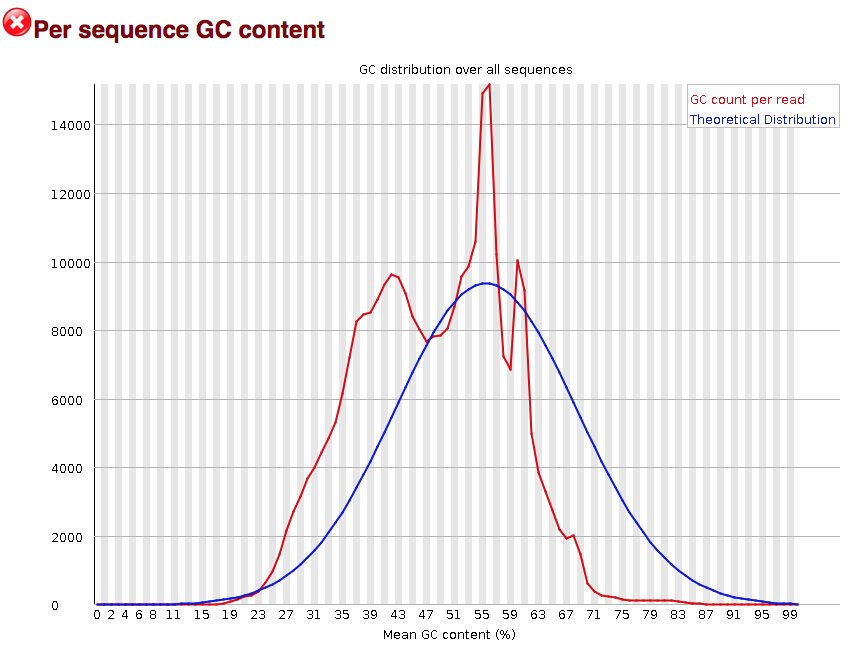
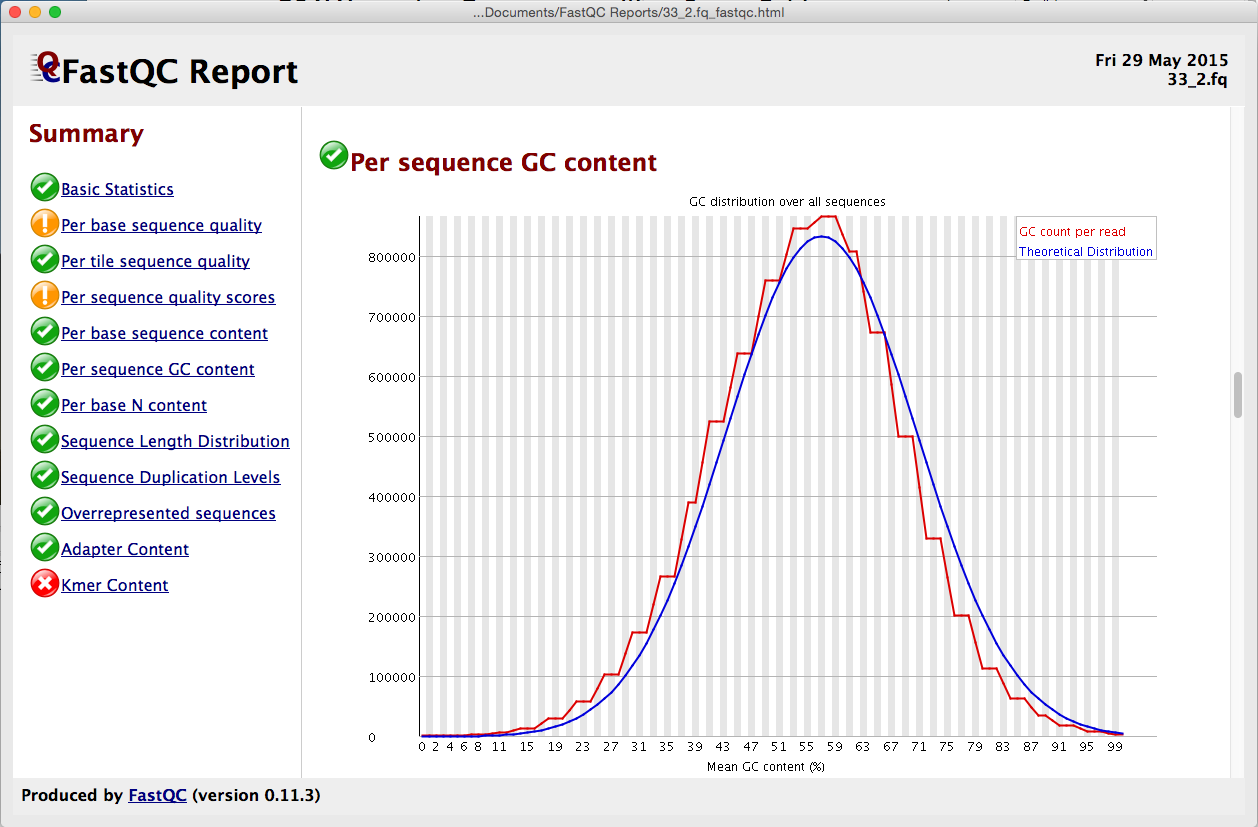
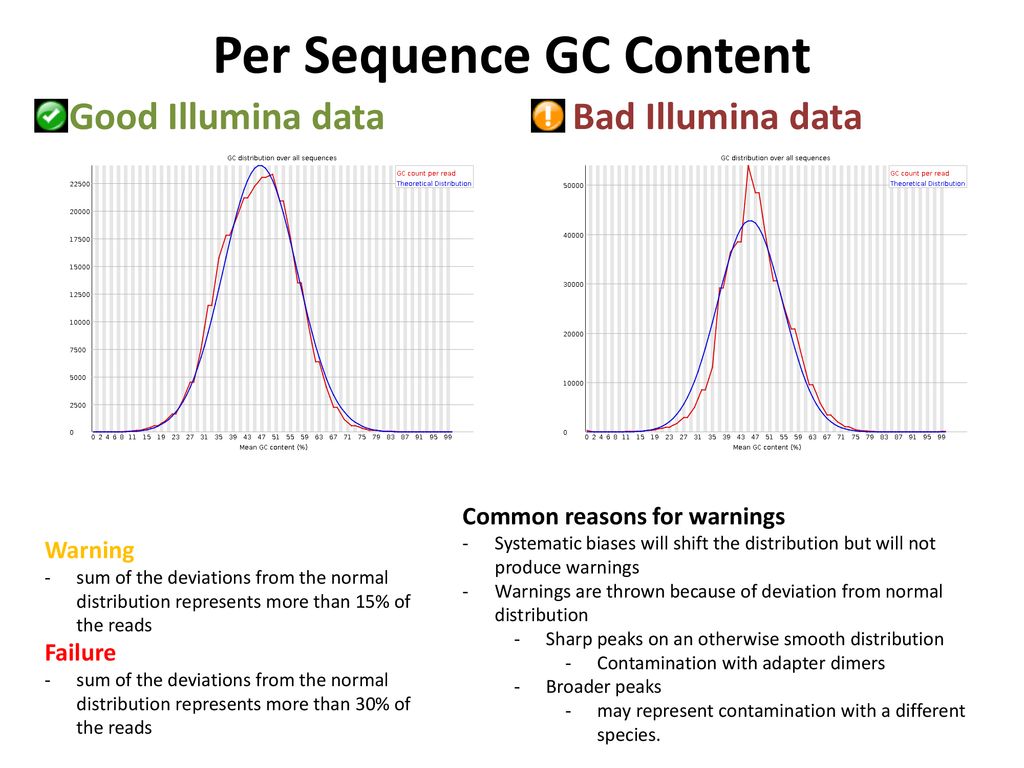
FastQC for RNA-Seq-Fails for per base seq content, gc content, duplication levels, adapter content