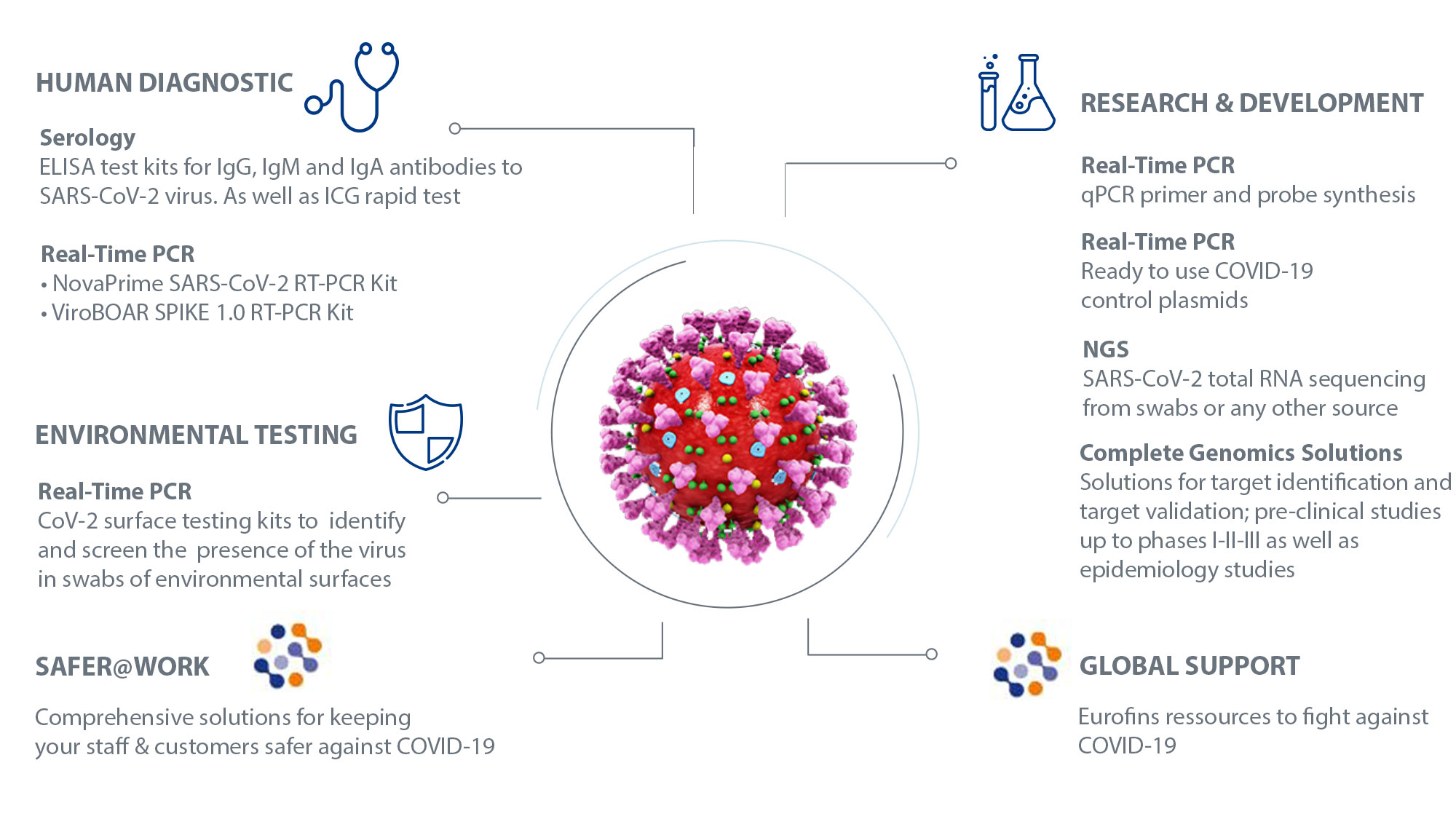
Massive and rapid COVID-19 testing is feasible by extraction-free SARS-CoV-2 RT-PCR | Nature Communications
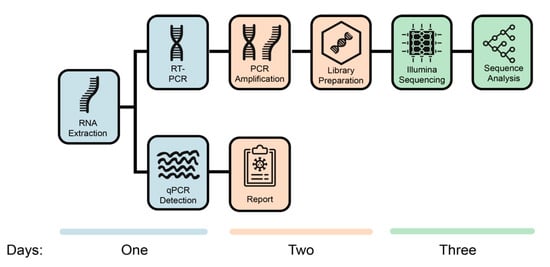
Genes | Free Full-Text | Whole Genome Sequencing of SARS-CoV-2: Adapting Illumina