
Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology

Well-characterized sequence features of eukaryote genomes and implications for ab initio gene prediction - ScienceDirect
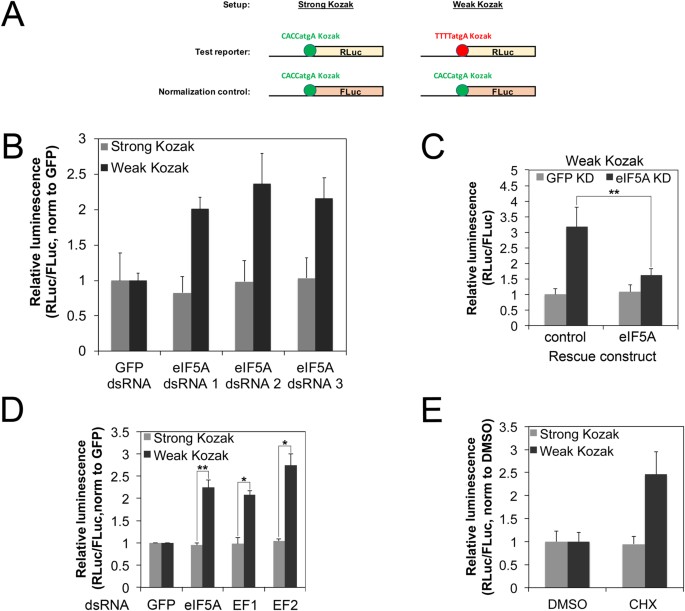
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports
The +4G Site in Kozak Consensus Is Not Related to the Efficiency of Translation Initiation | PLOS ONE

Rps26 directs mRNA-specific translation by recognition of Kozak sequence elements | Nature Structural & Molecular Biology

Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology
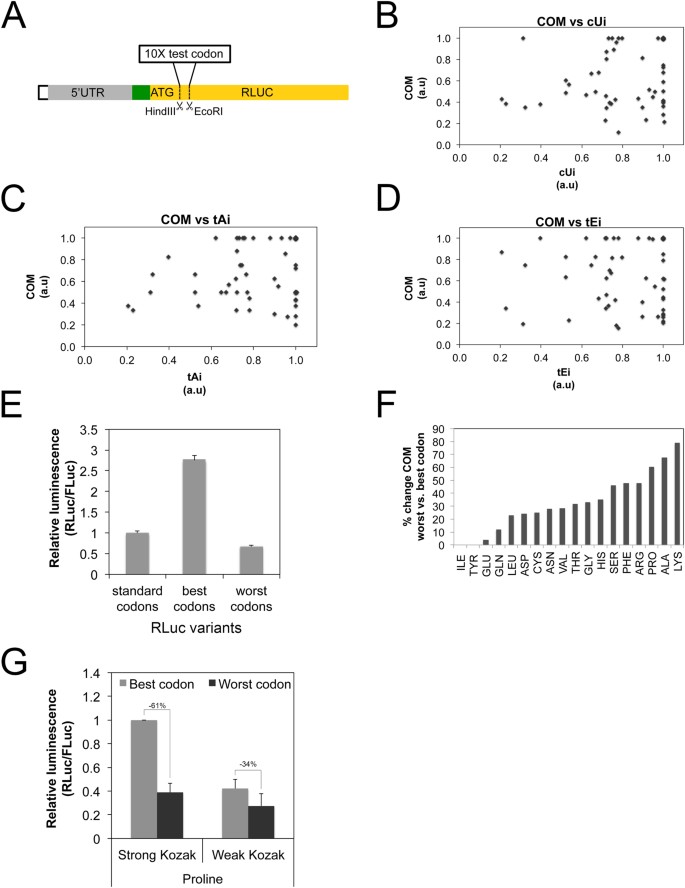
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports
Machine learning predicts translation initiation sites in neurologic diseases with nucleotide repeat expansions | PLOS ONE

Quantification and discovery of sequence determinants of protein‐per‐mRNA amount in 29 human tissues | Molecular Systems Biology

