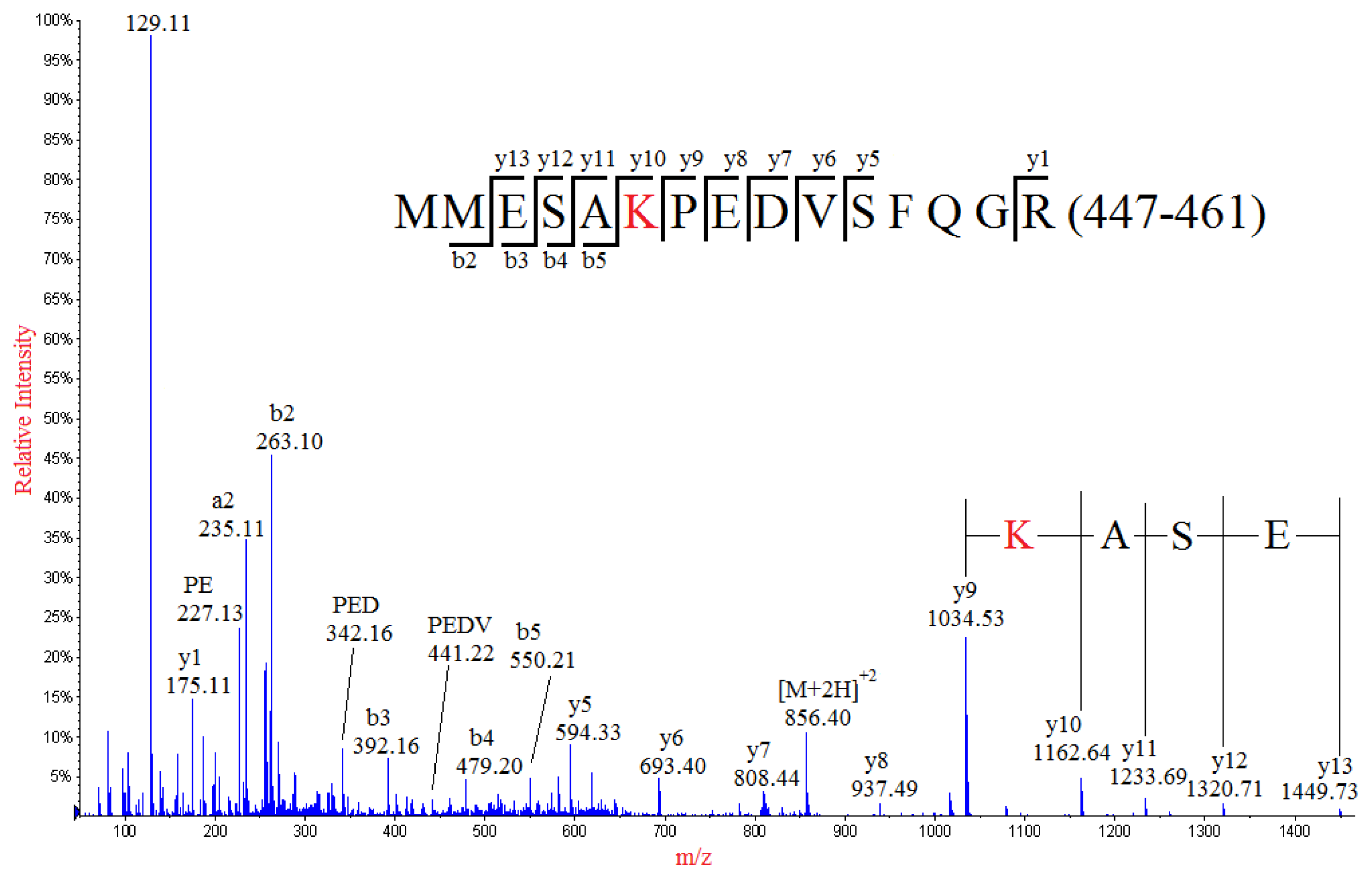
IJMS | Free Full-Text | Mass Spectrometry Analysis Coupled with de novo Sequencing Reveals Amino Acid Substitutions in Nucleocapsid Protein from Influenza A Virus

Protein/Peptide Sequence Analysis: Current Methodologies (English Edition) eBook : Bhown, A.S.: Amazon.de: Kindle Store

