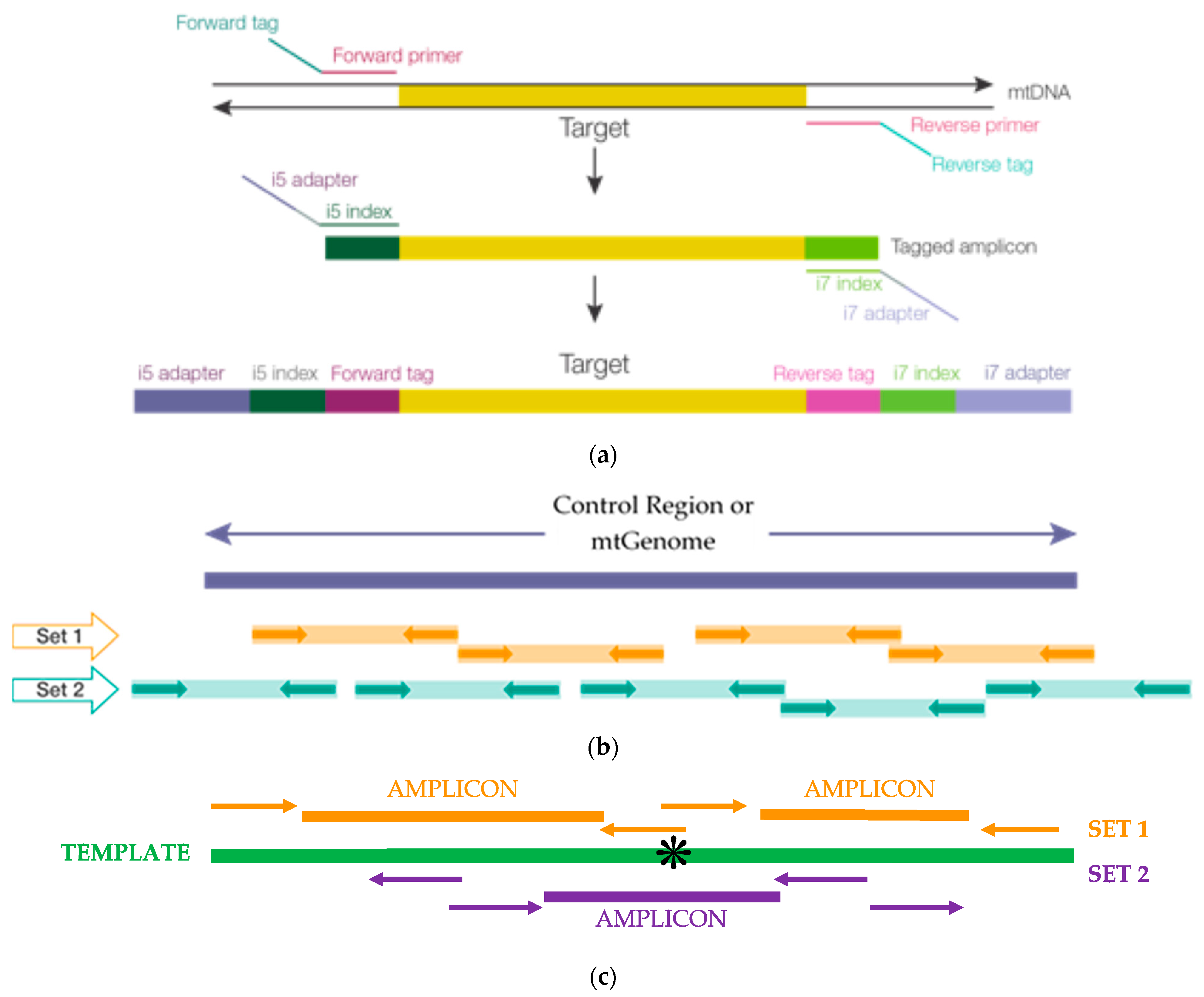
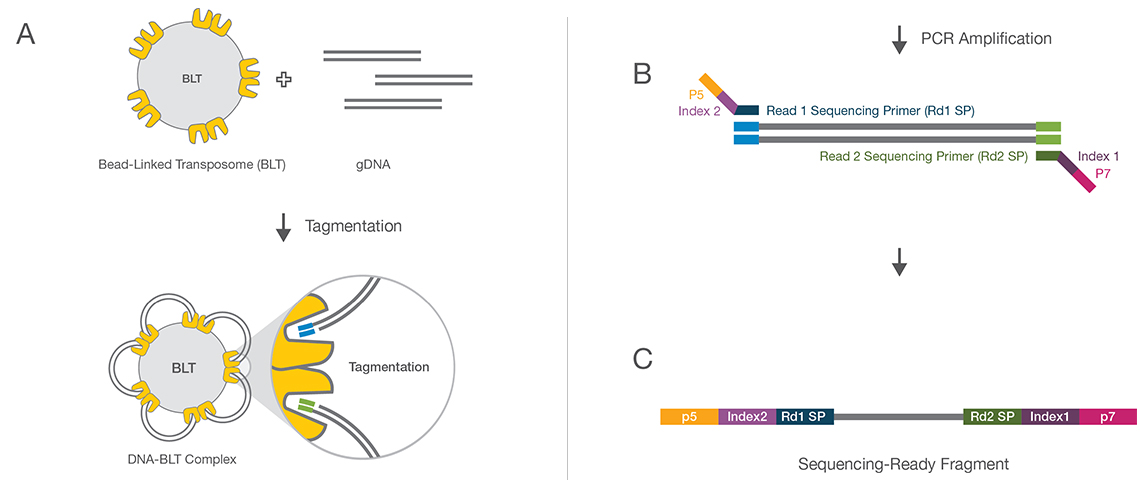
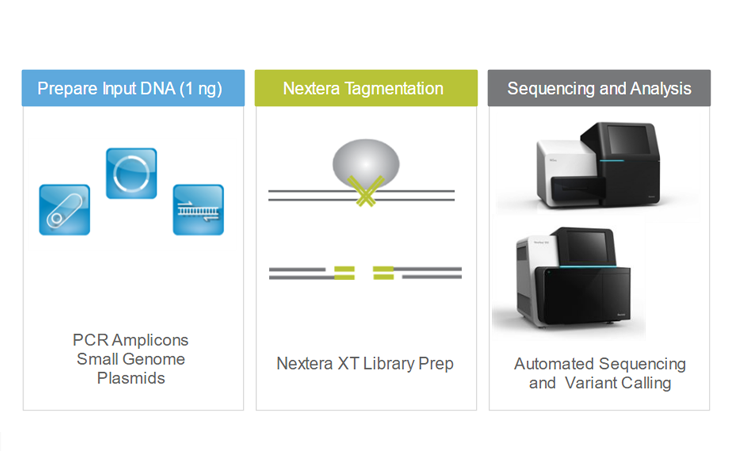
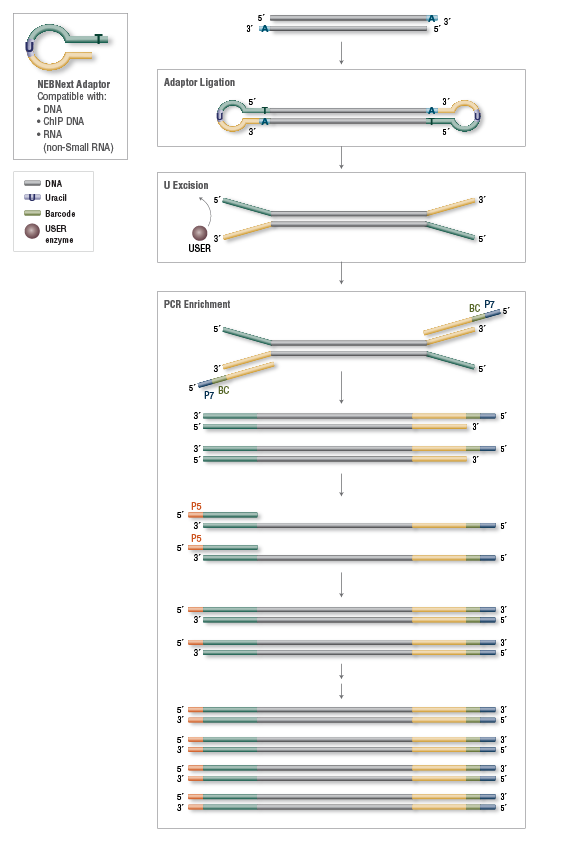
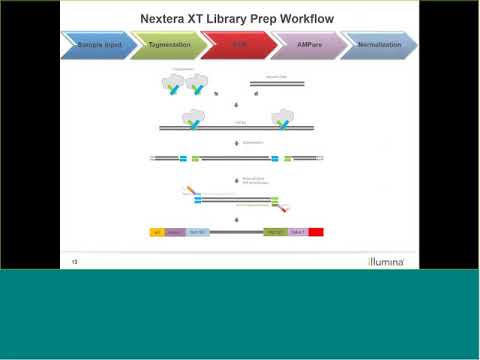
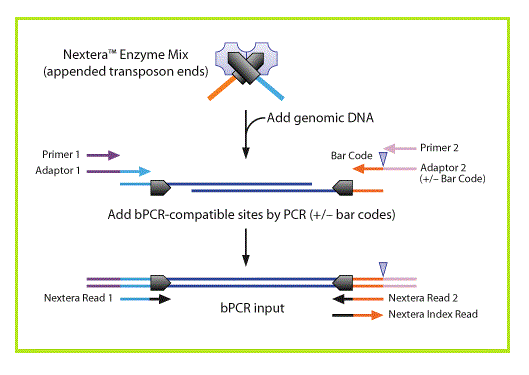
