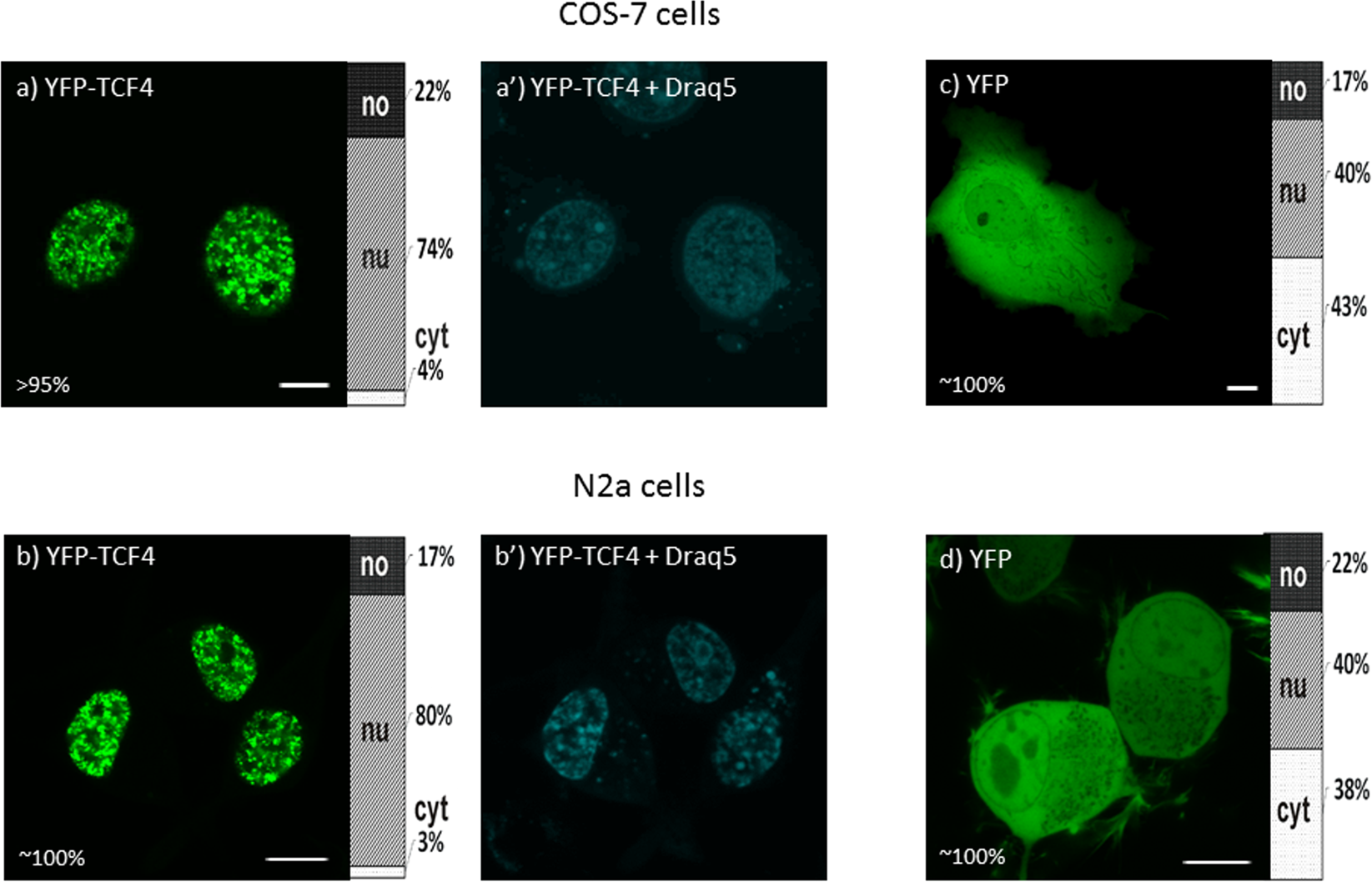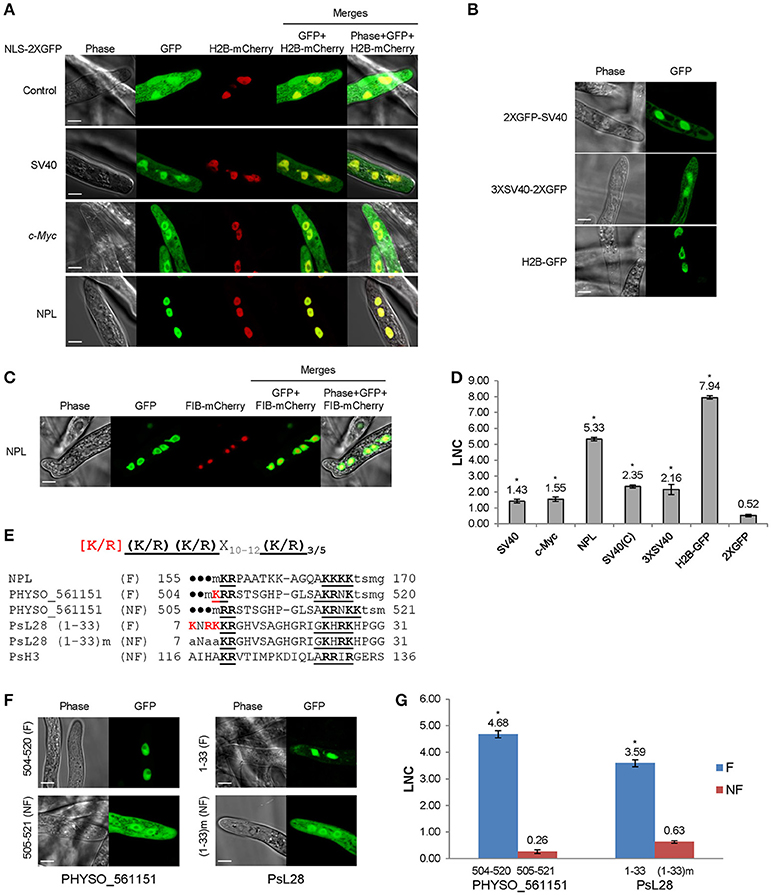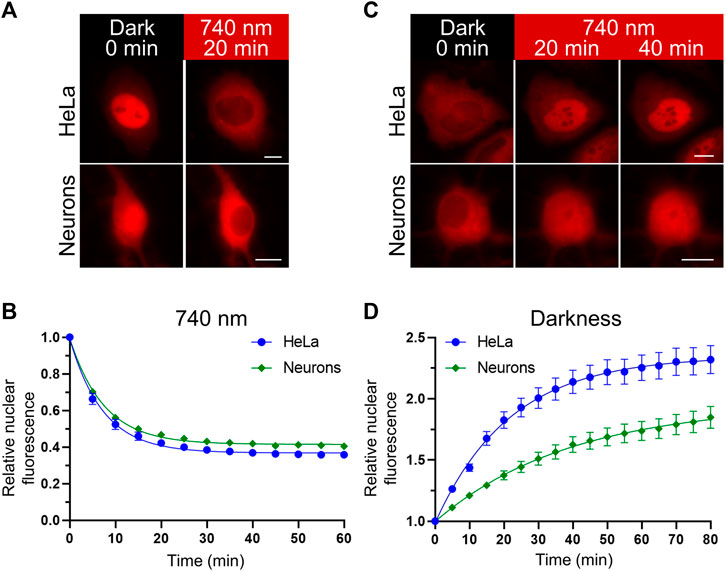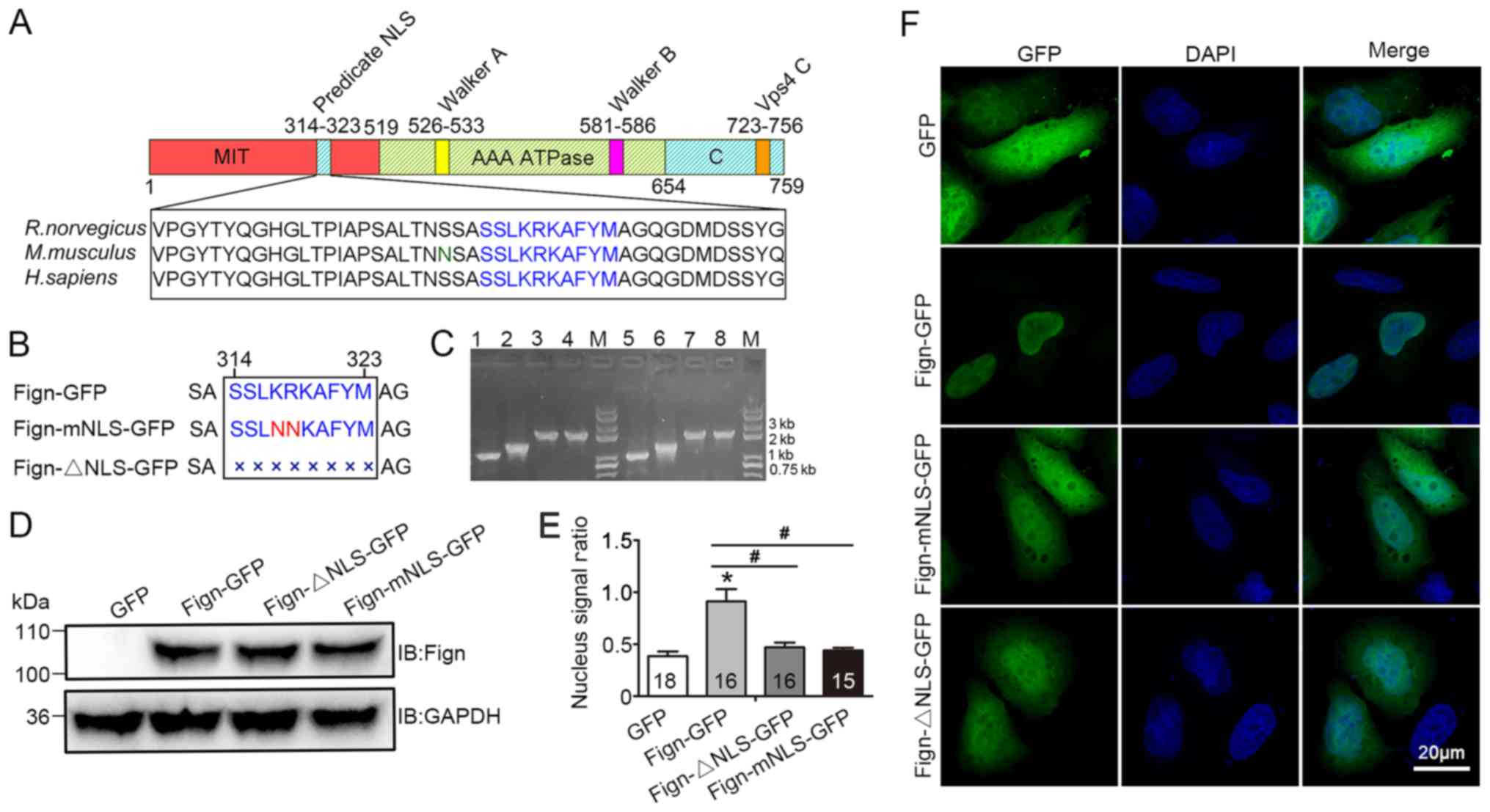
A nuclear localization signal is required for the nuclear translocation of Fign and its microtubule‑severing function

