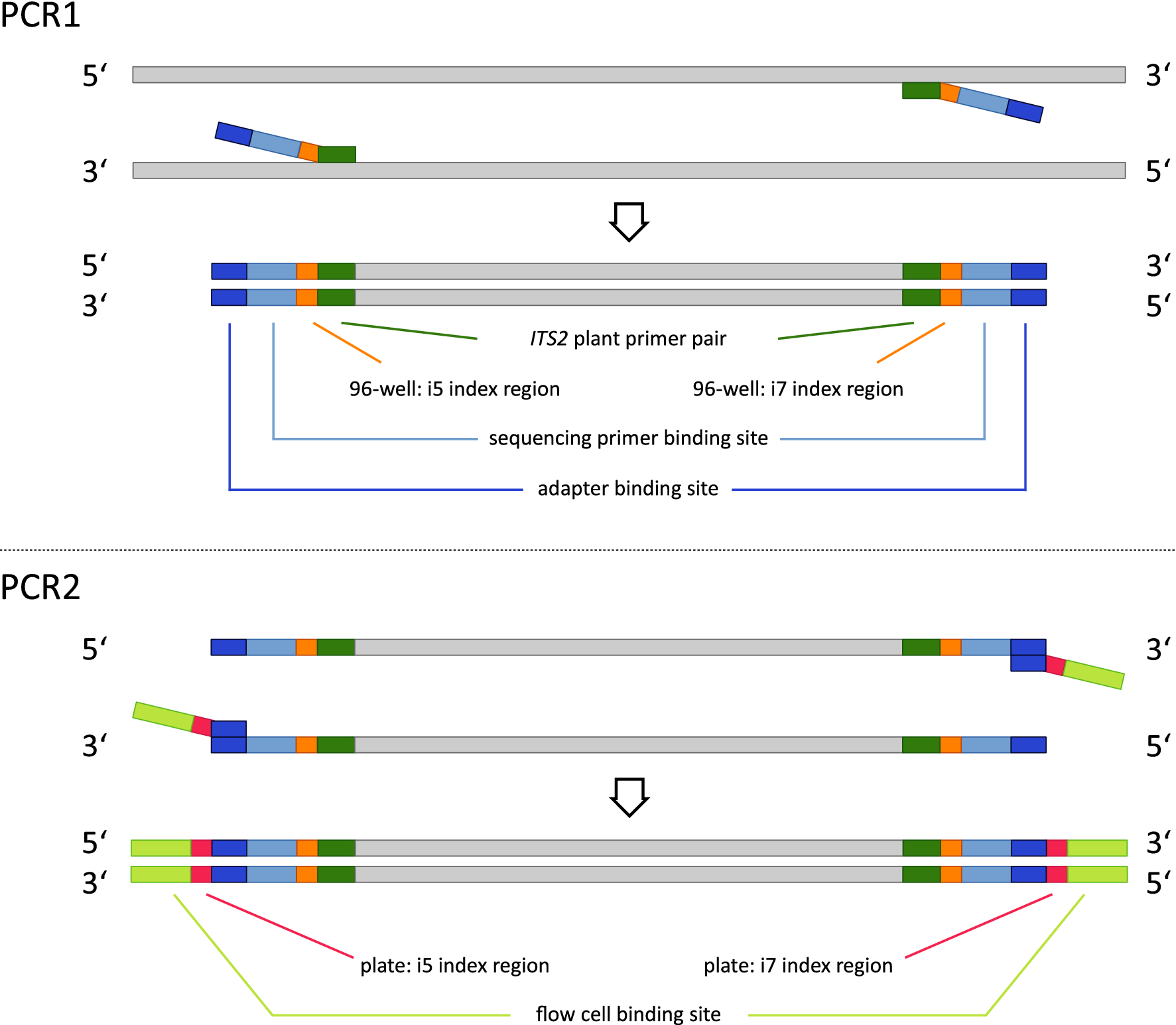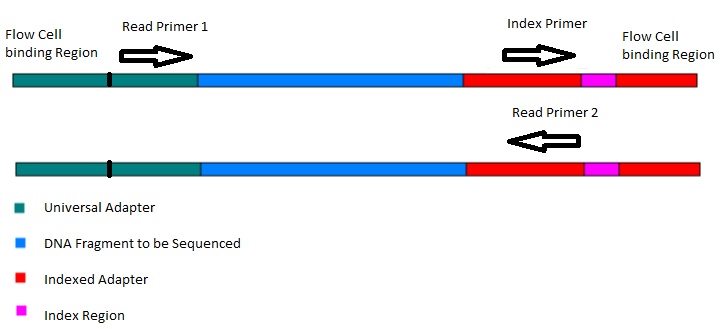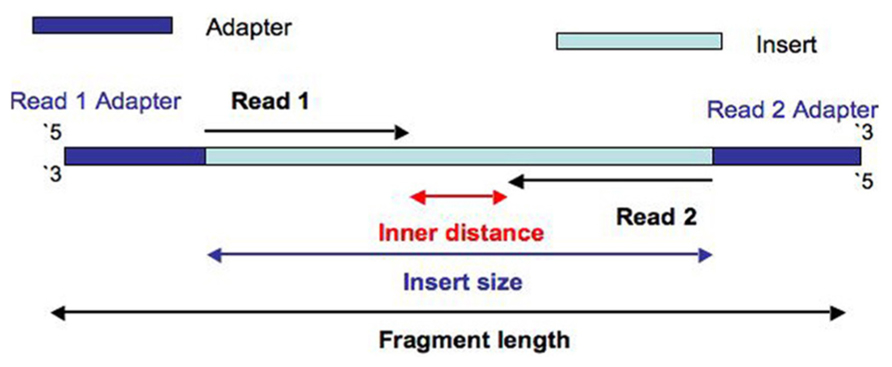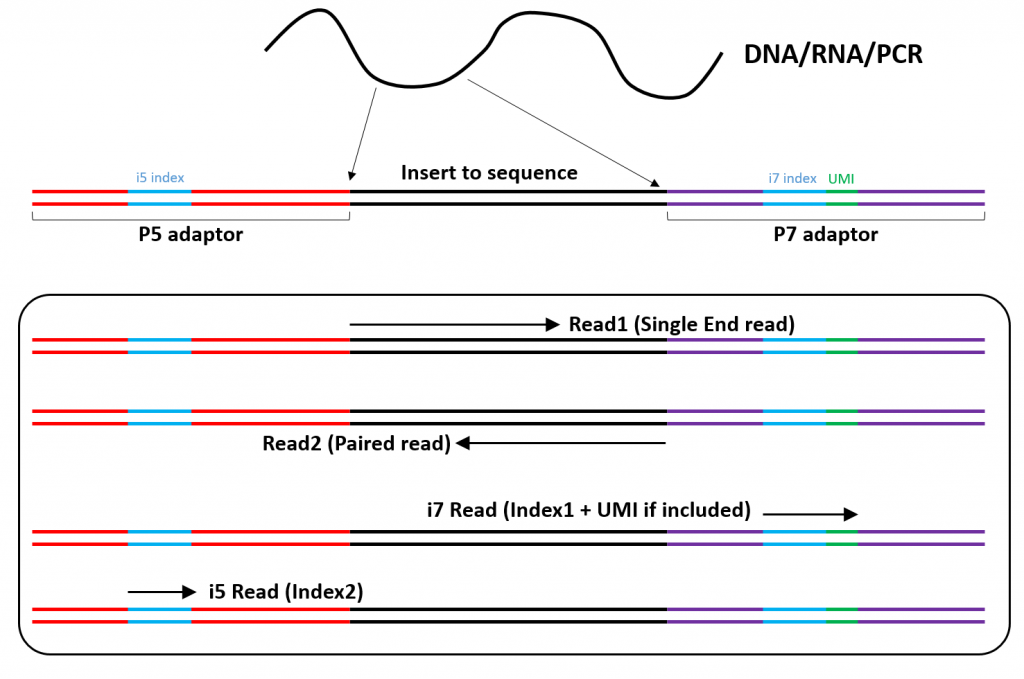