Protocols for Quick and Accurate Outbreak Investigation during a Pandemic
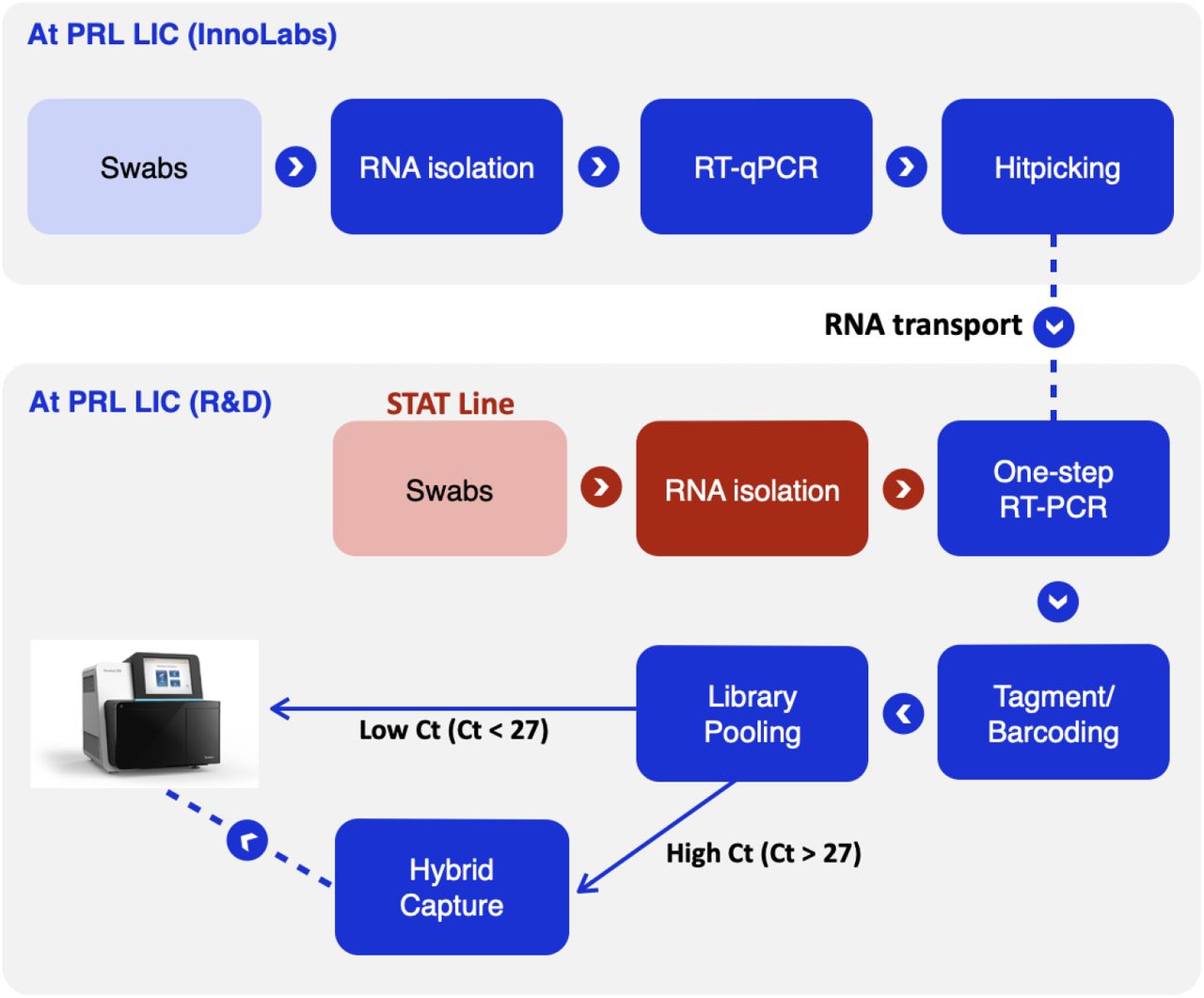
Development and scaling of a sequencing pipeline for genomic surveillance of SARS-CoV-2 in New York City | medRxiv
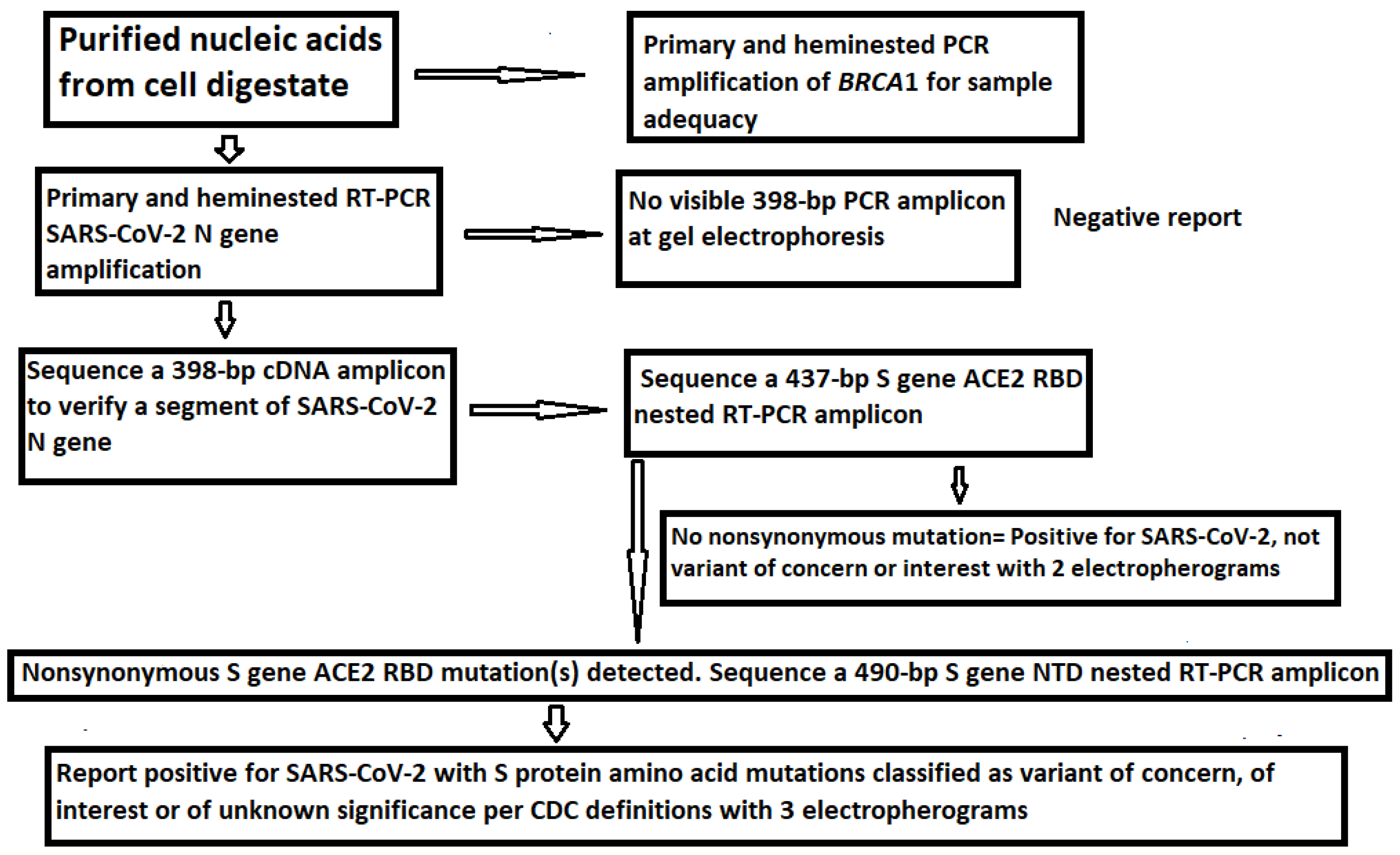