Determination of sequence and absolute configuration of peptide amino acids by HPLC–MS/CD-based detection of liberated N-terminus phenylthiohydantoin amino acids | Scientific Reports
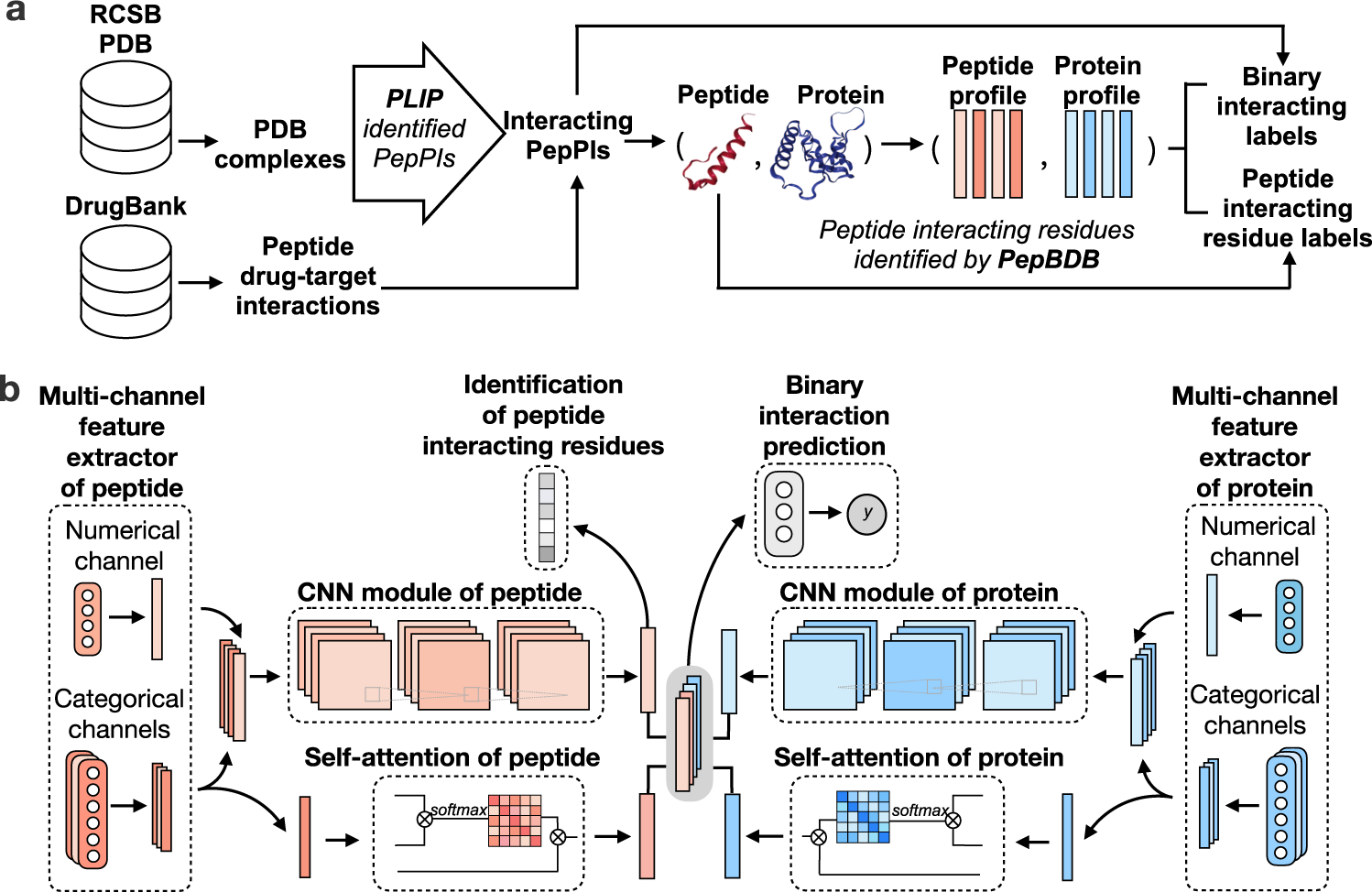
A deep-learning framework for multi-level peptide–protein interaction prediction | Nature Communications

Sequence analysis of the resistance peptides. (A) Alignments of the... | Download Scientific Diagram

Molecules | Free Full-Text | PepFun: Open Source Protocols for Peptide-Related Computational Analysis