and kmer content? | ResearchGate

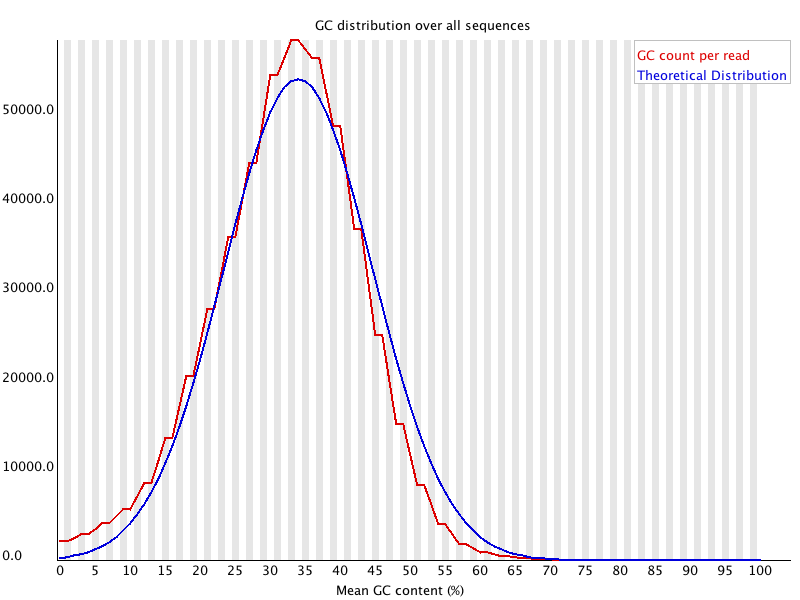
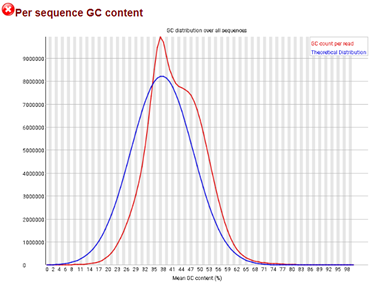
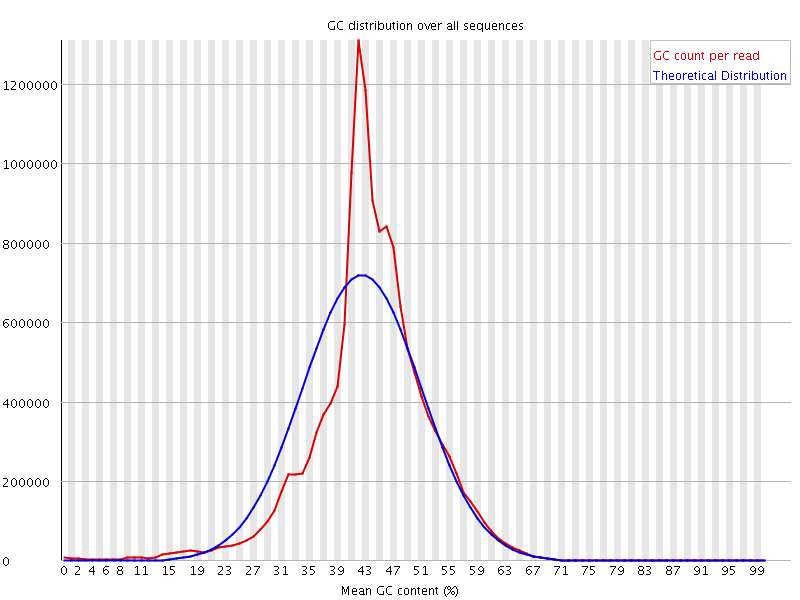
Representative graph of the mean GC content for raw data obtained from... | Download Scientific Diagram

SciBear🇺🇦 on Twitter: "(1/7) A quality control tool for raw #sequence data. Using #FastQC you may check: 🚀 Per base sequence #quality (do you see a drop in sequencing quality near the