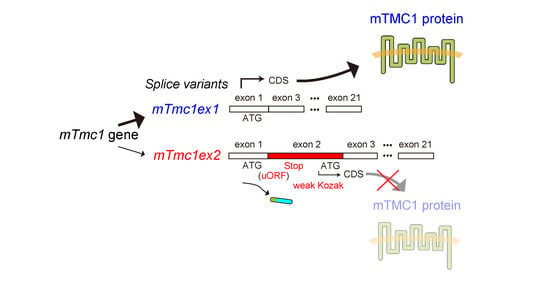
Identification of Alternatively Translated Tetherin Isoforms with Differing Antiviral and Signaling Activities | PLOS Pathogens
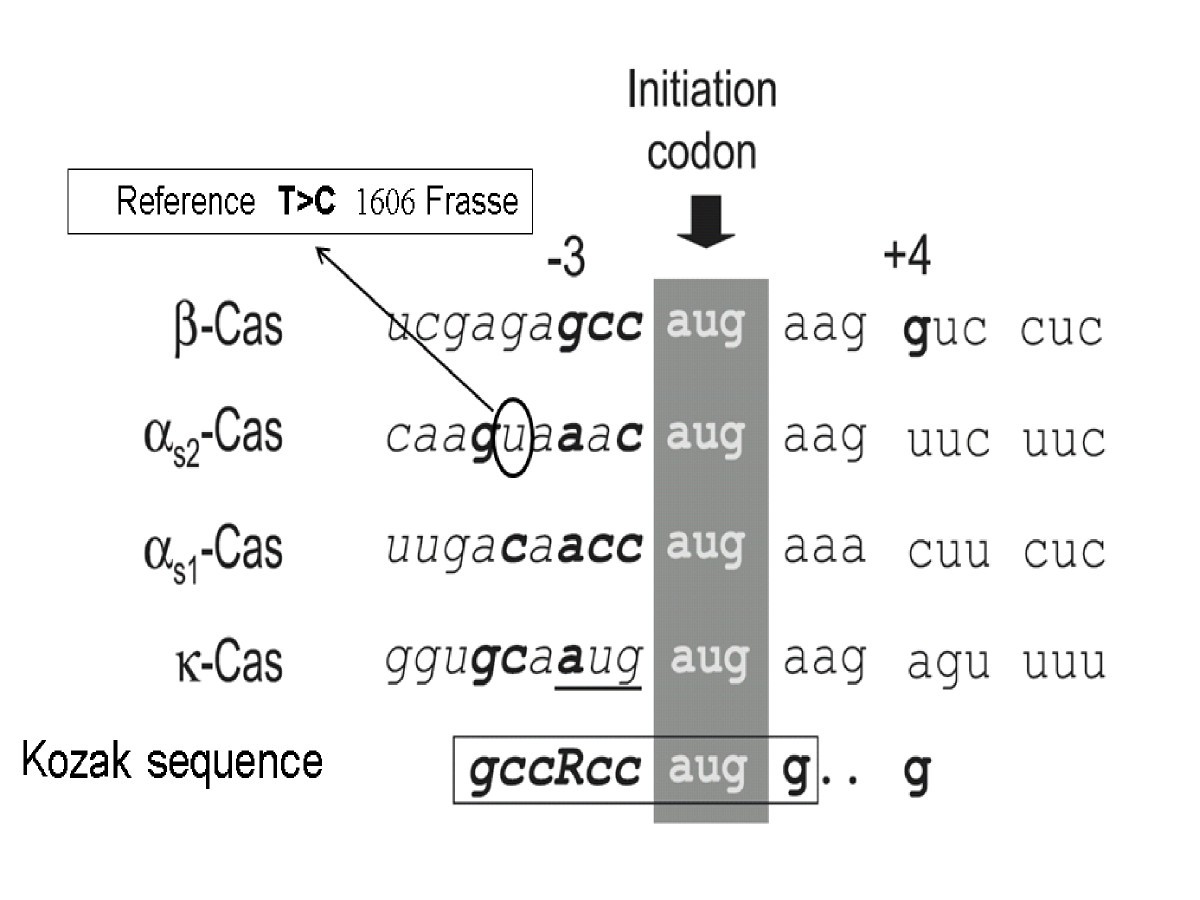
Molecular characterization of a long range haplotype affecting protein yield and mastitis susceptibility in Norwegian Red cattle | BMC Genomic Data | Full Text