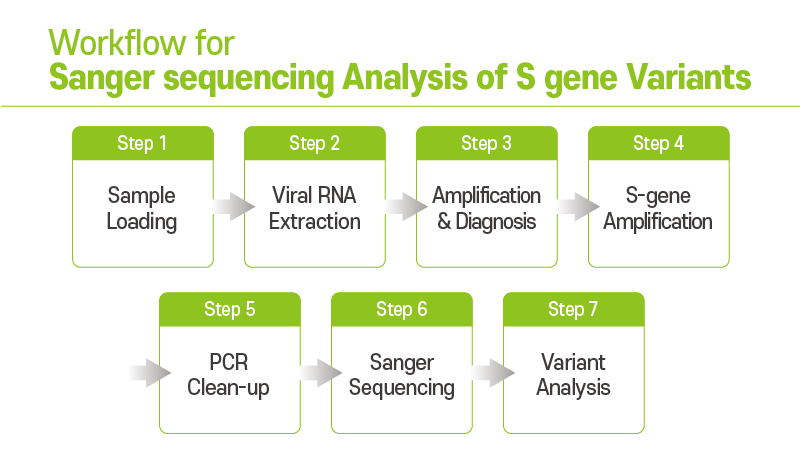
Viruses | Free Full-Text | A Routine Sanger Sequencing Target Specific Mutation Assay for SARS-CoV-2 Variants of Concern and Interest
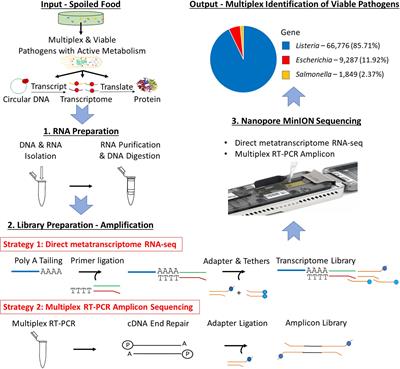
Frontiers | Direct Metatranscriptome RNA-seq and Multiplex RT-PCR Amplicon Sequencing on Nanopore MinION – Promising Strategies for Multiplex Identification of Viable Pathogens in Food
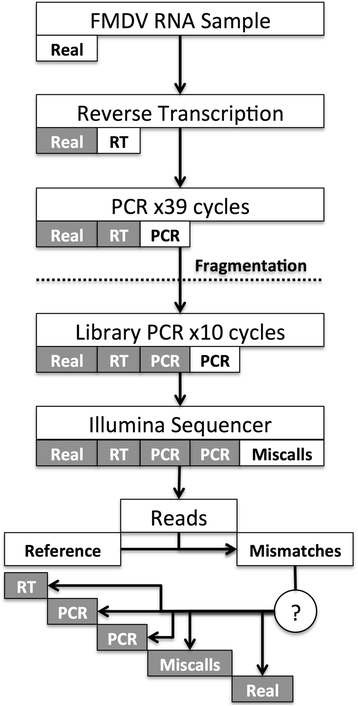
Distinguishing low frequency mutations from RT-PCR and sequence errors in viral deep sequencing data | BMC Genomics | Full Text
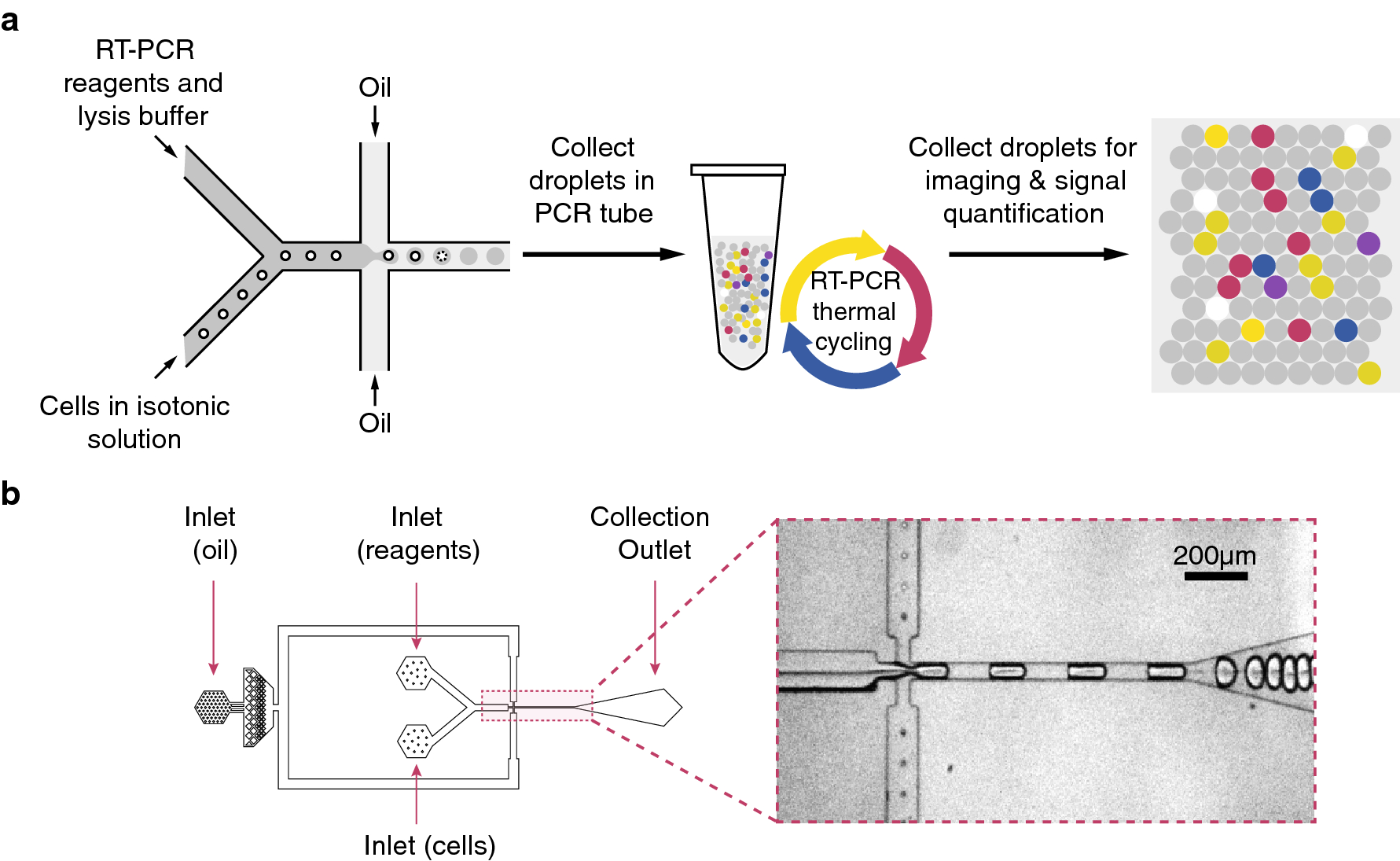
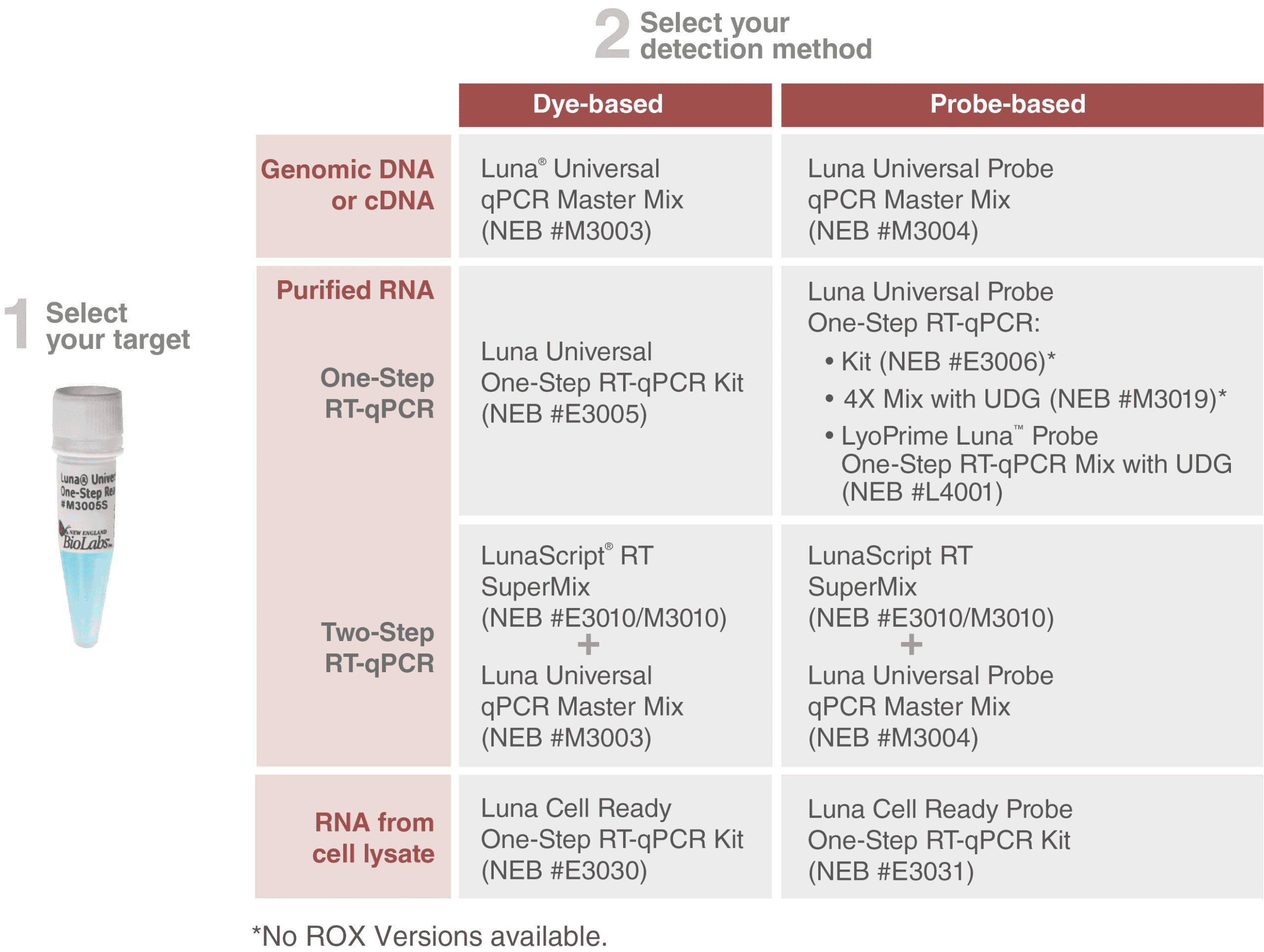
Microdroplet-based one-step RT-PCR for ultrahigh throughput single-cell multiplex gene expression analysis and rare cell detection | Scientific Reports
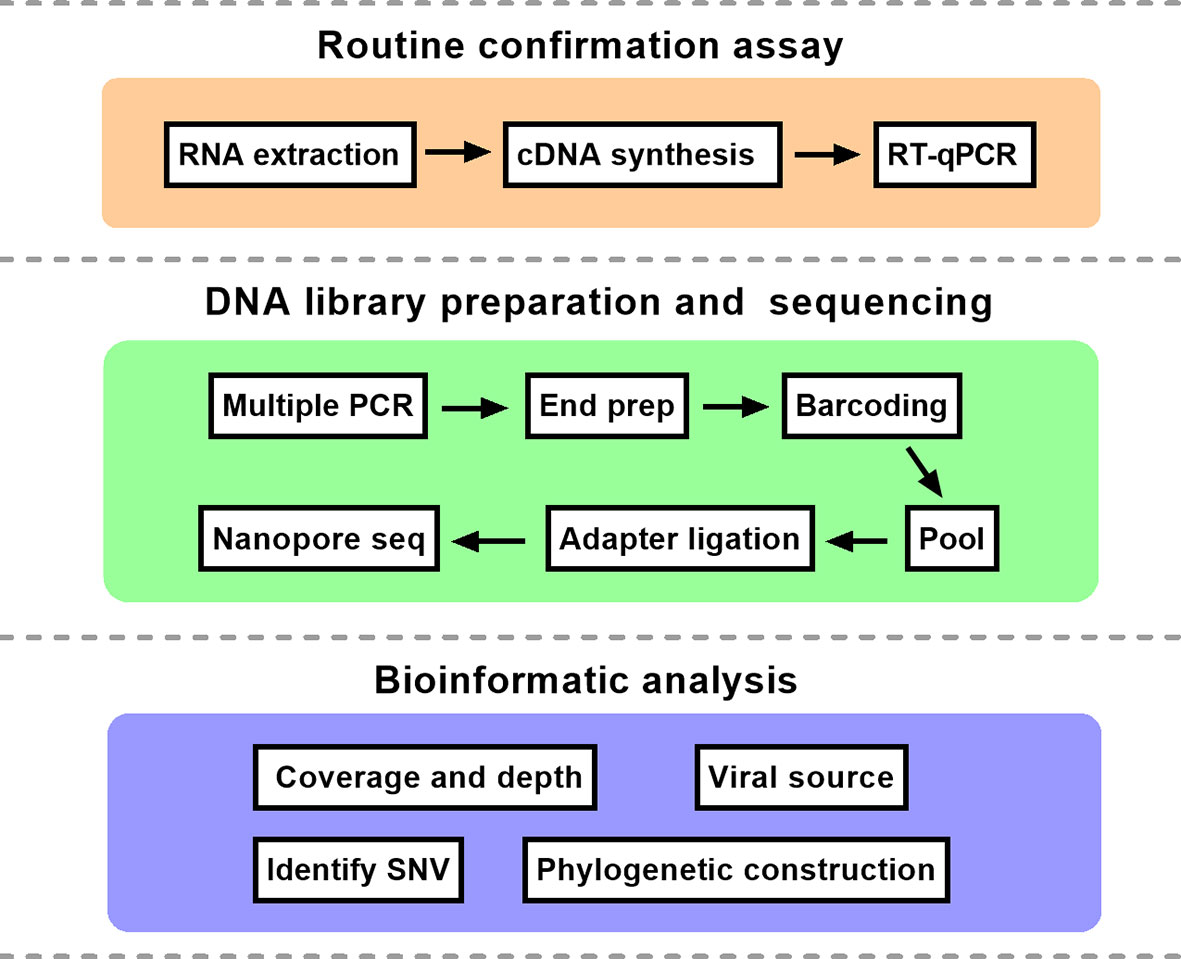
Frontiers | Genetic Surveillance of Five SARS-CoV-2 Clinical Samples in Henan Province Using Nanopore Sequencing

Rapid High-Throughput Whole-Genome Sequencing of SARS-CoV-2 by Using One-Step Reverse Transcription-PCR Amplification with an Integrated Microfluidic System and Next-Generation Sequencing | Journal of Clinical Microbiology

Genotyping of hepatitis C virus by nucleotide sequencing: A robust method for a diagnostic laboratory - ScienceDirect
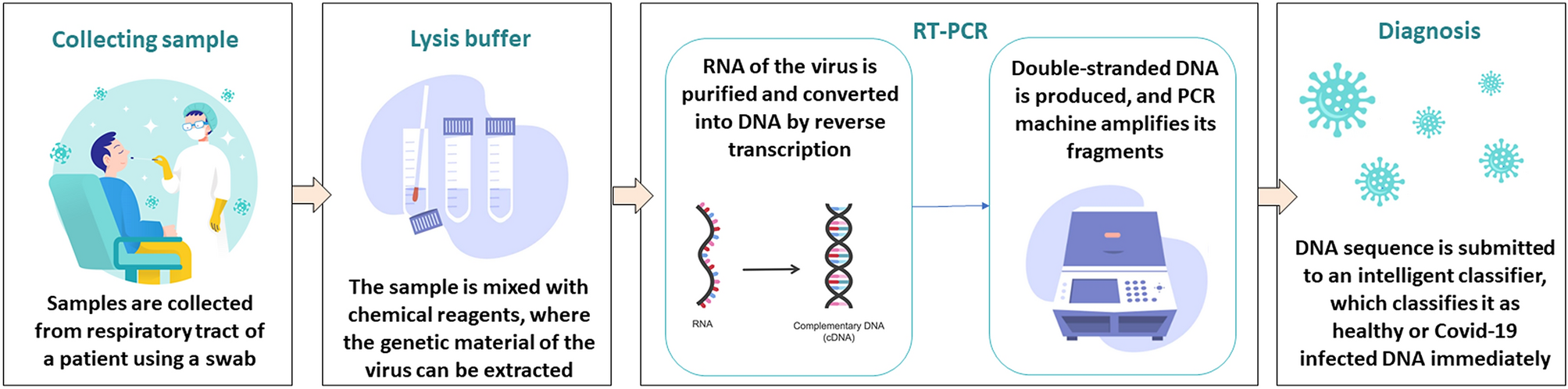
