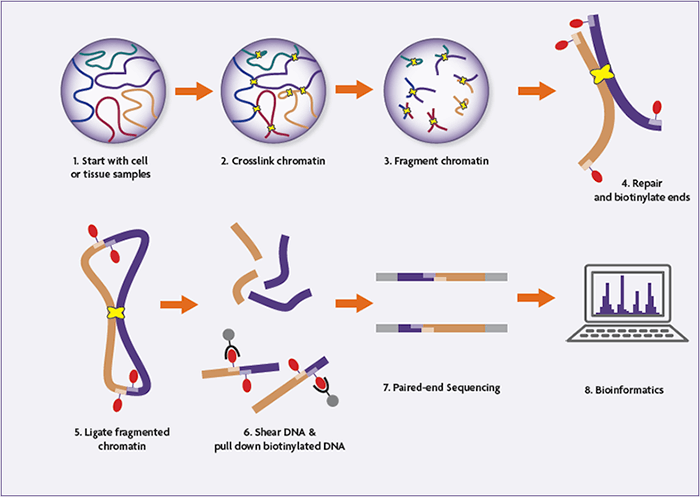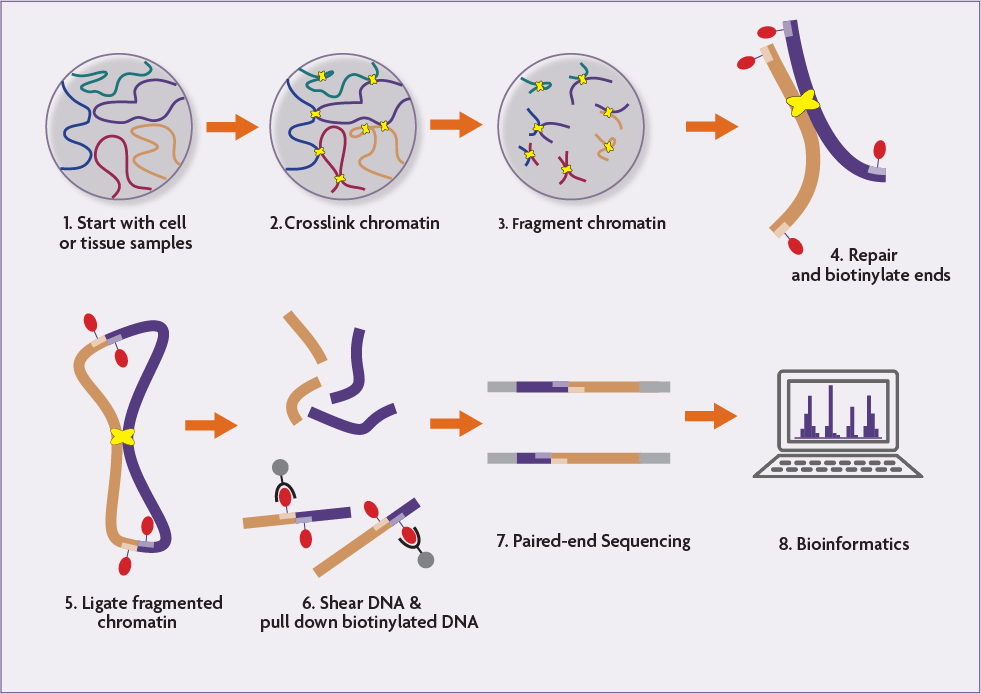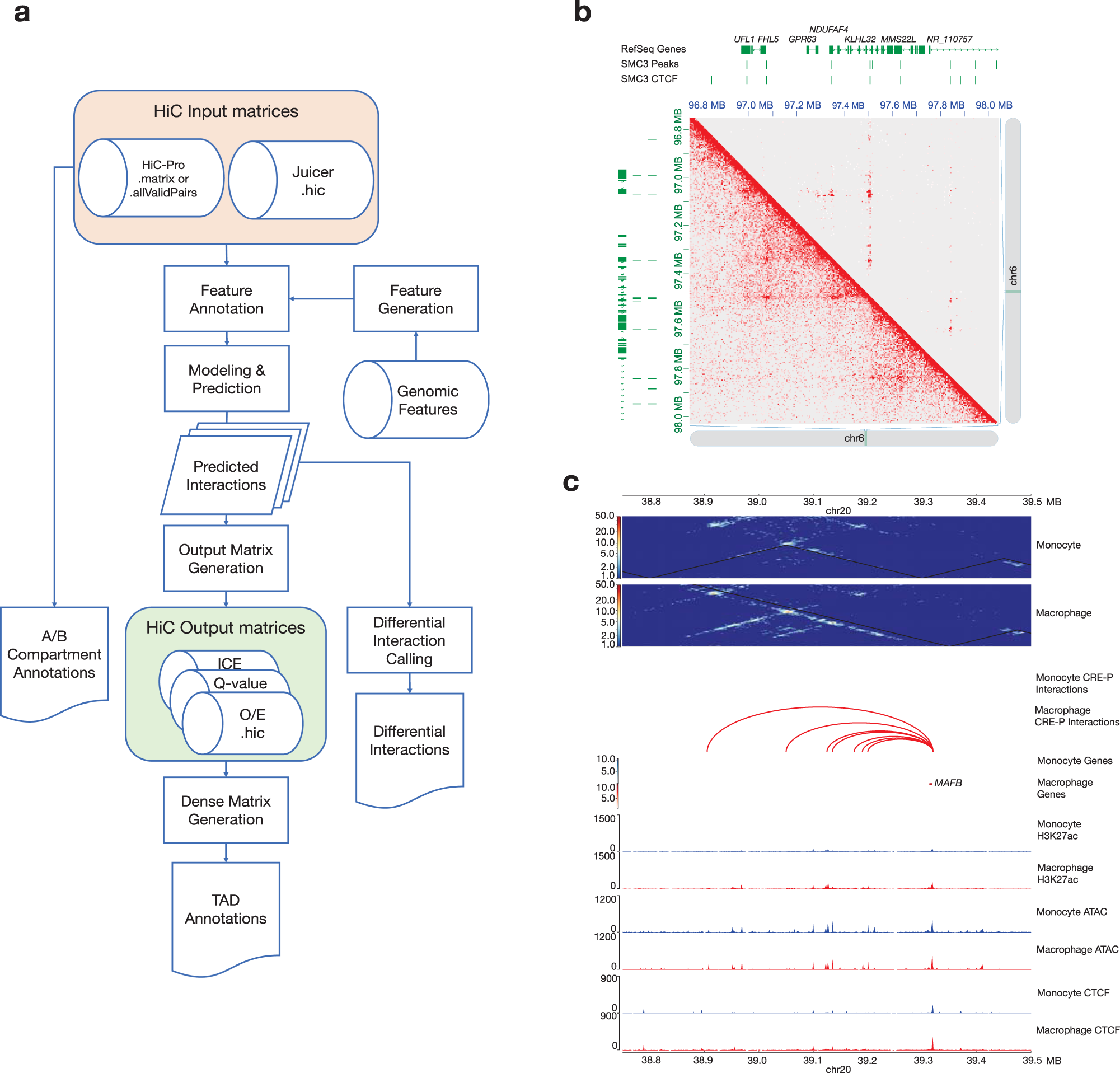
HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP | Nature Communications
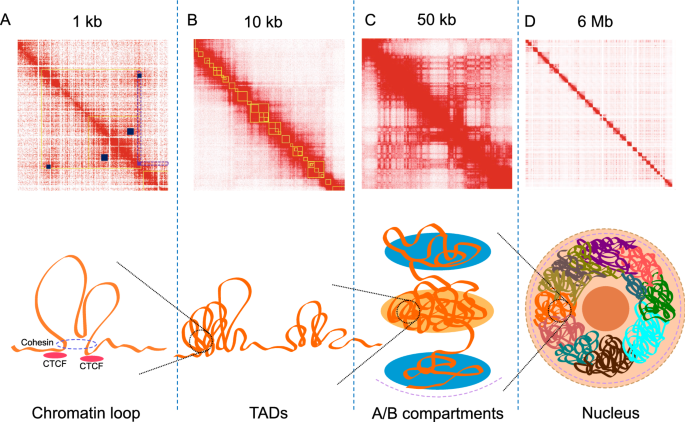
Seeing the forest through the trees: prioritising potentially functional interactions from Hi-C | Epigenetics & Chromatin | Full Text

