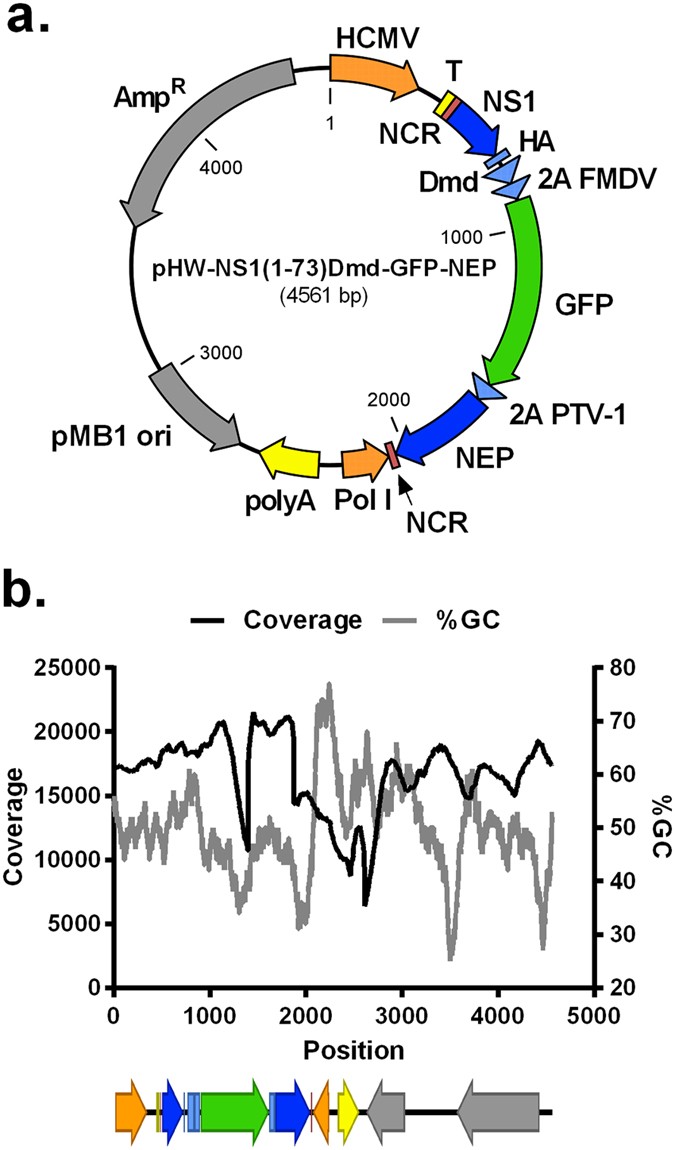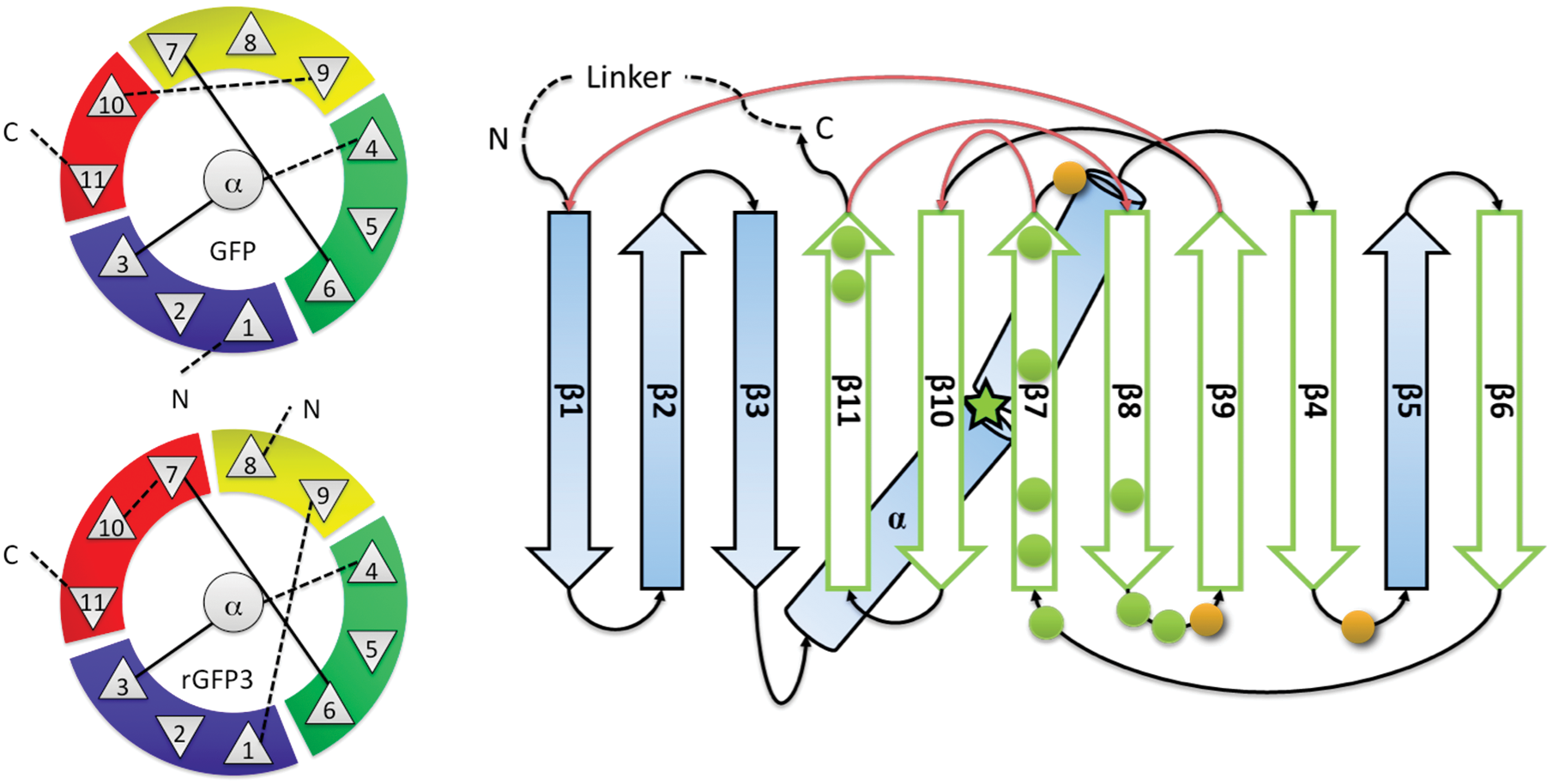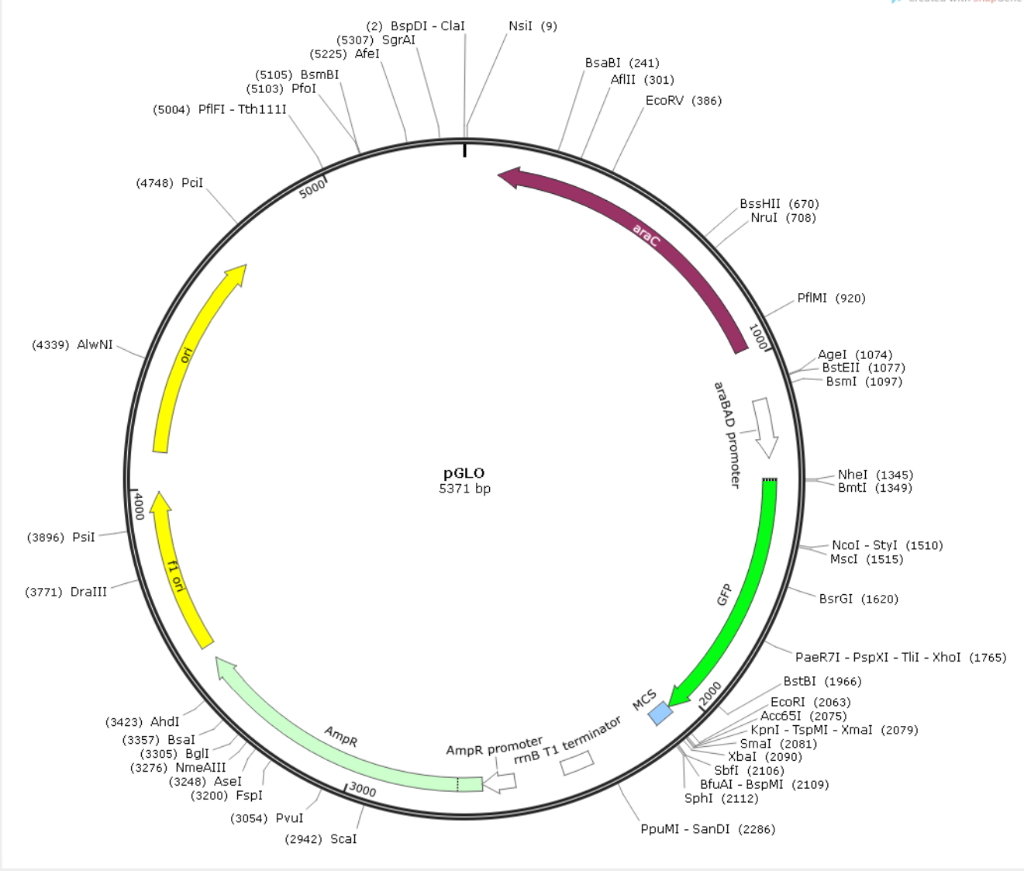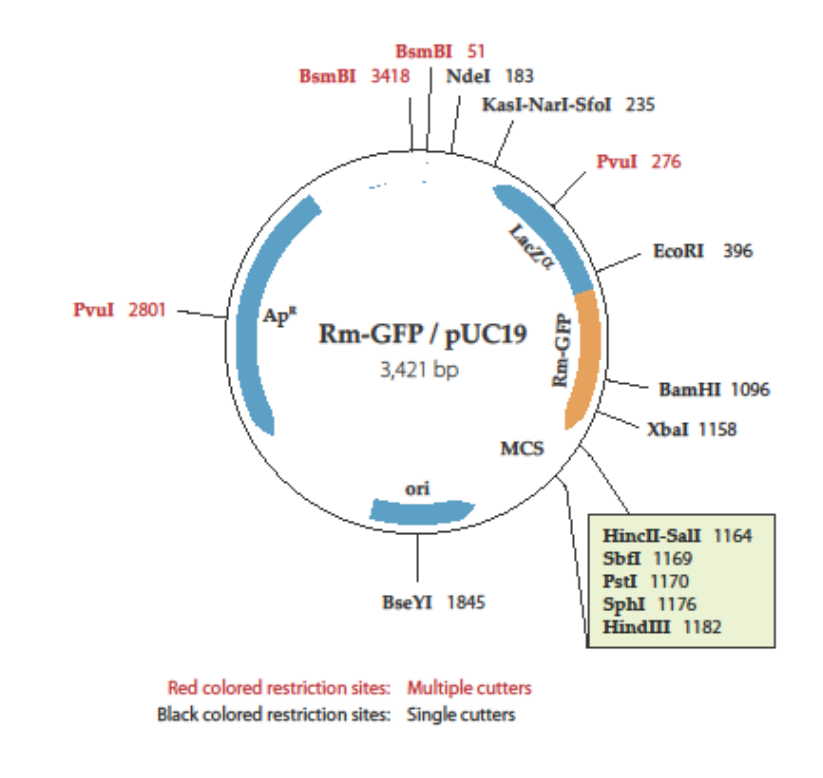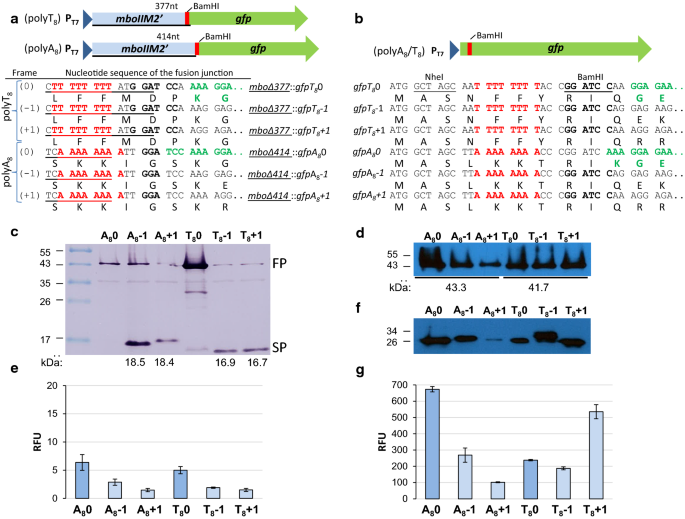
Evaluation of GFP reporter utility for analysis of transcriptional slippage during gene expression | Microbial Cell Factories | Full Text

Removal of a cryptic intron and subcellular localization of green fluorescent protein are required to mark transgenic Arabidopsis plants brightly | PNAS

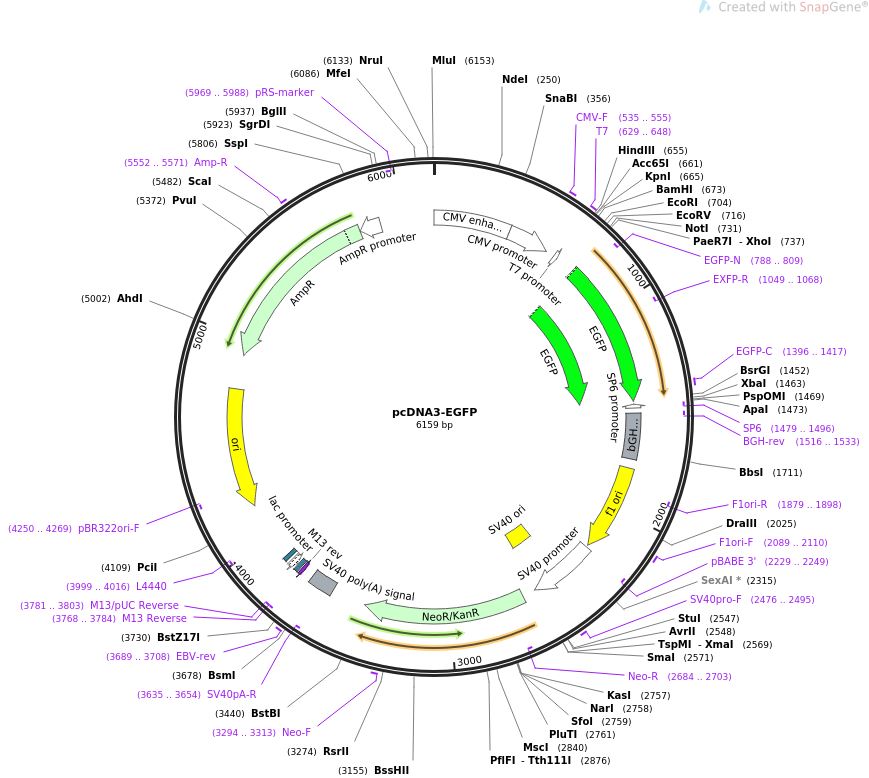
Construction of a plasmid coding for green fluorescent protein tagged cathepsin L and data on expression in colorectal carcinoma cells - ScienceDirect

Figure 10 from Comparative studies on the structure and stability of fluorescent proteins EGFP, zFP506, mRFP1, "dimer2", and DsRed1. | Semantic Scholar

Sequence alignment of asCP, GFP, and DsRed proteins. The numbering is... | Download Scientific Diagram

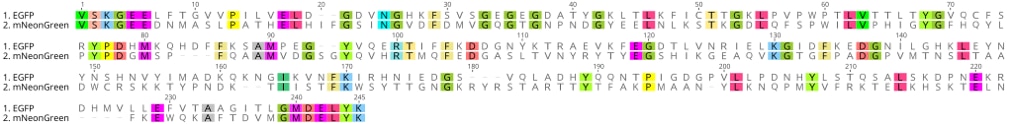
Alignment of amino acid sequence of BFP and GFP. The two proteins are... | Download Scientific Diagram