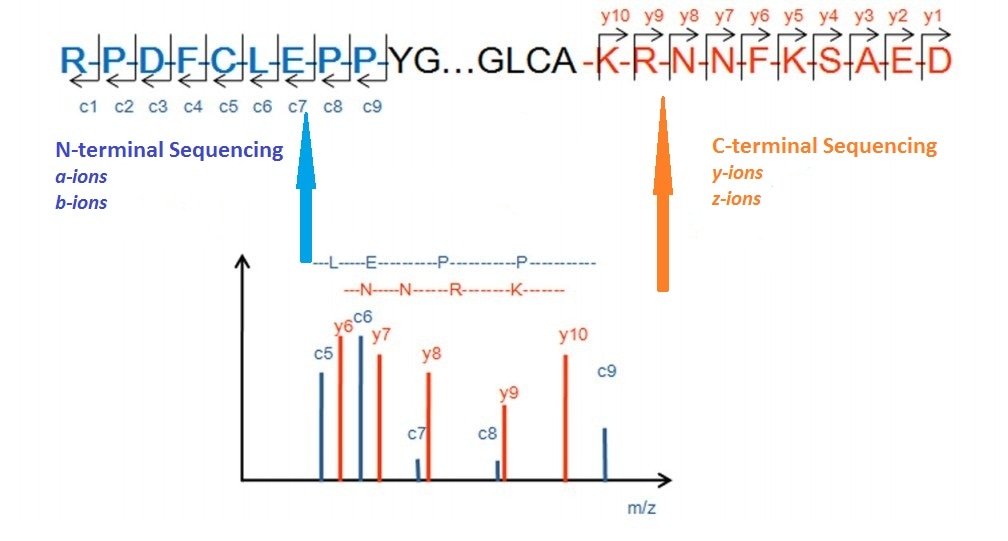


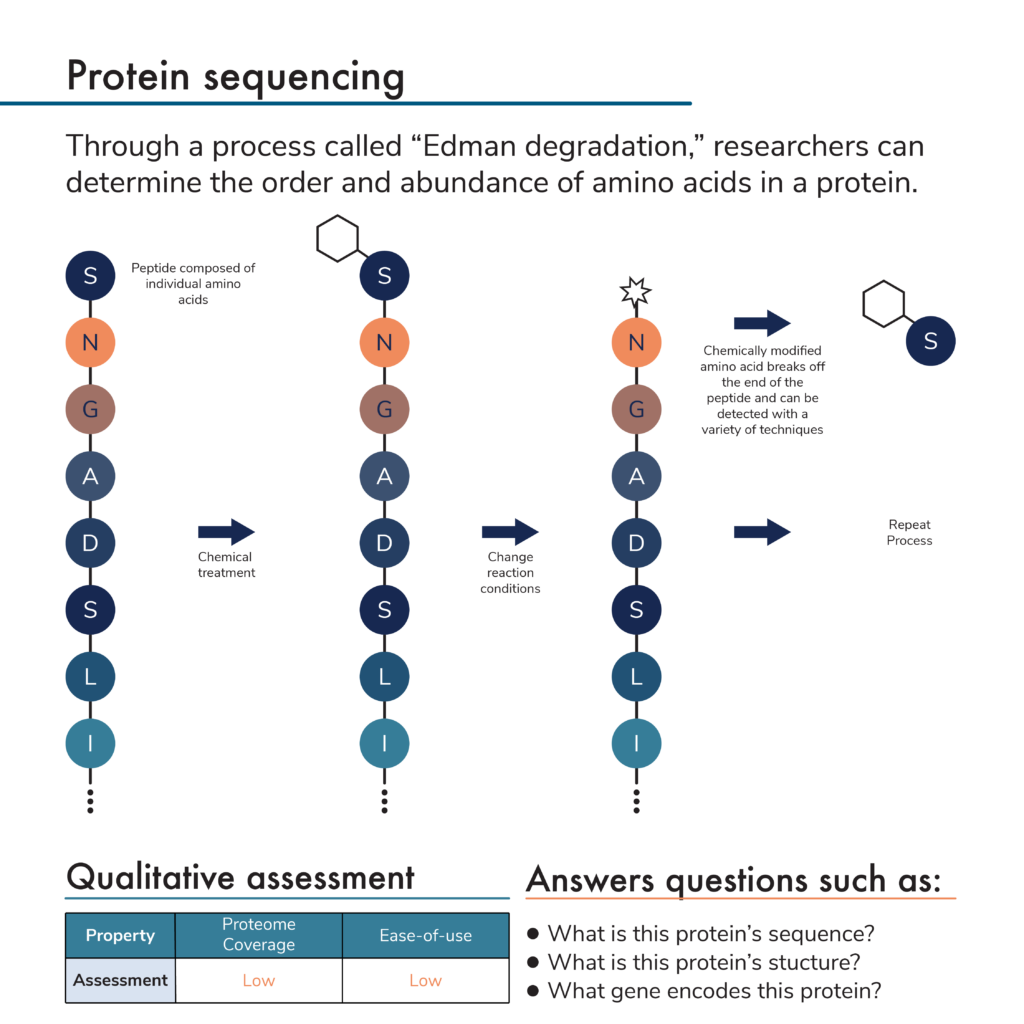




![PDF] Tandem mass spectrometry for peptide and protein sequence analysis. | Semantic Scholar PDF] Tandem mass spectrometry for peptide and protein sequence analysis. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/eddc094667af4da7b3698c26230b0b1fbbe08448/2-Figure2-1.png)