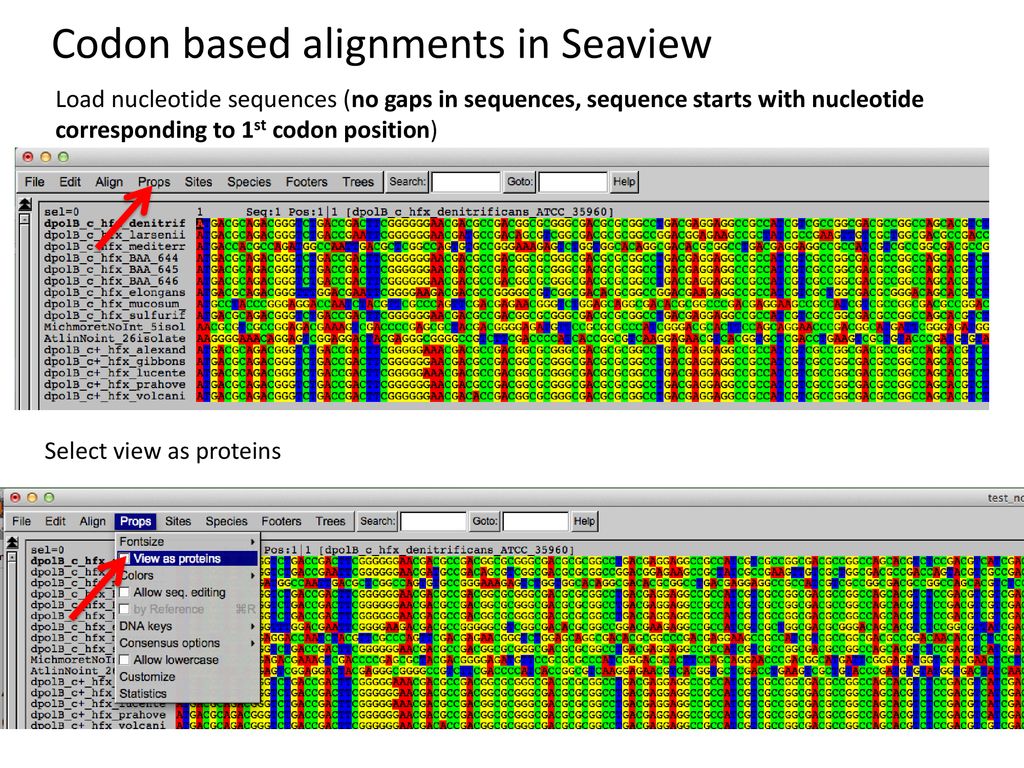
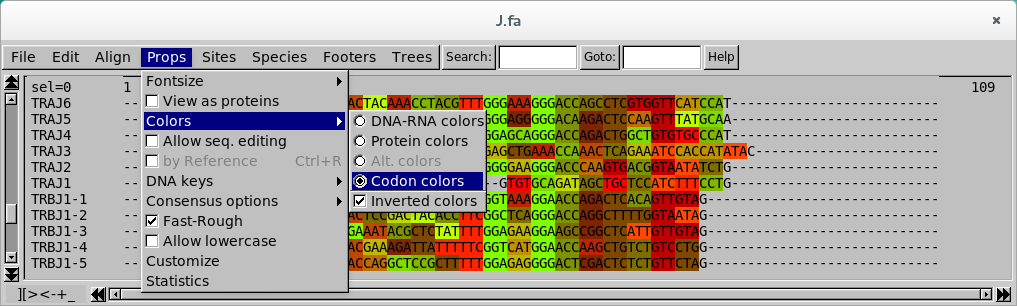
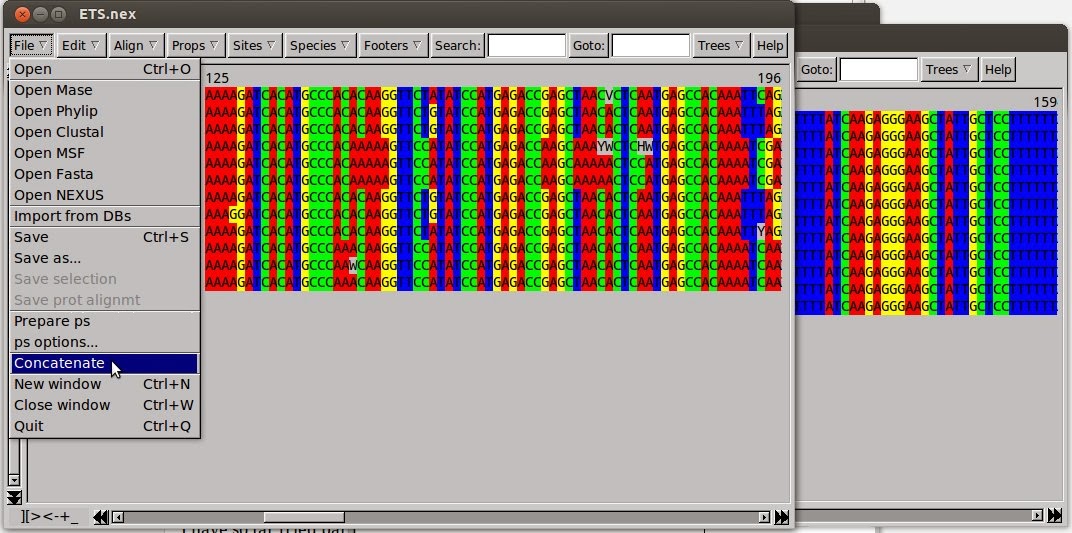
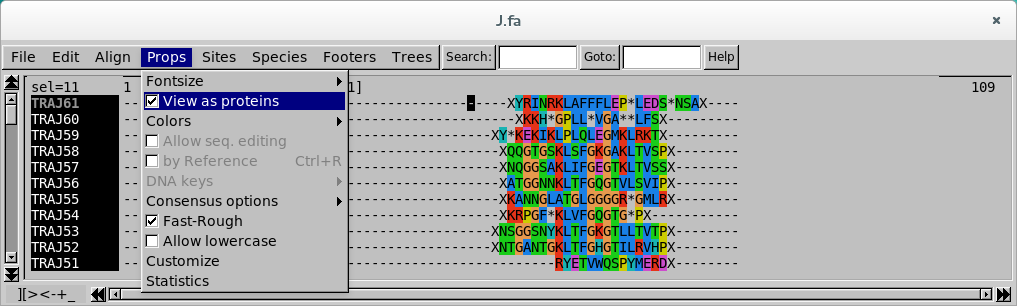
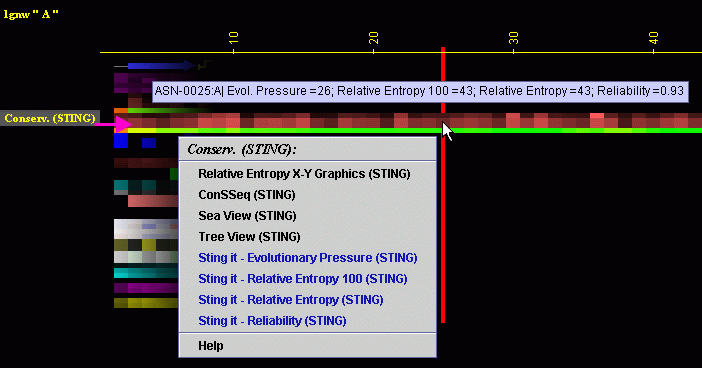
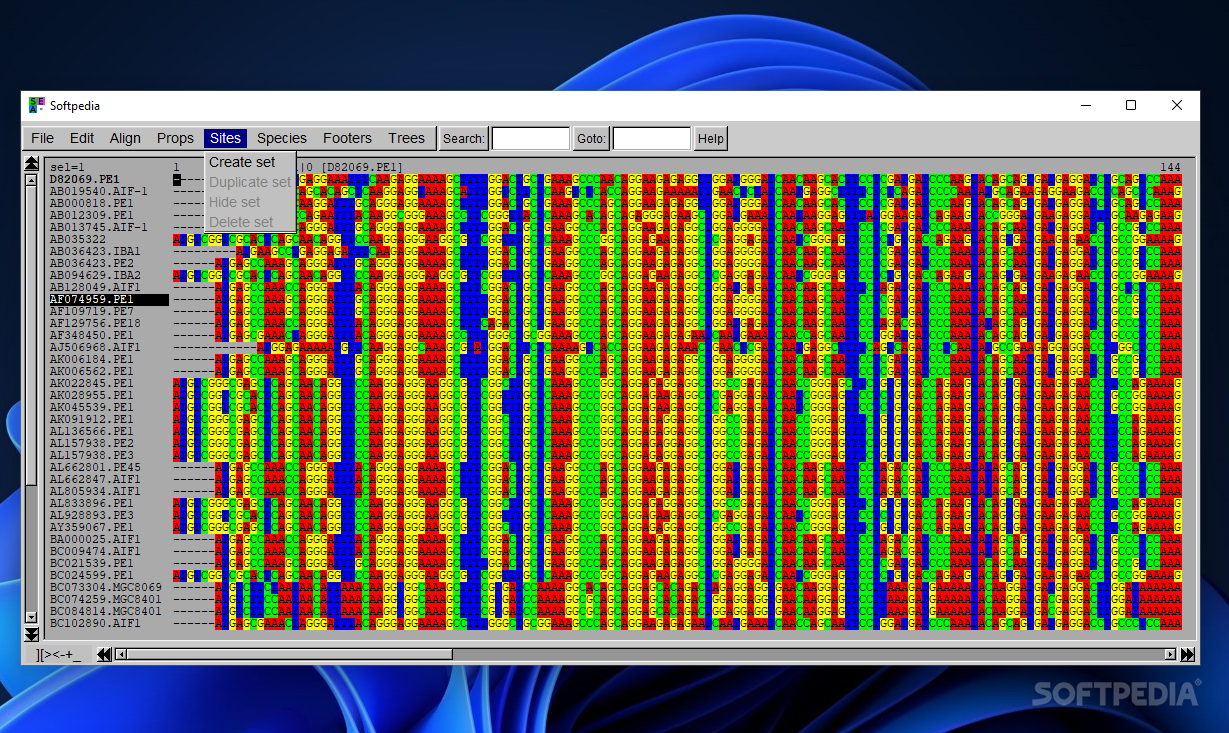
Seaview Version 5: A Multiplatform Software for Multiple Sequence Alignment, Molecular Phylogenetic Analyses, and Tree Reconciliation | SpringerLink