The flow charts of predicting NLS. (a) The flow chart of mining the... | Download Scientific Diagram

Discovering nuclear targeting signal sequence through protein language learning and multivariate analysis - ScienceDirect
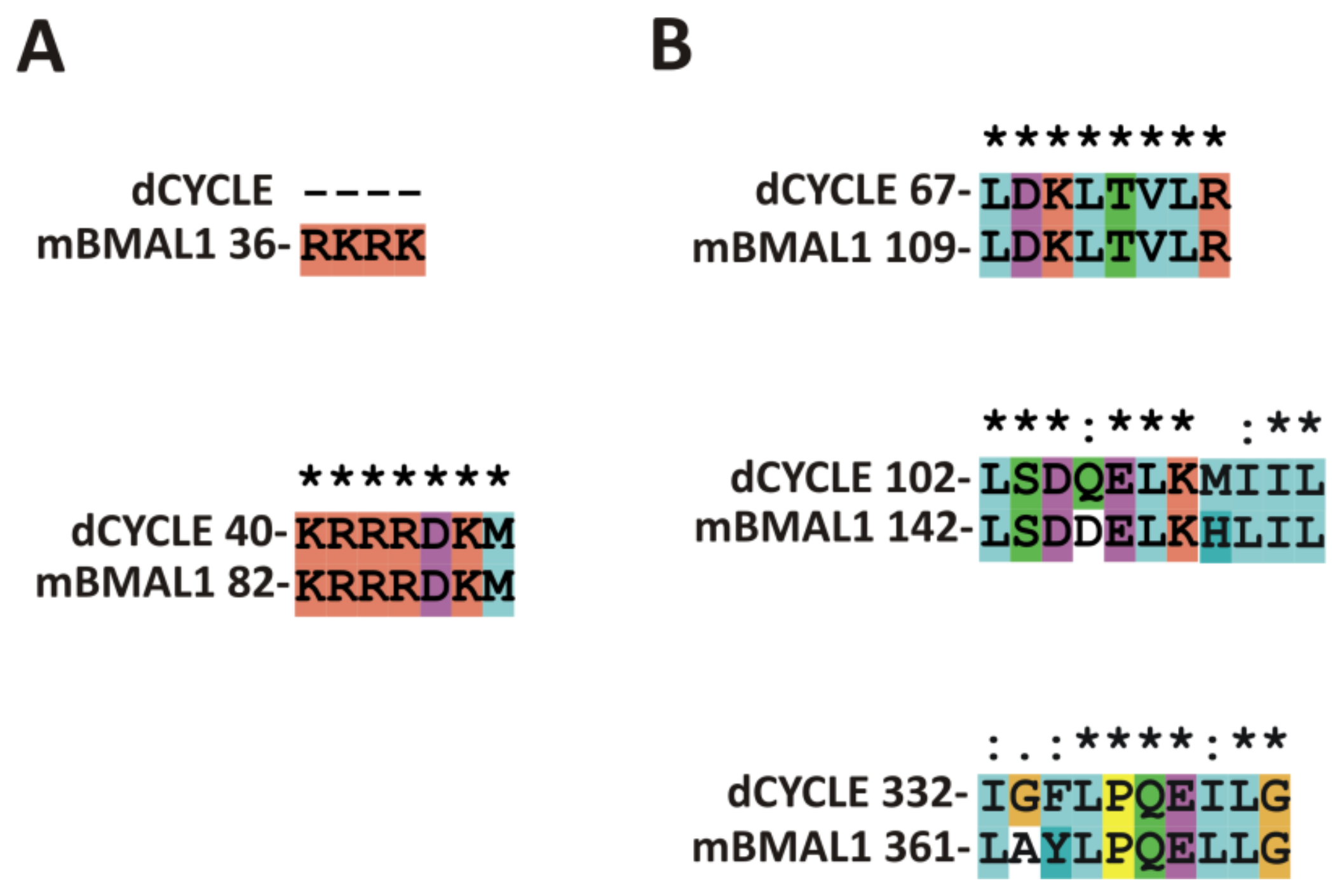
IJMS | Free Full-Text | Subcellular Localization