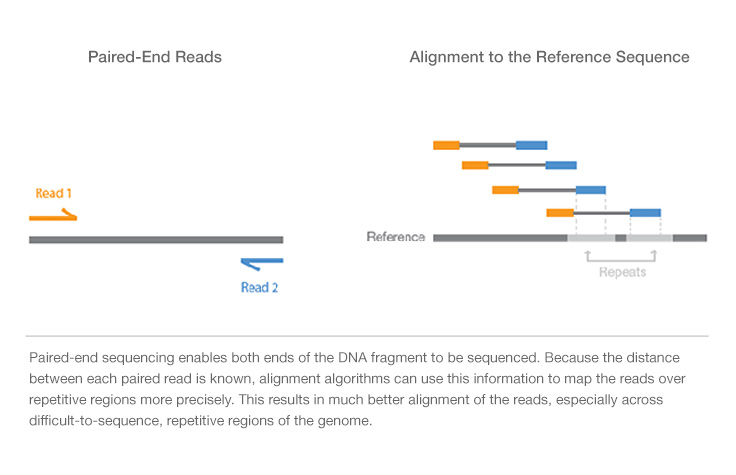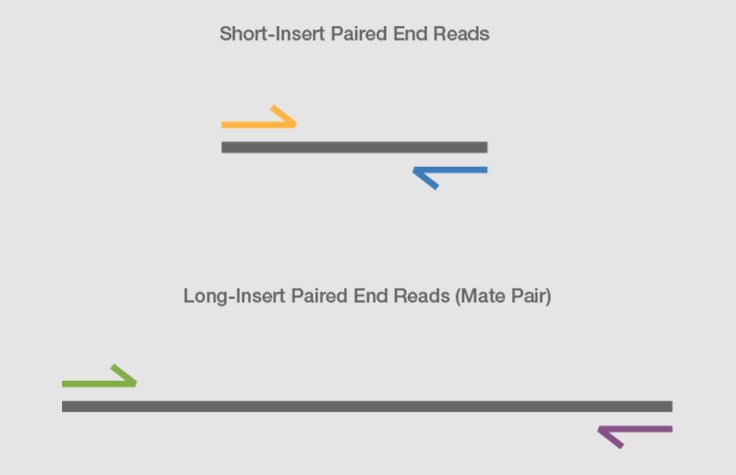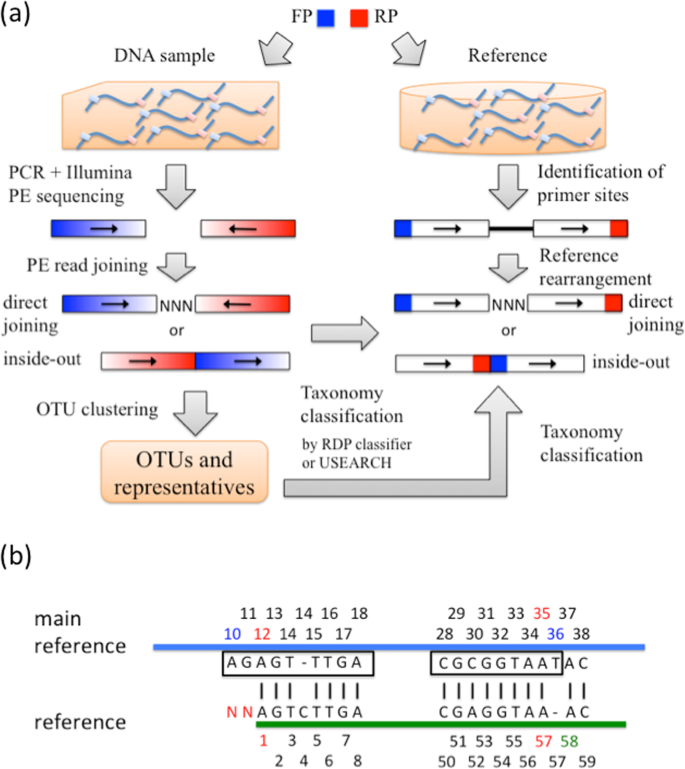
Joining Illumina paired-end reads for