Identification of Putative Chemosensory Receptor Genes from the Athetis dissimilis Antennal Transcriptome | PLOS ONE

Biochemical properties and subcellular localization of six members of the HXK family in maize and its metabolic contribution to embryo germination | BMC Plant Biology | Full Text

Seaview Version 5: A Multiplatform Software for Multiple Sequence Alignment, Molecular Phylogenetic Analyses, and Tree Reconciliation | SpringerLink
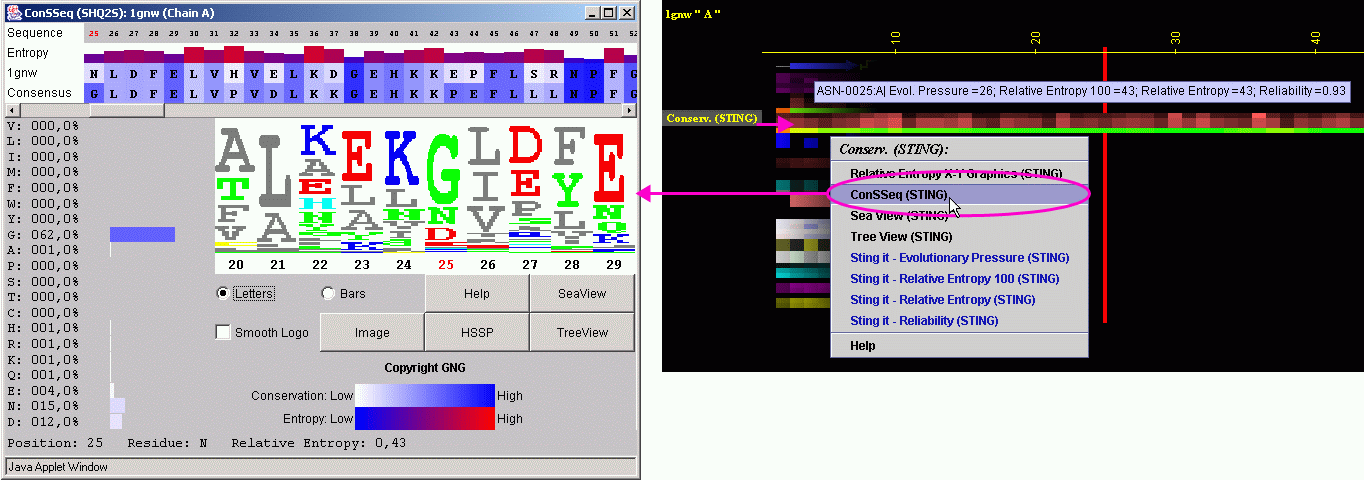
![PDF] SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny | Semantic Scholar PDF] SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1da2dc26a075432c92196131a5c1f0bd63f053f8/5-Figure3-1.png)
PDF] SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny | Semantic Scholar
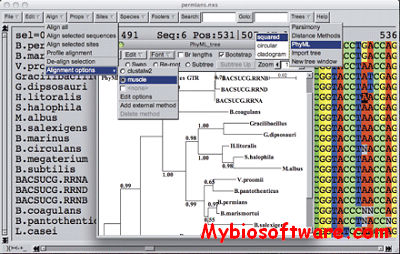
SeaView 4.7 – Sequence Alignment and Phylogenetic Tree Building – My Biosoftware – Bioinformatics Softwares Blog