Signals of bHLH-PAS Proteins: Their Significance, Current State of Knowledge and Future Perspectives
Identification of the nuclear localization sequence (NLS) in PARN. (A)... | Download Scientific Diagram

Cells | Free Full-Text | Bioinformatics and Functional Analysis of a New Nuclear Localization Sequence of the Influenza A Virus Nucleoprotein
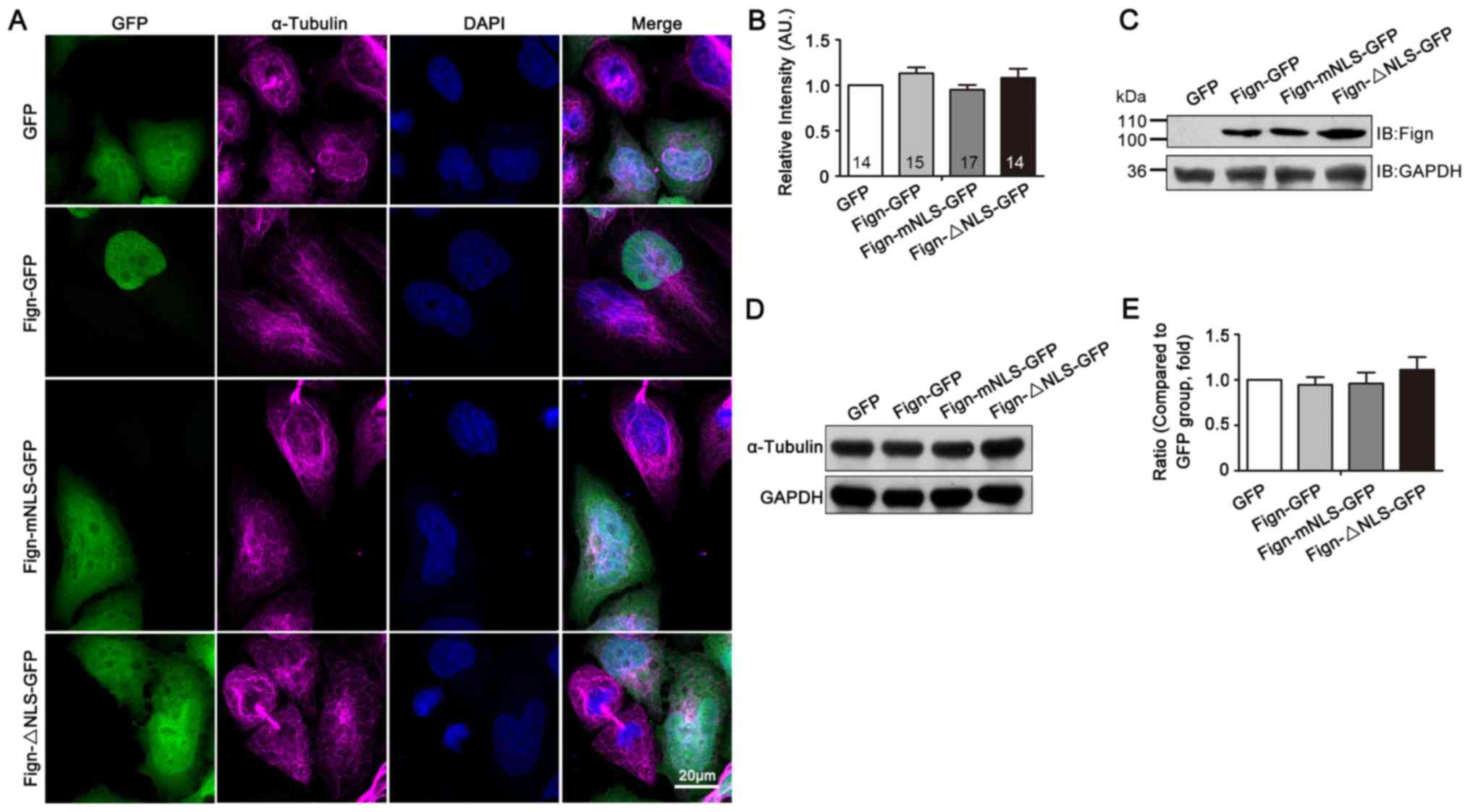
A nuclear localization signal is required for the nuclear translocation of Fign and its microtubule‑severing function

Potential NLS sequences in TPs of Streptomyces. (A) The archetypal Tpg... | Download Scientific Diagram

Prediction results for putative NLS, NES and DNA-binding motifs on CAV... | Download Scientific Diagram
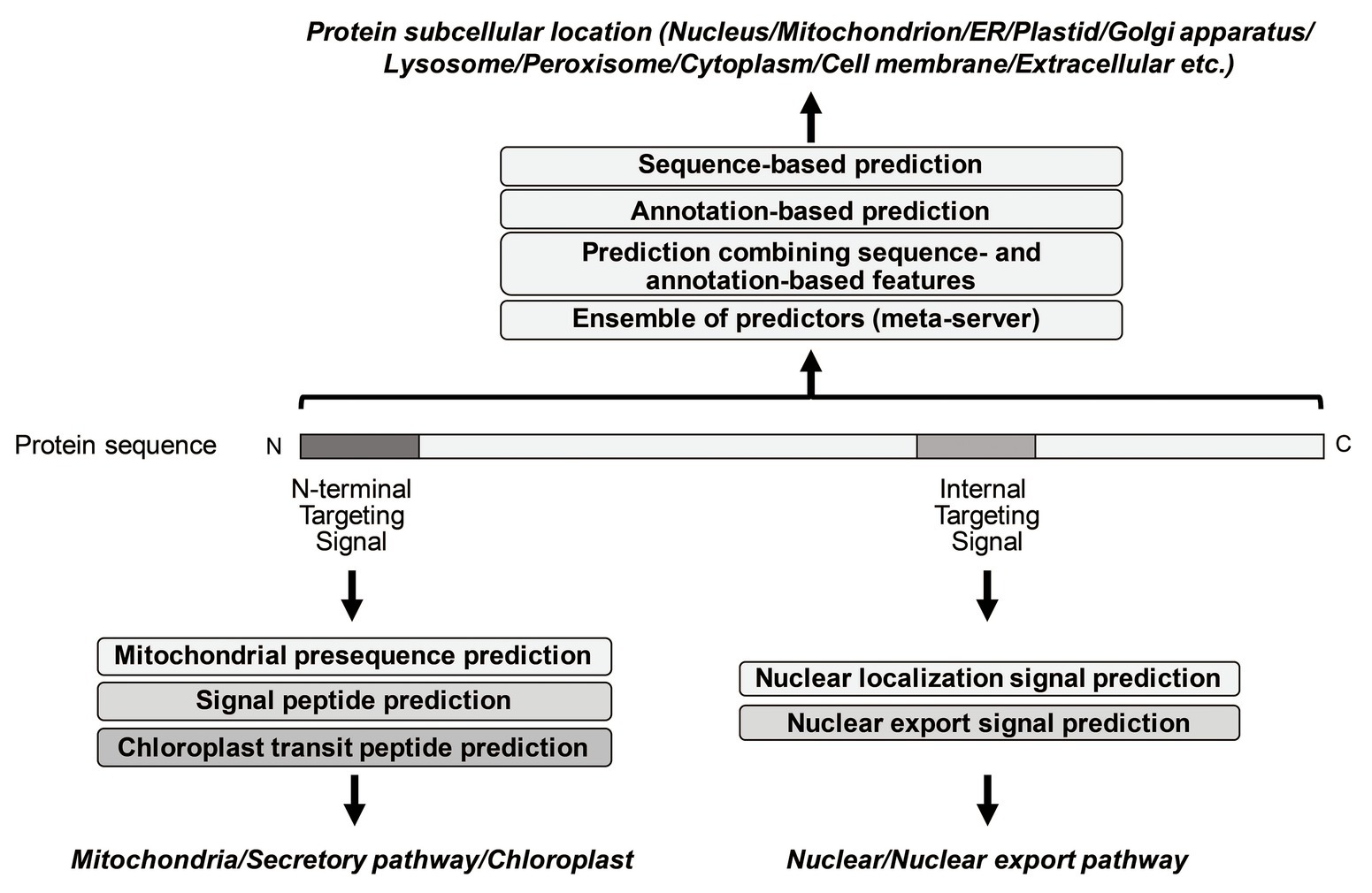
Frontiers | Tools for the Recognition of Sorting Signals and the Prediction of Subcellular Localization of Proteins From Their Amino Acid Sequences
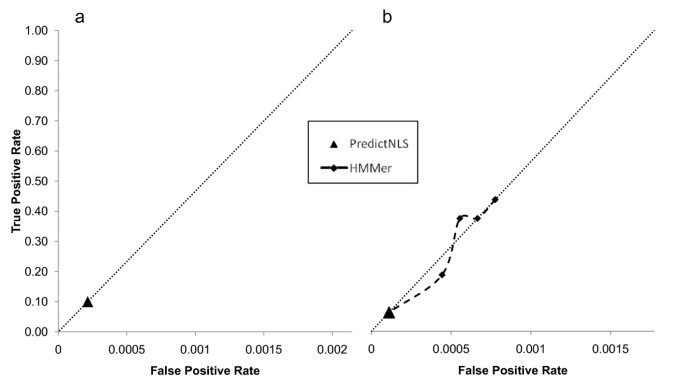
NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction | BMC Bioinformatics | Full Text

Identification of a bipartite nuclear localization signal in the silkworm Masc protein - Sugano - 2016 - FEBS Letters - Wiley Online Library