Covid-19 diagnosis by combining RT-PCR and pseudo-convolutional machines to characterize virus sequences | Scientific Reports
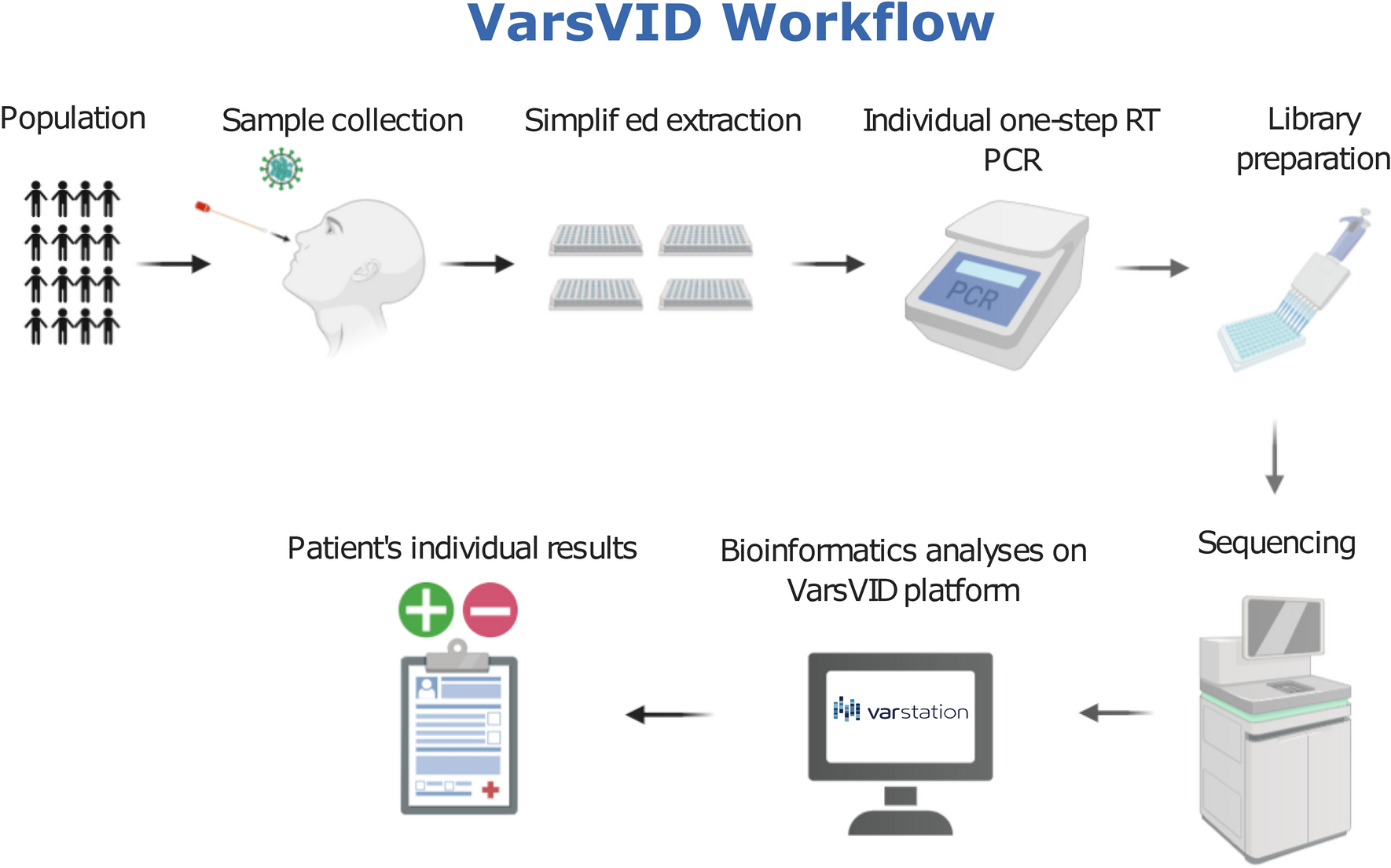
Mass molecular testing for COVID19 using NGS-based technology and a highly scalable workflow | Scientific Reports