classifying phylogenetic marker sequences | BMC Bioinformatics | Full Text
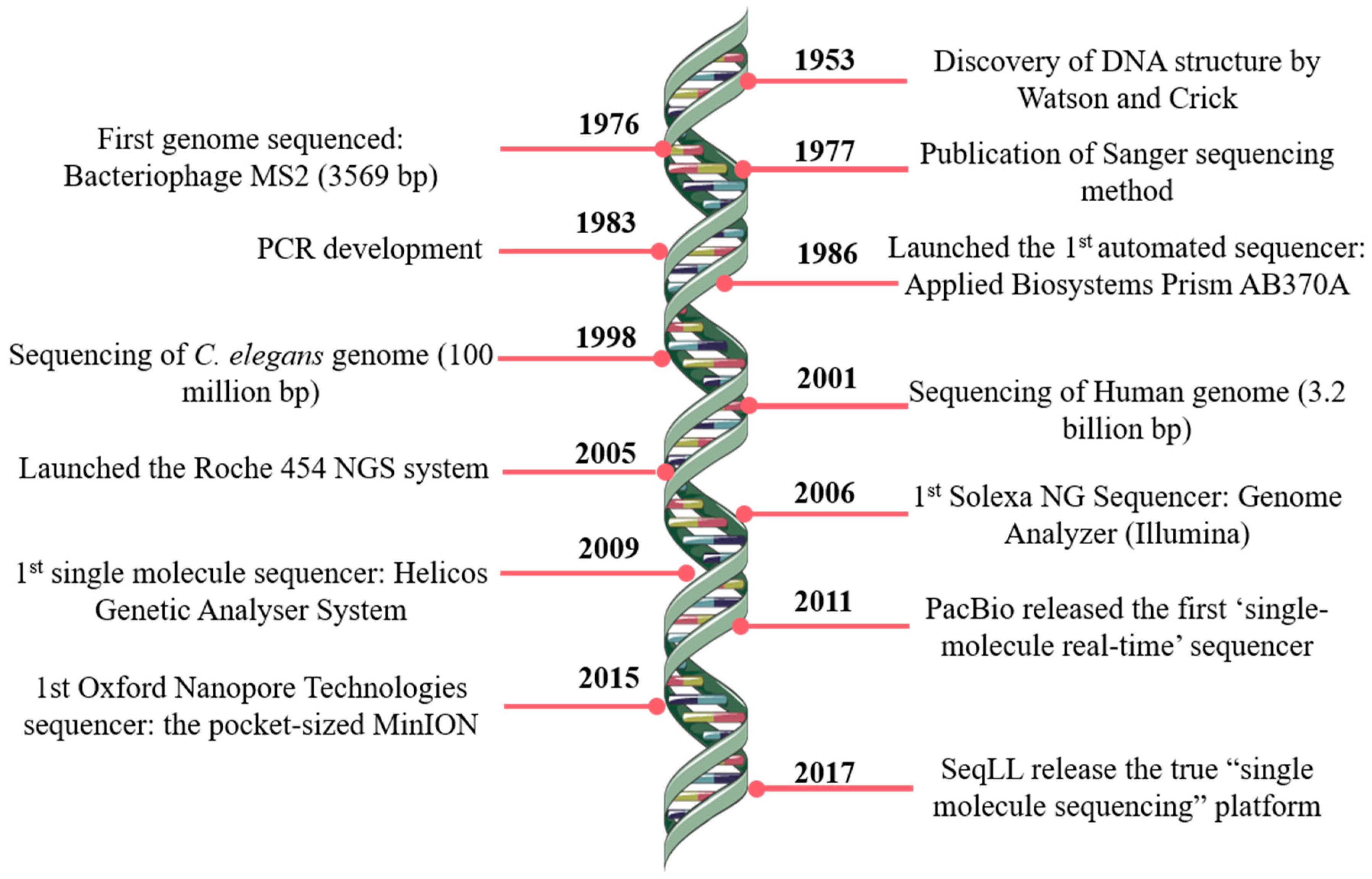
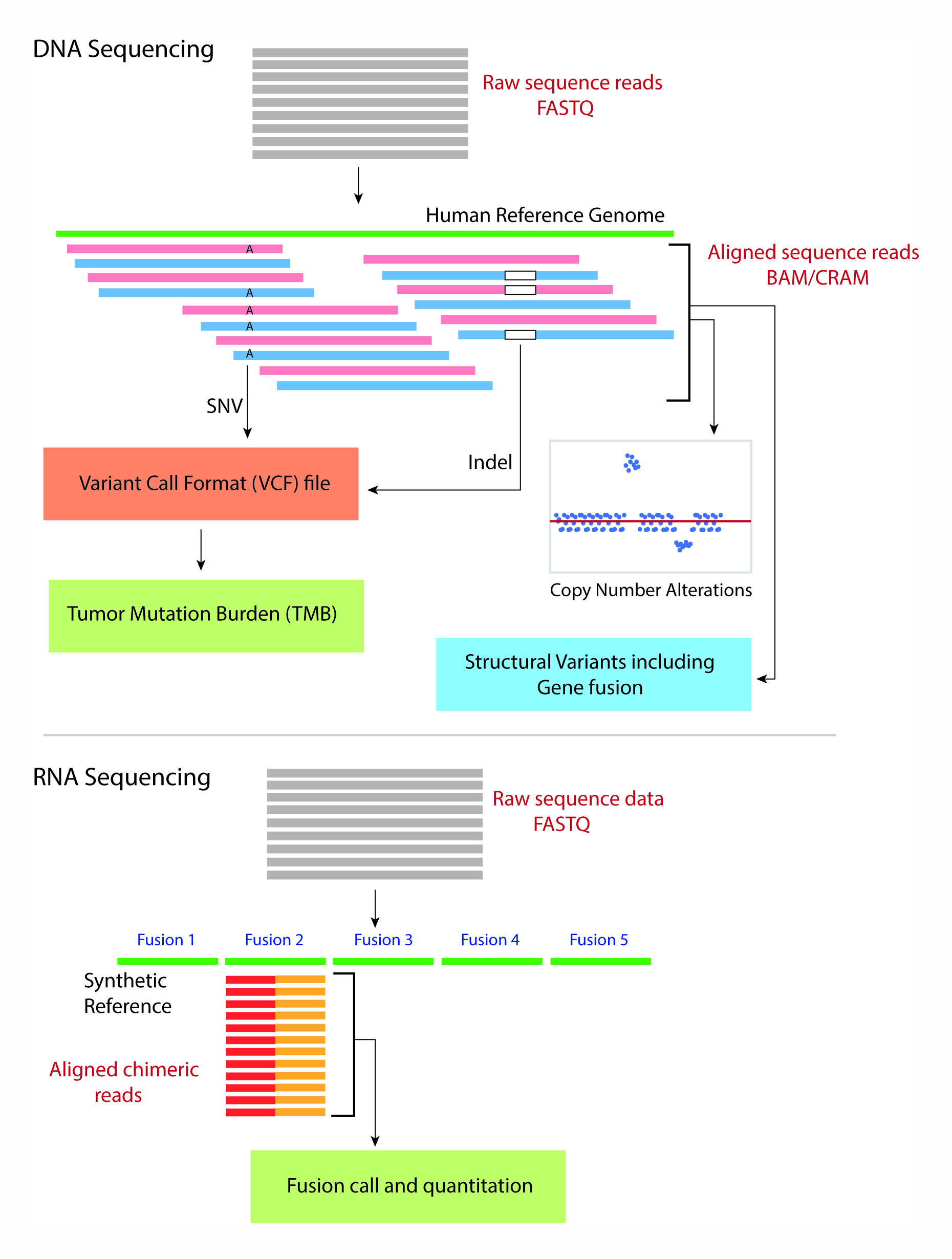
JCM | Free Full-Text | Bioinformatics and Computational Tools for Next-Generation Sequencing Analysis in Clinical Genetics
Major short-read and long-read sequencing technologies. (A) Illumina... | Download Scientific Diagram

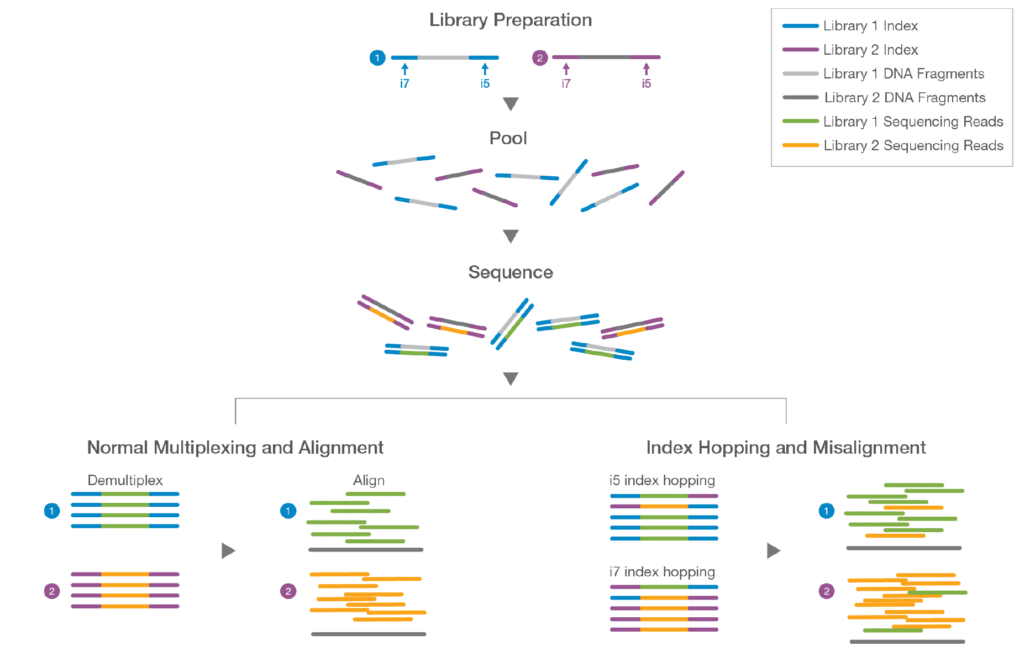
Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge
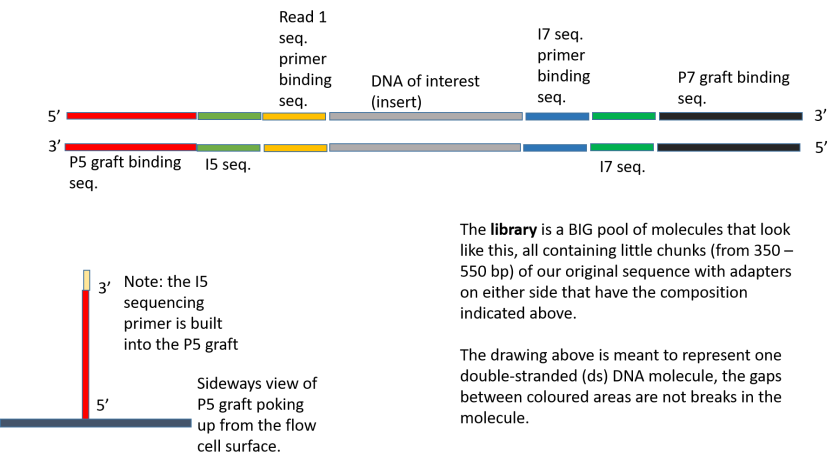
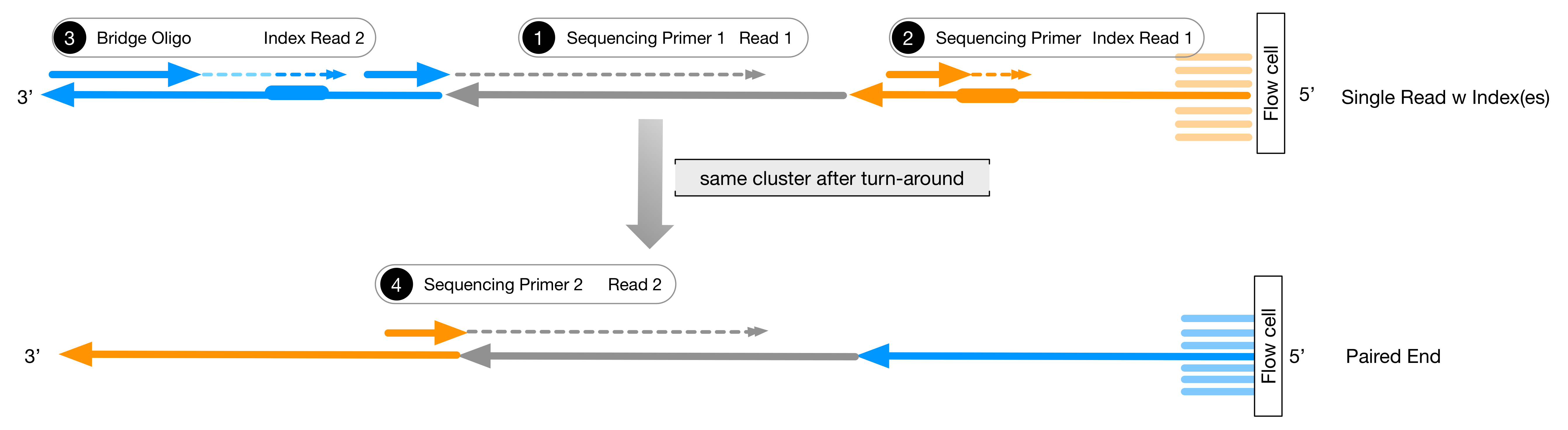
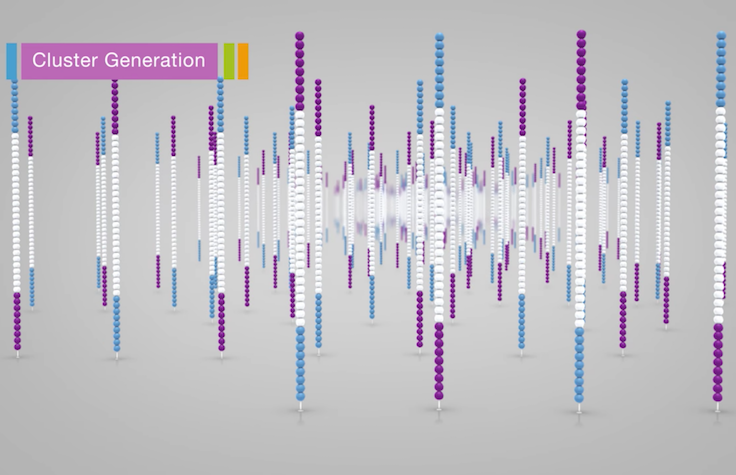
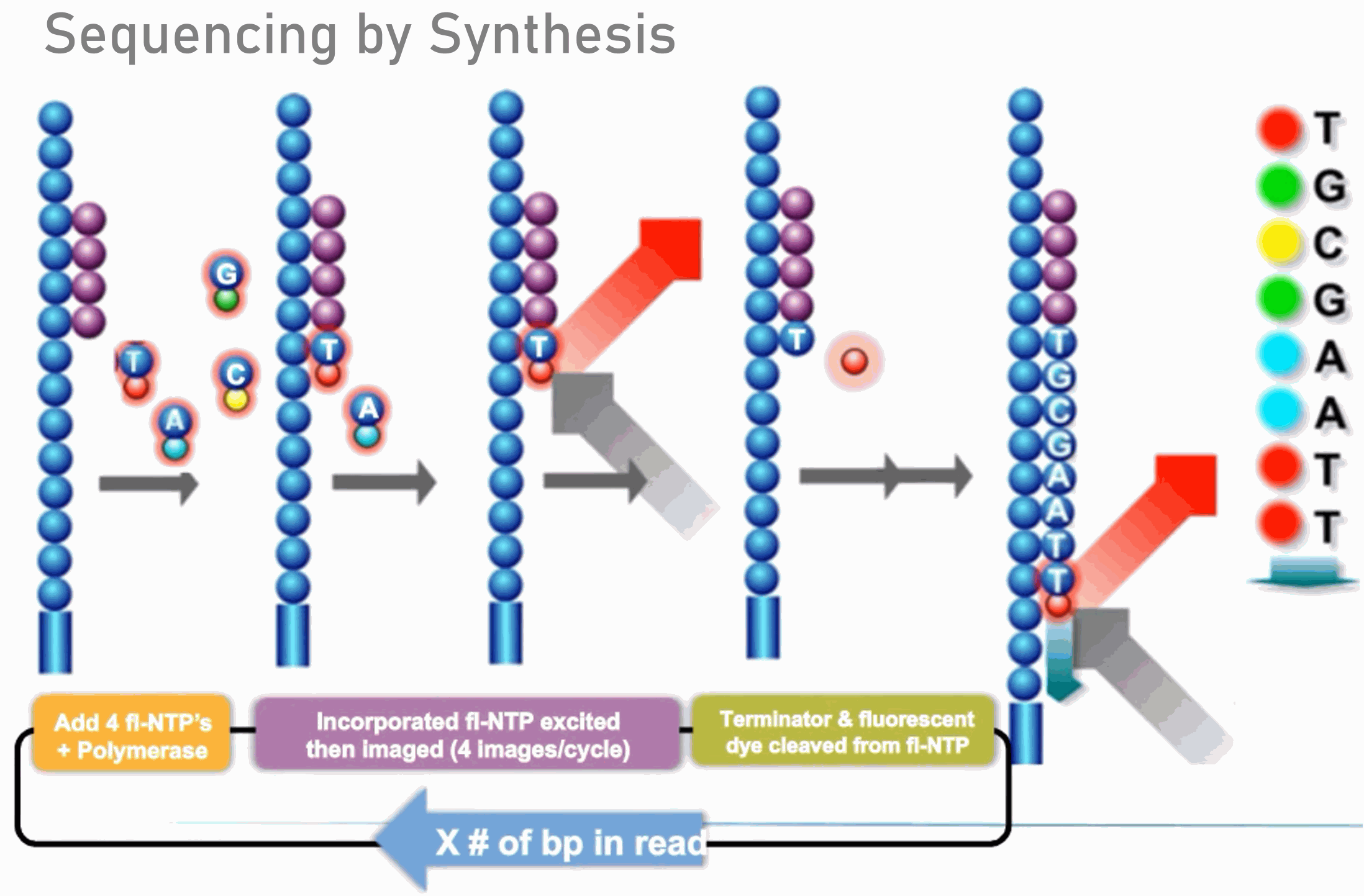
