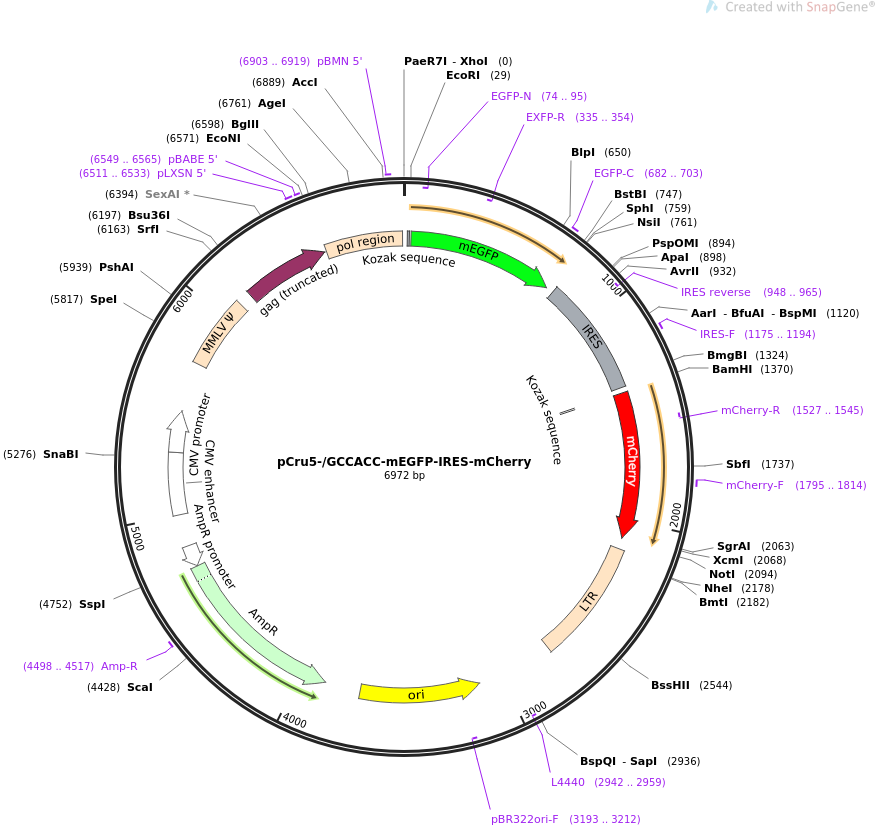
Kozak sequence improved the expression of 2A downstream gene by its... | Download Scientific Diagram

Fine-tuning the expression of pathway gene in yeast using a regulatory library formed by fusing a synthetic minimal promoter with different Kozak variants | Microbial Cell Factories | Full Text

IRES-Dependent Second Gene Expression Is Significantly Lower Than Cap-Dependent First Gene Expression in a Bicistronic Vector: Molecular Therapy

Nucleotides upstream of the Kozak sequence strongly influence gene expression in the yeast S. cerevisiae | Journal of Biological Engineering | Full Text
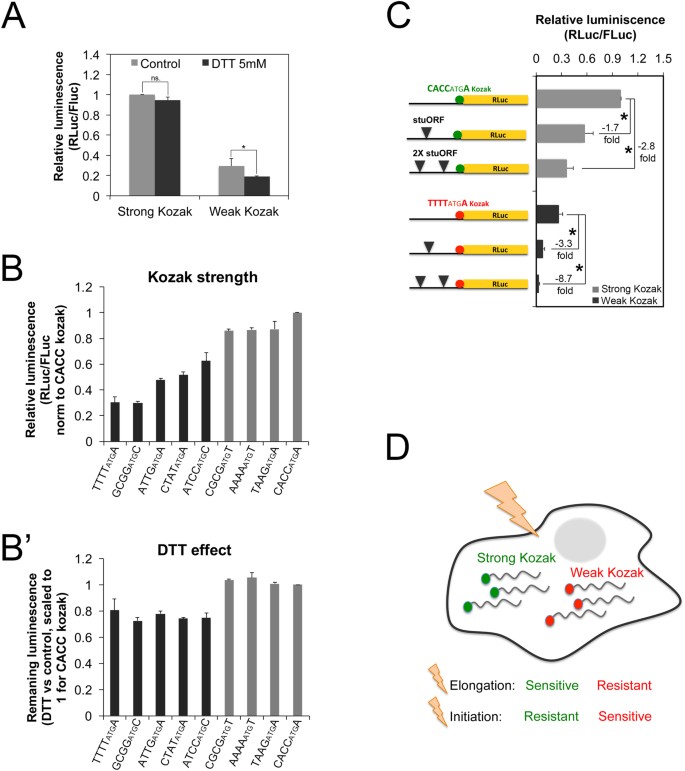
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports

Tailoring translational strength using Kozak sequence variants improves bispecific antibody assembly and reduces product‐related impurities in CHO cells - Blanco - 2020 - Biotechnology and Bioengineering - Wiley Online Library

Enhancement of protein expression in insect cells by a lobster tropomyosin cDNA leader sequence - ScienceDirect

Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv
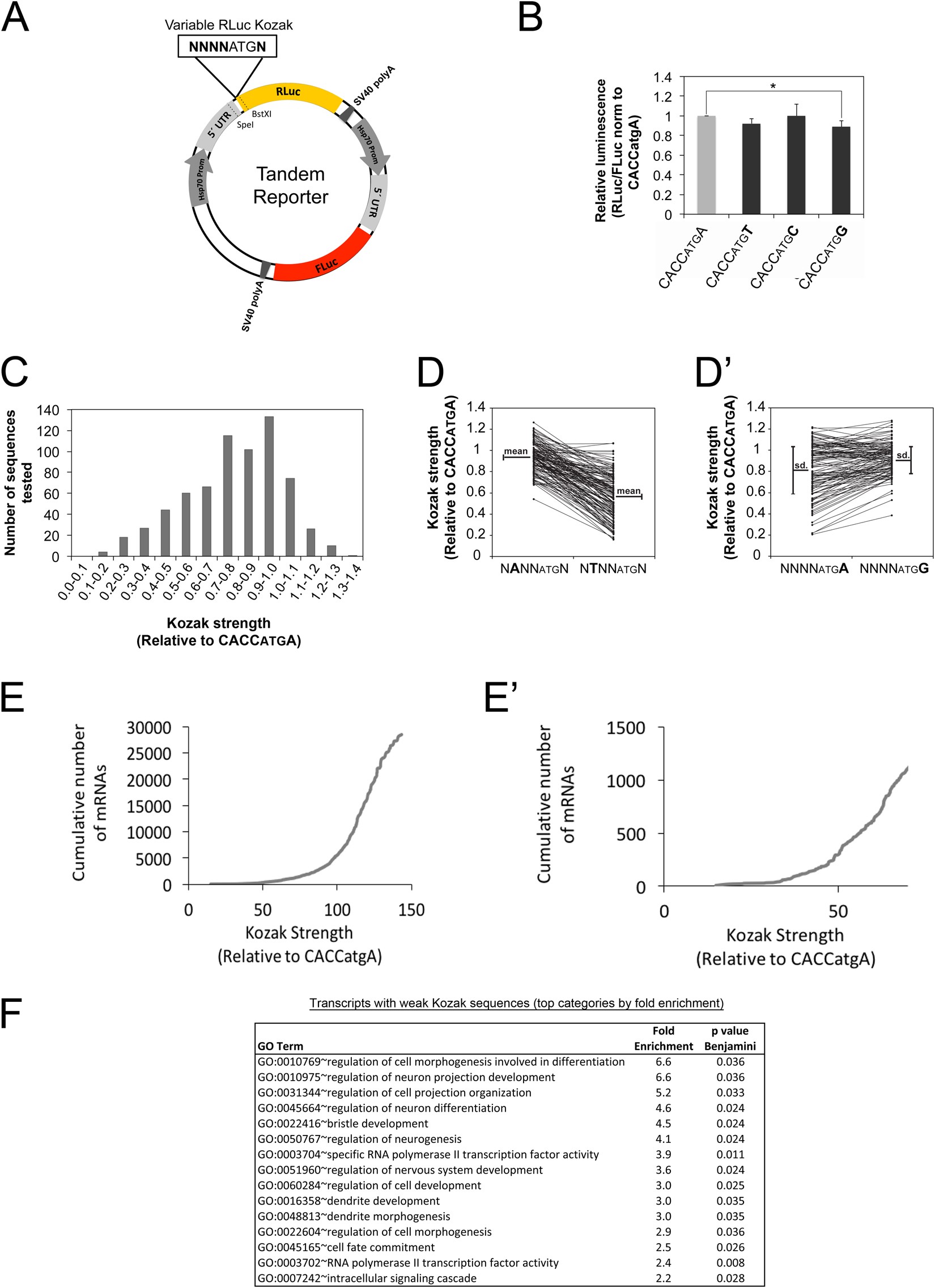
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports