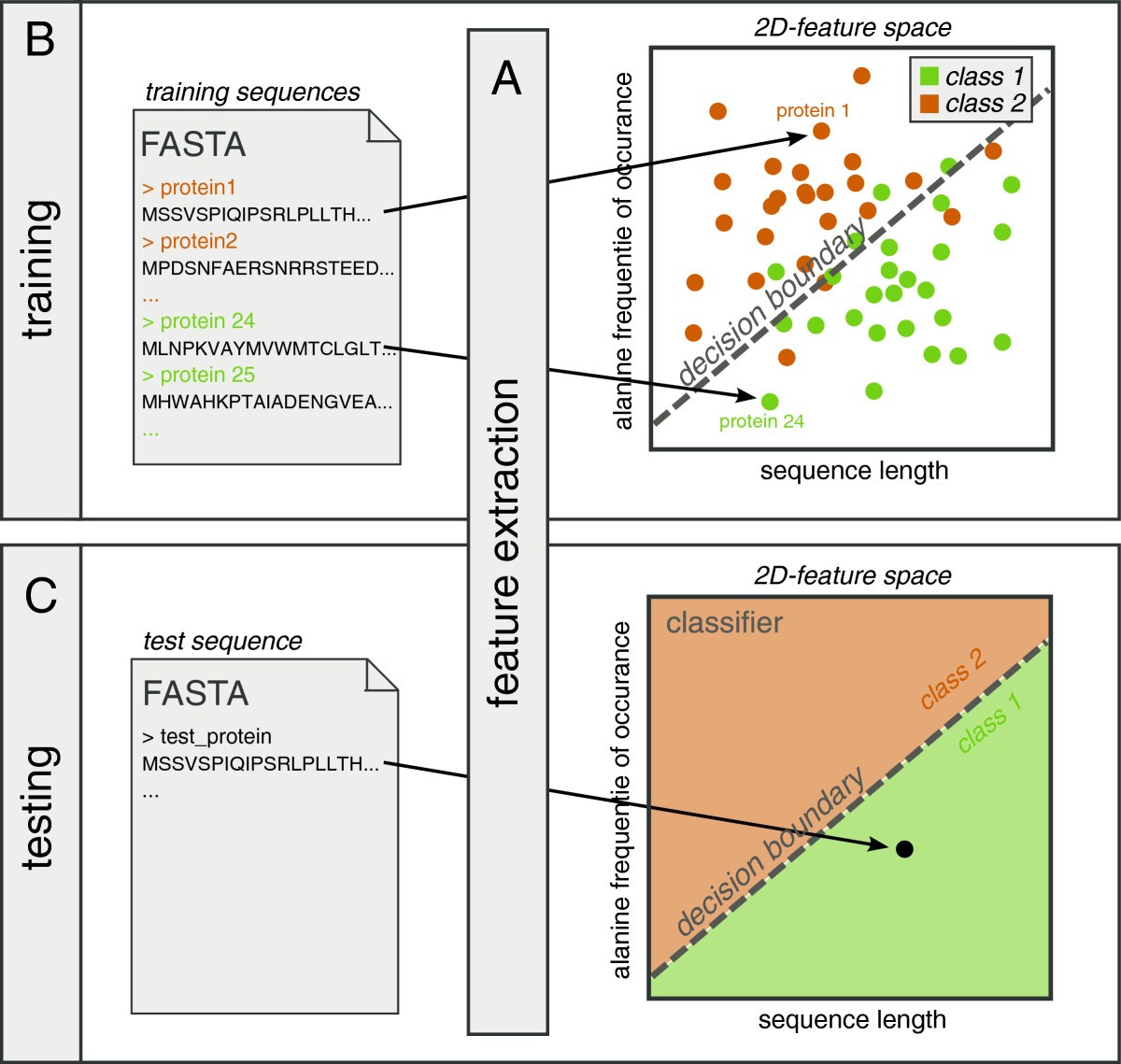
SPiCE: a web-based tool for sequence-based protein classification and exploration | BMC Bioinformatics | Full Text

Co‐Evolutionary Fitness Landscapes for Sequence Design - Tian - 2018 - Angewandte Chemie International Edition - Wiley Online Library
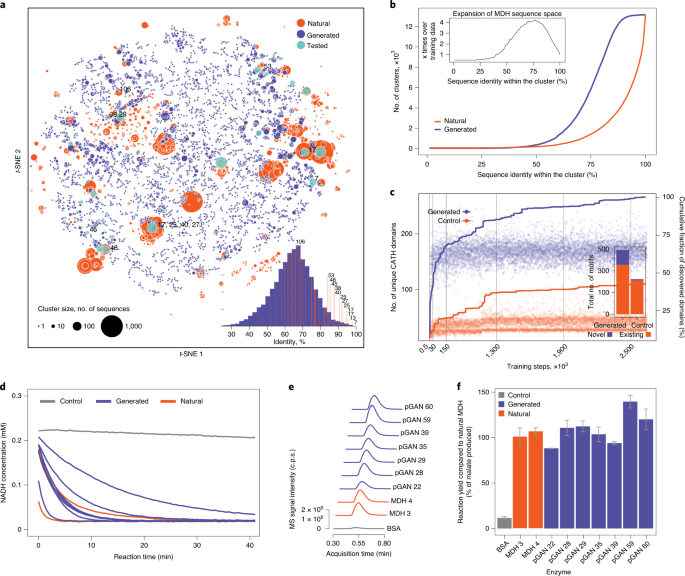
Expanding functional protein sequence spaces using generative adversarial networks | Nature Machine Intelligence

