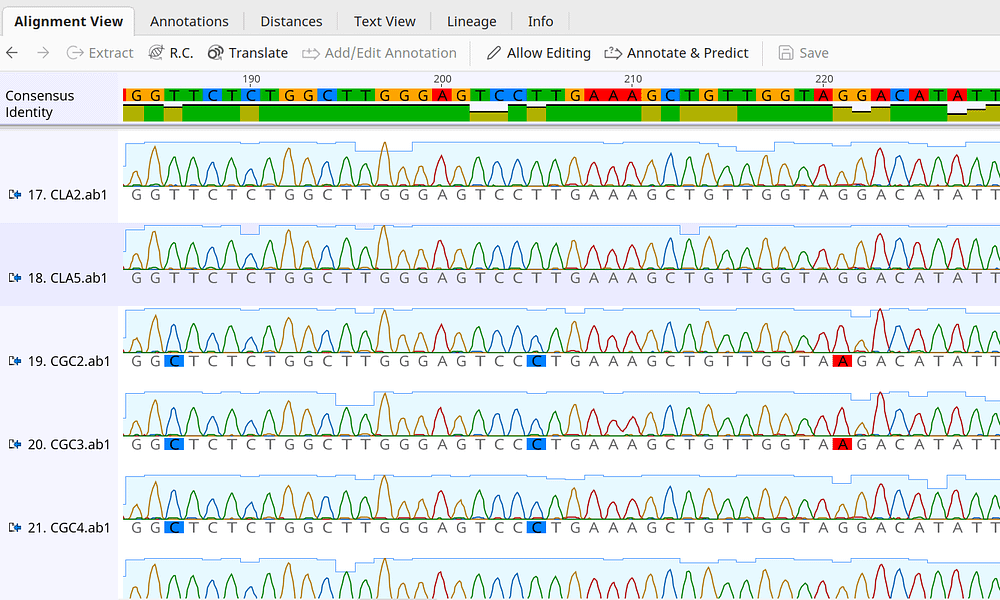
Research Article Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling
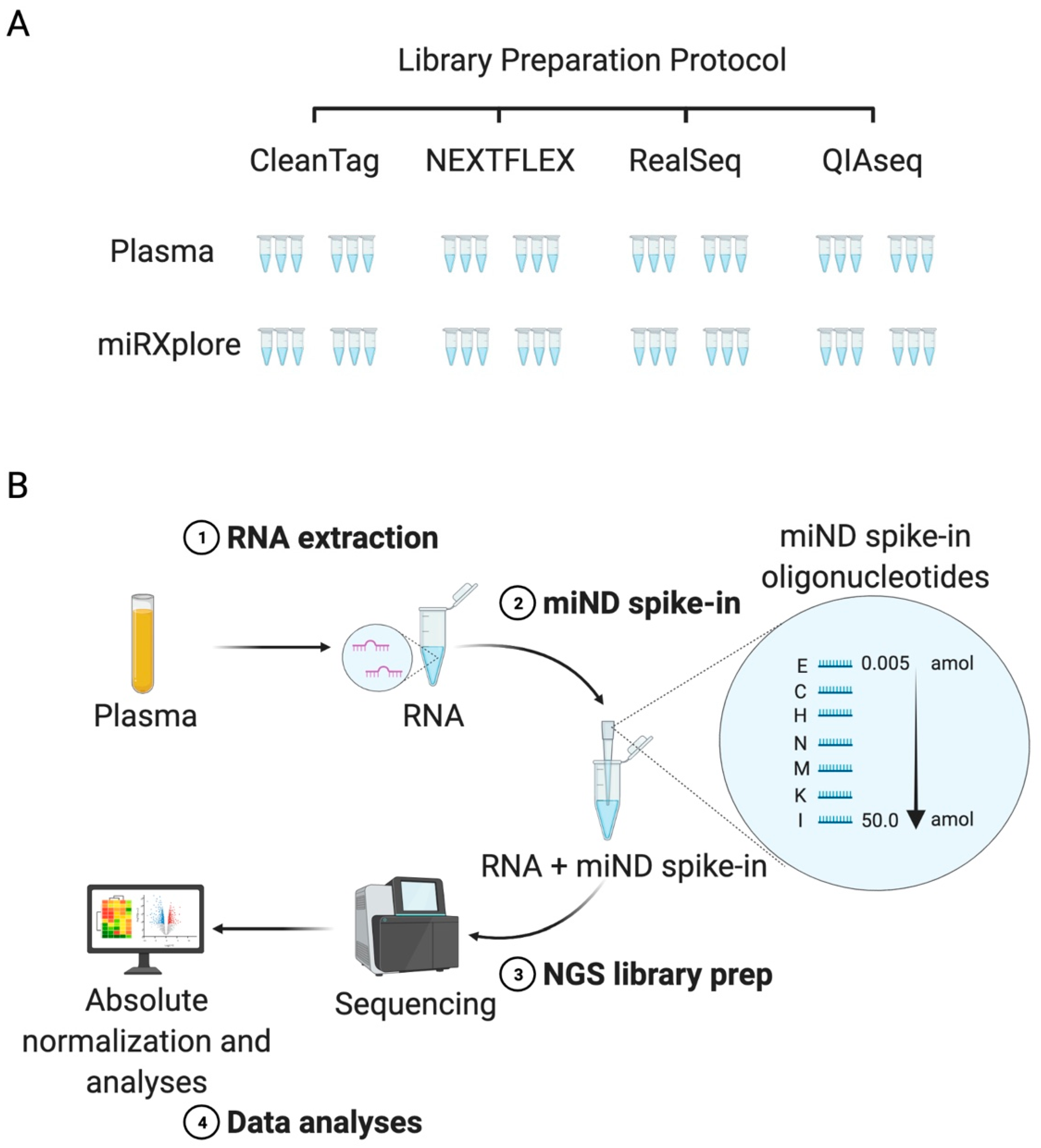
IJMS | Free Full-Text | A MicroRNA Next-Generation-Sequencing Discovery Assay (miND) for Genome-Scale Analysis and Absolute Quantitation of Circulating MicroRNA Biomarkers
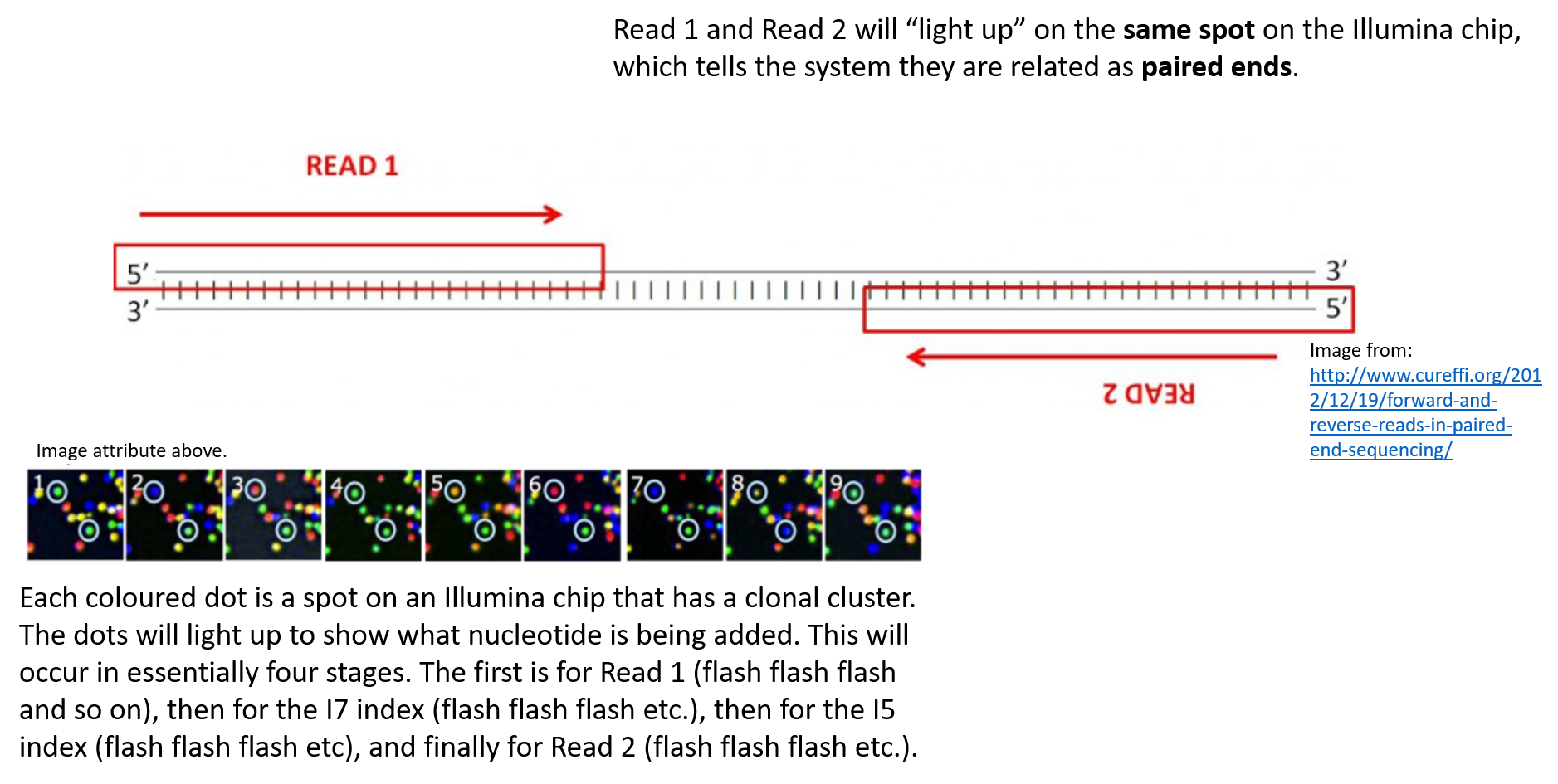
Illumina Sequencing (for Dummies) -An overview on how our samples are sequenced. – kscbioinformatics

Low-Cost, User-Friendly, All-Integrated Smartphone-Based Microplate Reader for Optical-Based Biological and Chemical Analyses | Analytical Chemistry
Decoding of Superimposed Traces Produced by Direct Sequencing of Heterozygous Indels | PLOS Computational Biology
Multi-amplicon microbiome data analysis pipelines for mixed orientation sequences using QIIME2: Assessing reference database, variable region and pre-processing bias in classification of mock bacterial community samples | PLOS ONE