Illumina Sequencing (for Dummies) -An overview on how our samples are sequenced. – kscbioinformatics

The NaS workflow. Inputs are the Illumina short reads and the MinION®... | Download Scientific Diagram
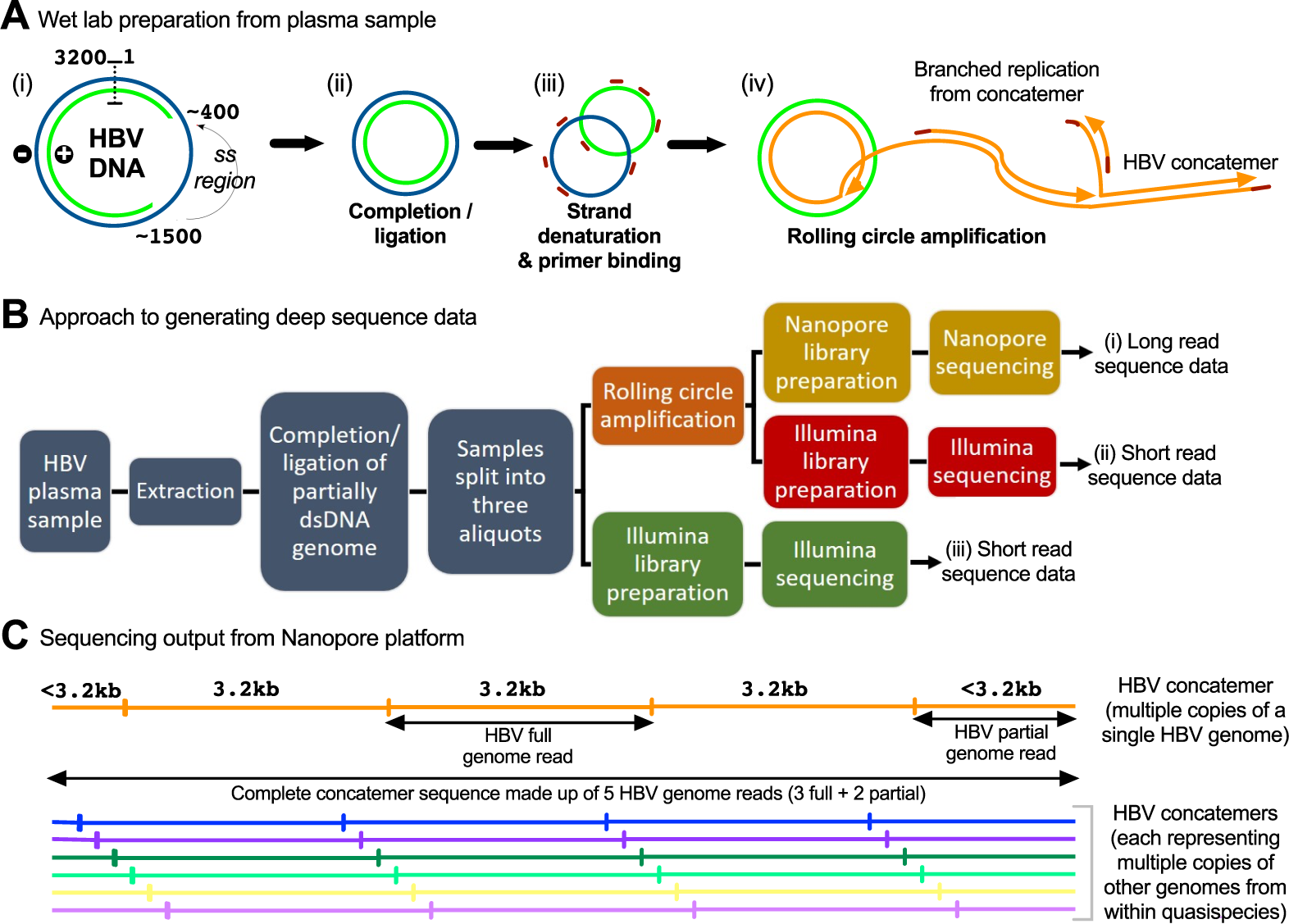
Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV) | Scientific Reports