Machine learning predicts translation initiation sites in neurologic diseases with expanded repeats | bioRxiv
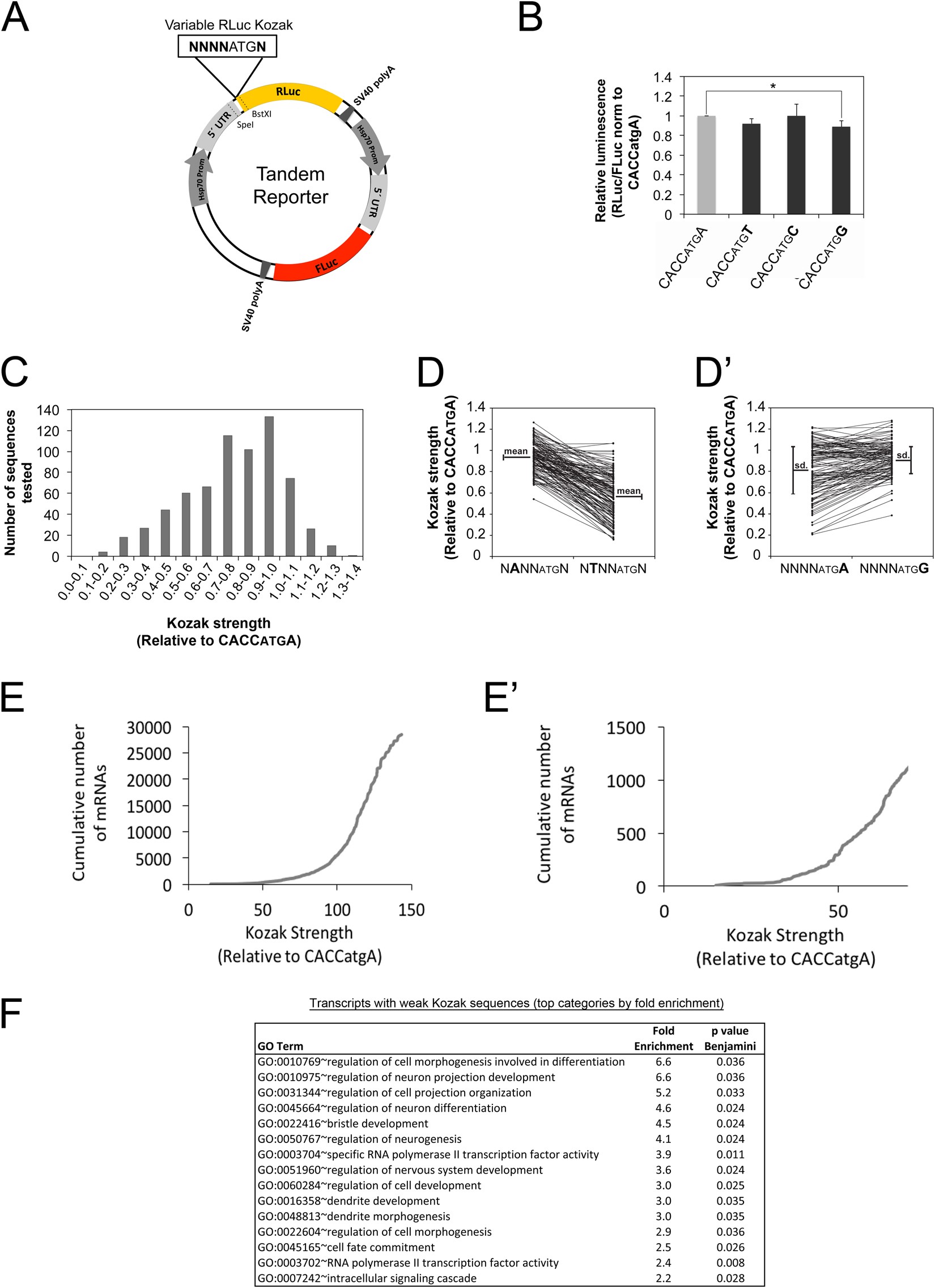
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports

The effect of changing the +4 position of the Kozak sequence on the... | Download Scientific Diagram

Plants | Free Full-Text | Modulation of the Translation Efficiency of Heterologous mRNA and Target Protein Stability in a Plant System: The Case Study of Interferon-αA

Relationship between consensus Kozak sequence at the true start ATG for... | Download Scientific Diagram

DeepTIS: Improved translation initiation site prediction in genomic sequence via a two-stage deep learning model - ScienceDirect

Kozak consensus sequence surrounding start methionin in the 440 cCDS... | Download Scientific Diagram

Analysis of Kozak consensus sequences surrounding the start codon AUG... | Download Scientific Diagram