Efficient Exploration of Sequence Space by Sequence-Guided Protein Engineering and Design | Biochemistry

How much of protein sequence space has been explored by life on Earth? | Journal of The Royal Society Interface
Benchmarking Inverse Statistical Approaches for Protein Structure and Design with Exactly Solvable Models | PLOS Computational Biology

Figure 1 from Synthetic biology for the directed evolution of protein biocatalysts: navigating sequence space intelligently | Semantic Scholar

The space of foldable HP sequences gives a glimpse of what real protein... | Download Scientific Diagram
![Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ] Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/cs-684/1/fig-2-full.png)
Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]

Neural networks to learn protein sequence–function relationships from deep mutational scanning data | PNAS

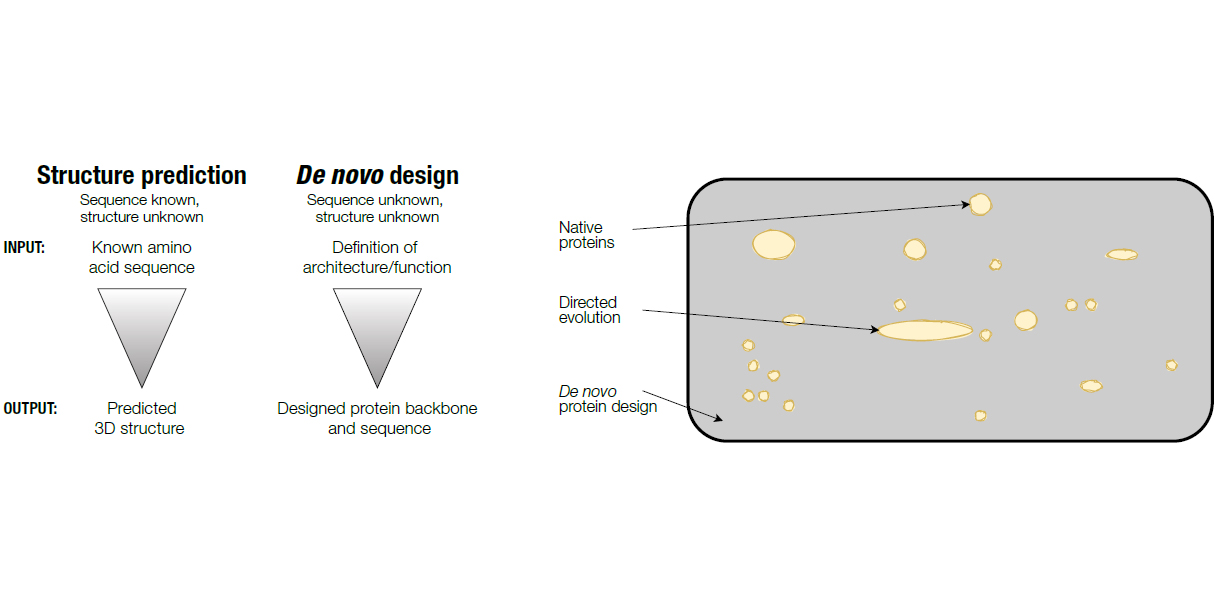


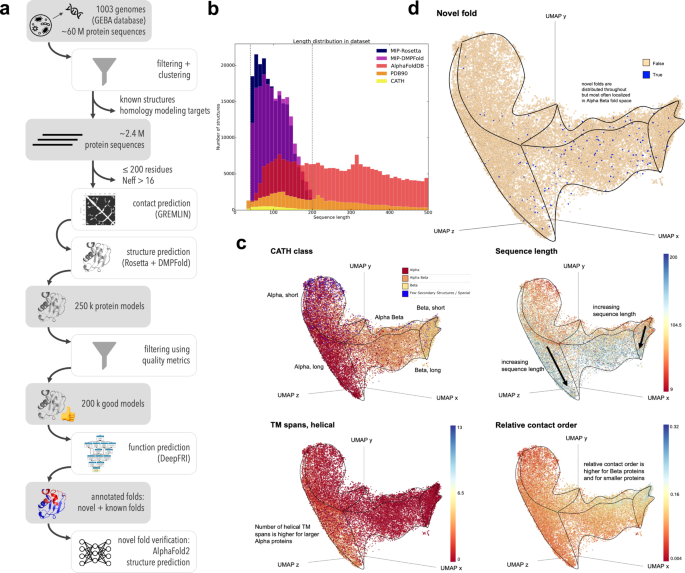
![PDF] On the universe of protein folds. | Semantic Scholar PDF] On the universe of protein folds. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/4feb83e934a4dd006a898c2eece9d007fcb001c5/16-Figure4-1.png)