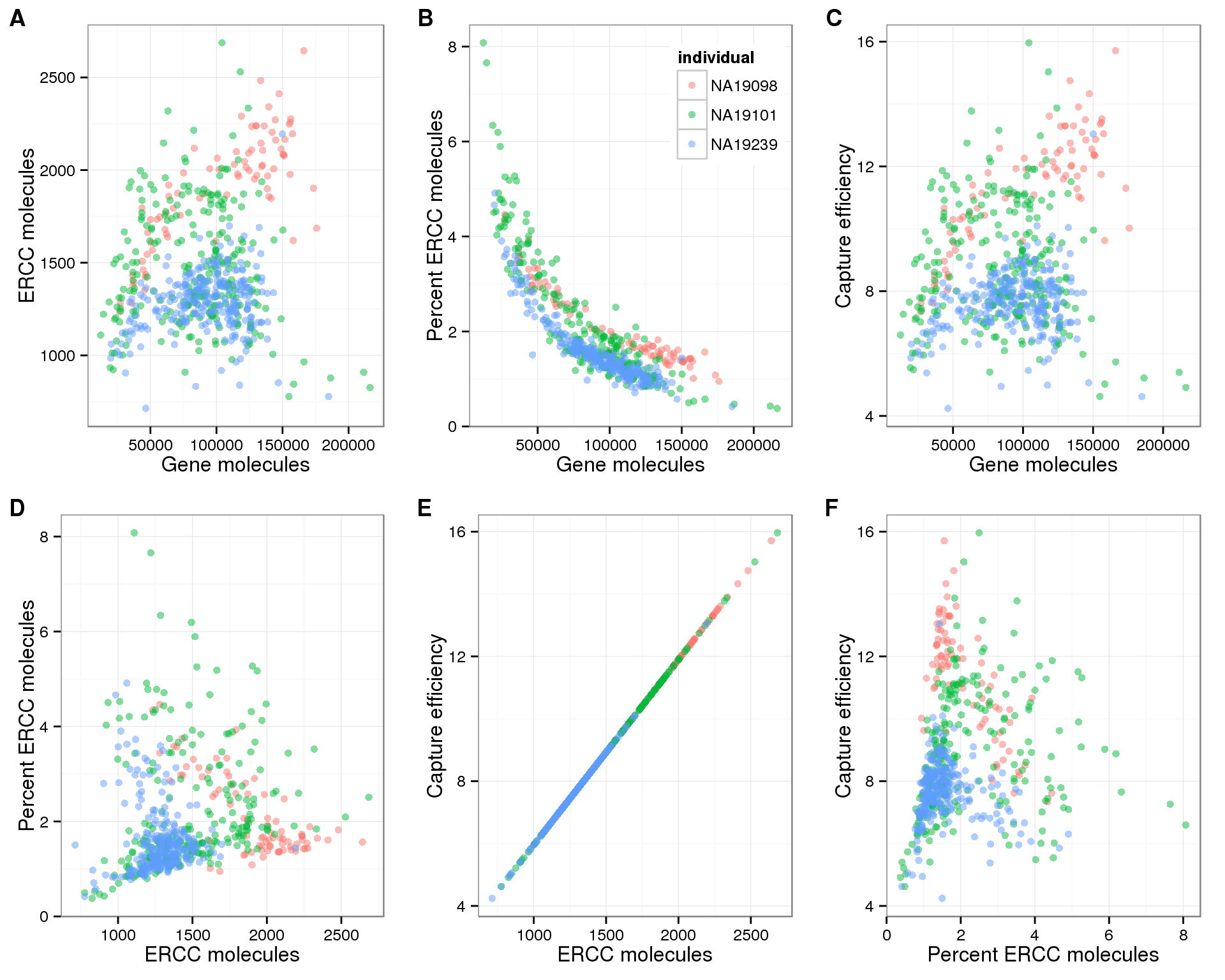
Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures | Nature Communications
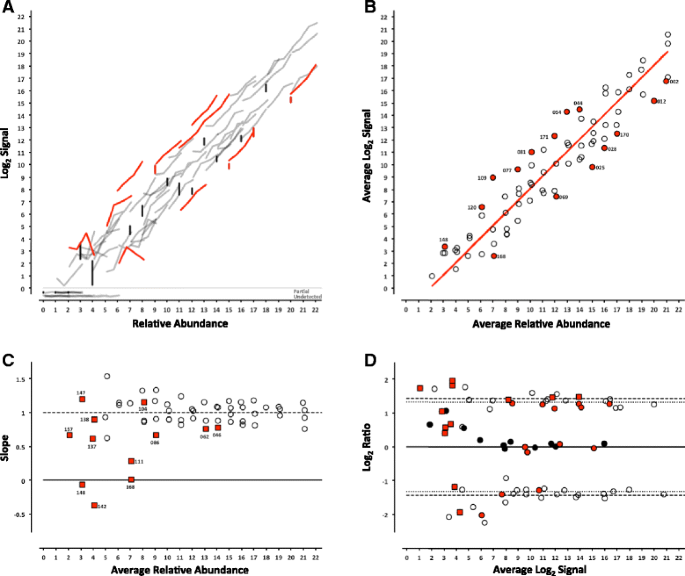
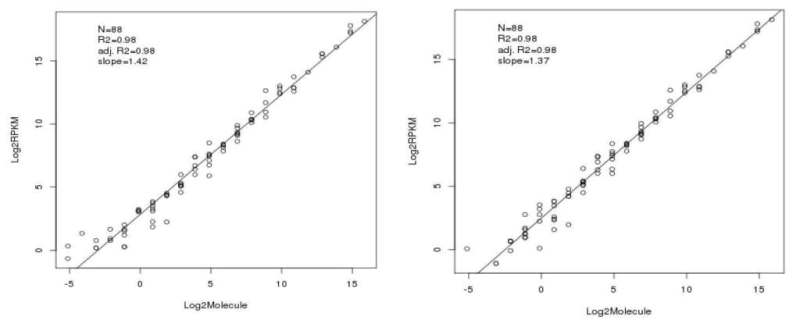
Evaluation of the External RNA Controls Consortium (ERCC) reference material using a modified Latin square design | BMC Biotechnology | Full Text
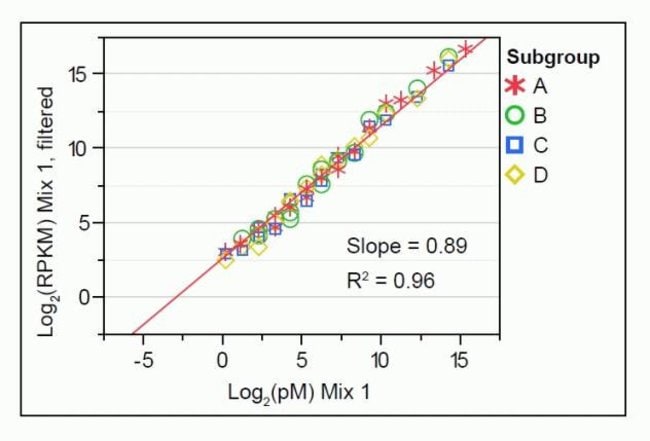
Using ERCC Spike-In Control Transcripts Provides Confidence in Agilent Microarray and RNA-Seq Gene Expression Data