Comparing library preparation methods for SARS-CoV-2 multiplex amplicon sequencing on the Illumina MiSeq platform | bioRxiv
Application of Nextera DNA Flex to bacterial amplicons. a Libraries... | Download Scientific Diagram
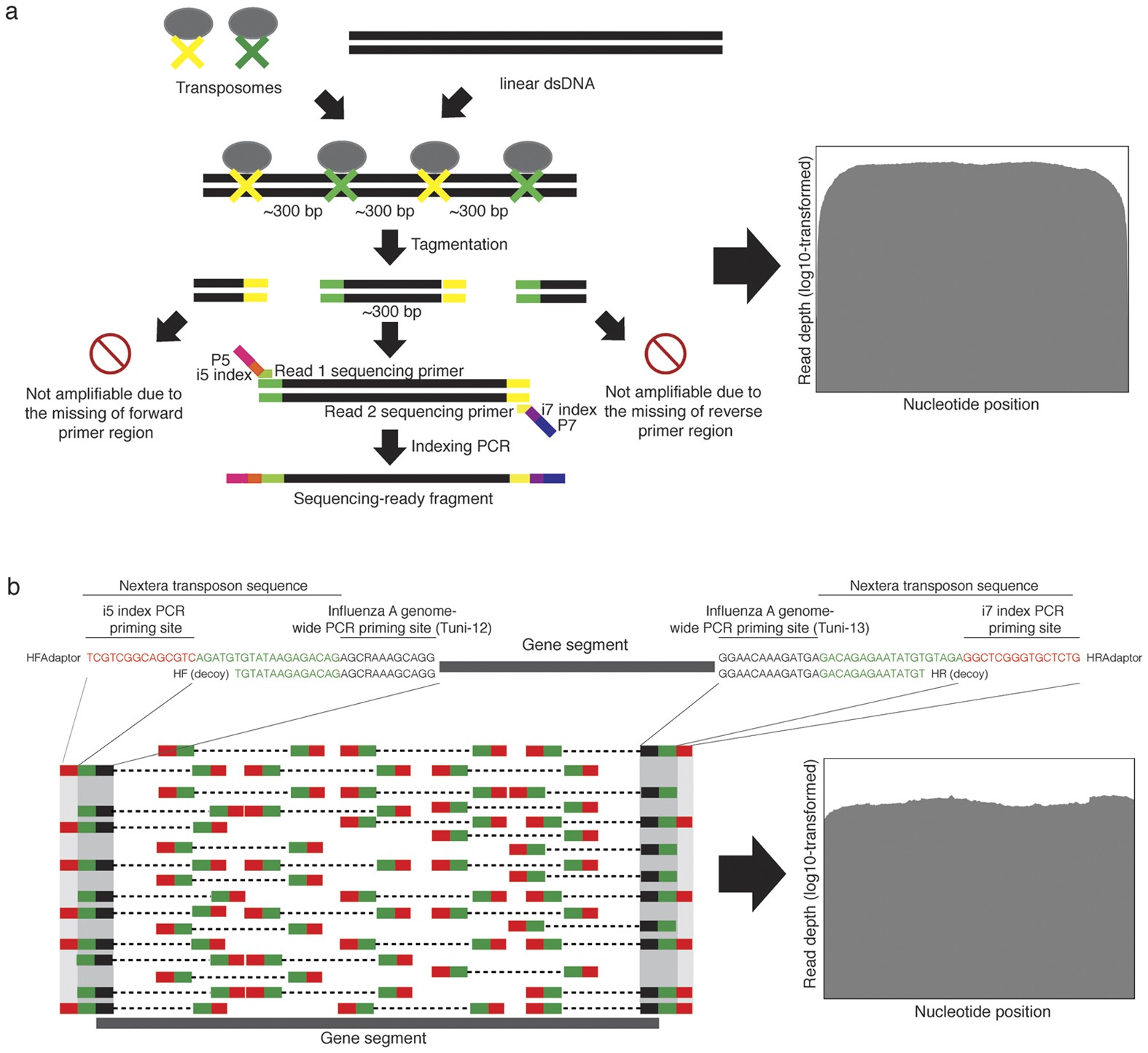
Contamination-controlled high-throughput whole genome sequencing