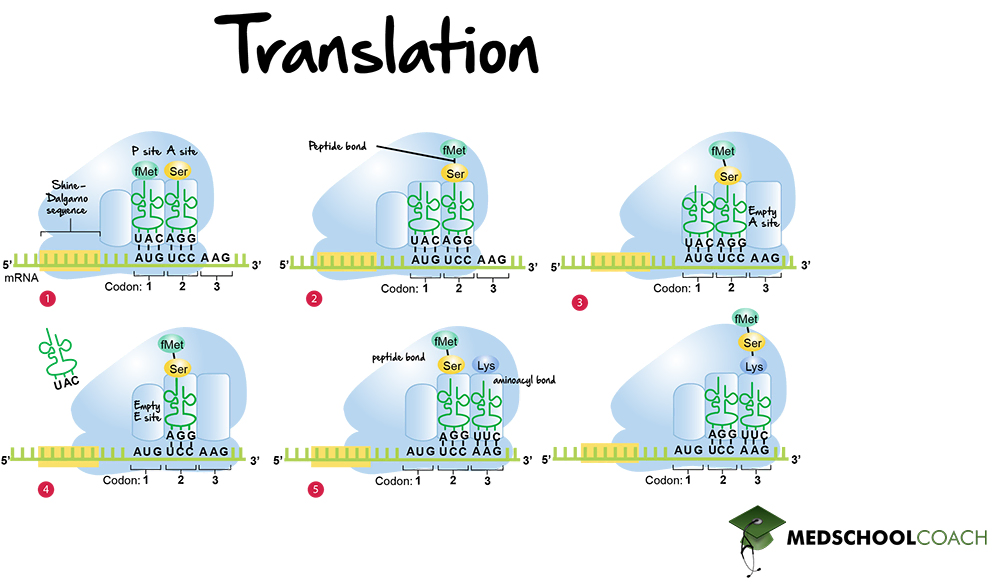
BIOL308 Lecture Notes - Fall 2017, Lecture 10 - Kozak Consensus Sequence, Ribosome-Binding Site, Start Codon

Relationship between consensus Kozak sequence at the true start ATG for... | Download Scientific Diagram

Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv
Single baseâ•'pair substitutions at the translation initiation sites of human genes as a cause of inherited disease<link

Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology
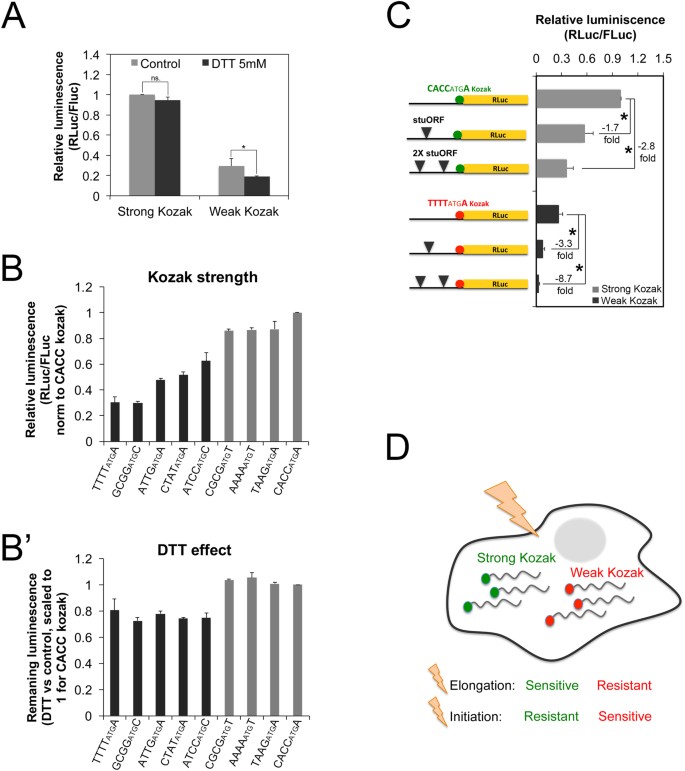
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports
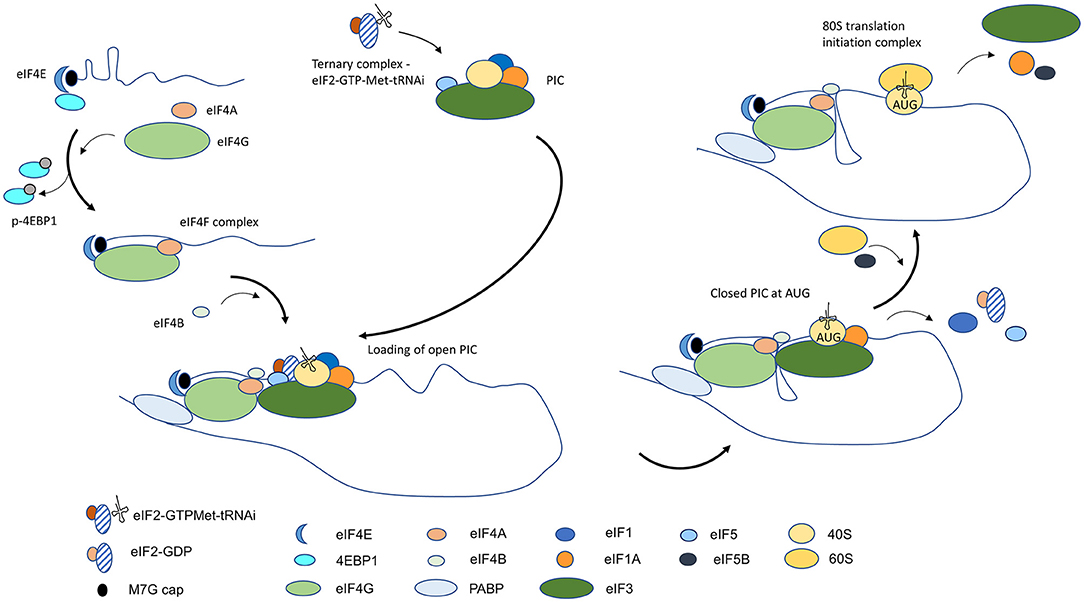
Frontiers | Disorder at the Start: The Contribution of Dysregulated Translation Initiation to Cancer Therapy Resistance

Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology

Fine-tuning the expression of pathway gene in yeast using a regulatory library formed by fusing a synthetic minimal promoter with different Kozak variants | Microbial Cell Factories | Full Text

What is the Difference Between Shine Dalgarno and Kozak Sequence | Compare the Difference Between Similar Terms
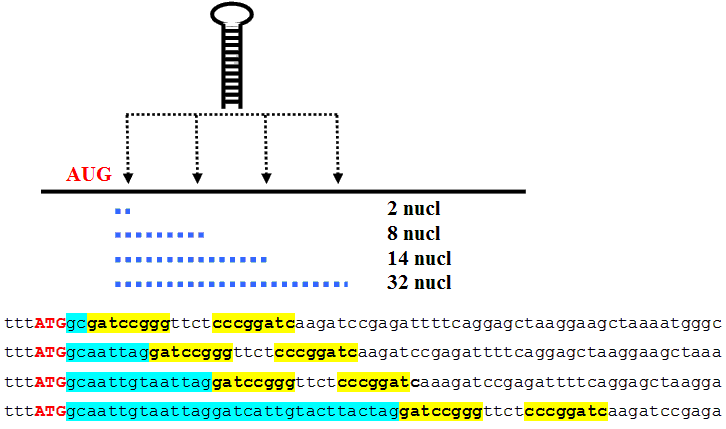
AUG_hairpin: program for prediction of a downstream hairpin potentially increasing initiation of translation at start AUG codon in a suboptimal context.
An Upstream Open Reading Frame Modulates Ebola Virus Polymerase Translation and Virus Replication | PLOS Pathogens