Liquid chromatin Hi-C characterizes compartment-dependent chromatin interaction dynamics | Nature Genetics

Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C | Nature Genetics

Hi‐C 3.0: Improved Protocol for Genome‐Wide Chromosome Conformation Capture - Lafontaine - 2021 - Current Protocols - Wiley Online Library

Hi-C 2.0: An Optimized Hi-C Procedure for High-Resolution Genome-Wide Mapping of Chromosome Conformation | bioRxiv

Comparative Hi-C Reveals that CTCF Underlies Evolution of Chromosomal Domain Architecture: Cell Reports
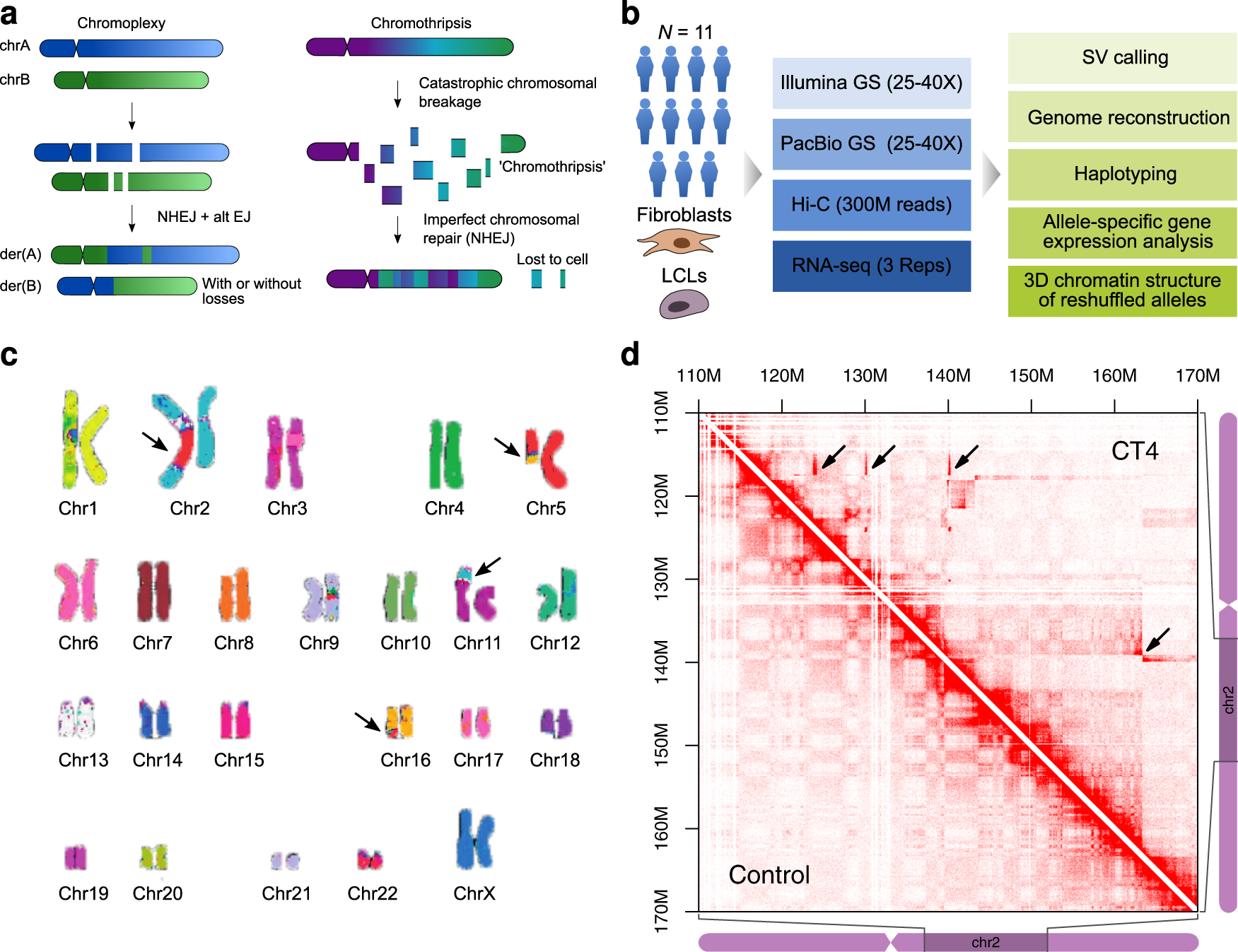
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes | Nature Communications

BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions | Nature Communications