for influenza A viruses using the MiSeq sequencer | Scientific Reports
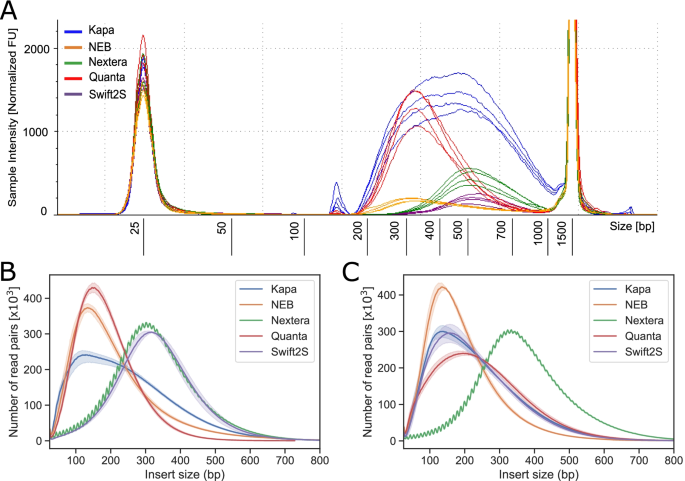
Optimization of enzymatic fragmentation is crucial to maximize genome coverage: a comparison of library preparation methods for Illumina sequencing | BMC Genomics | Full Text