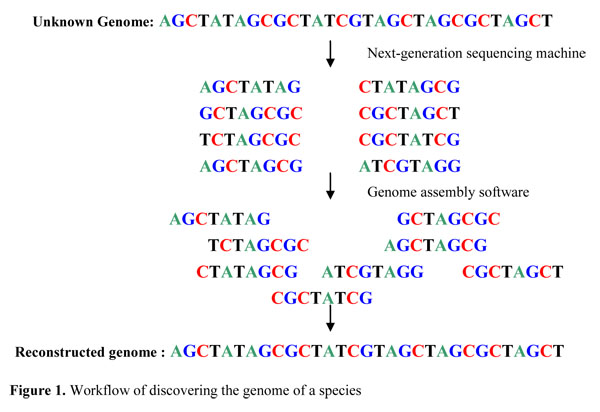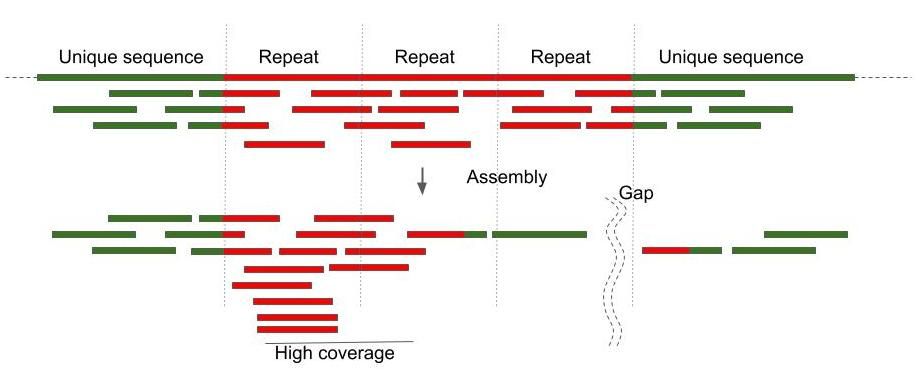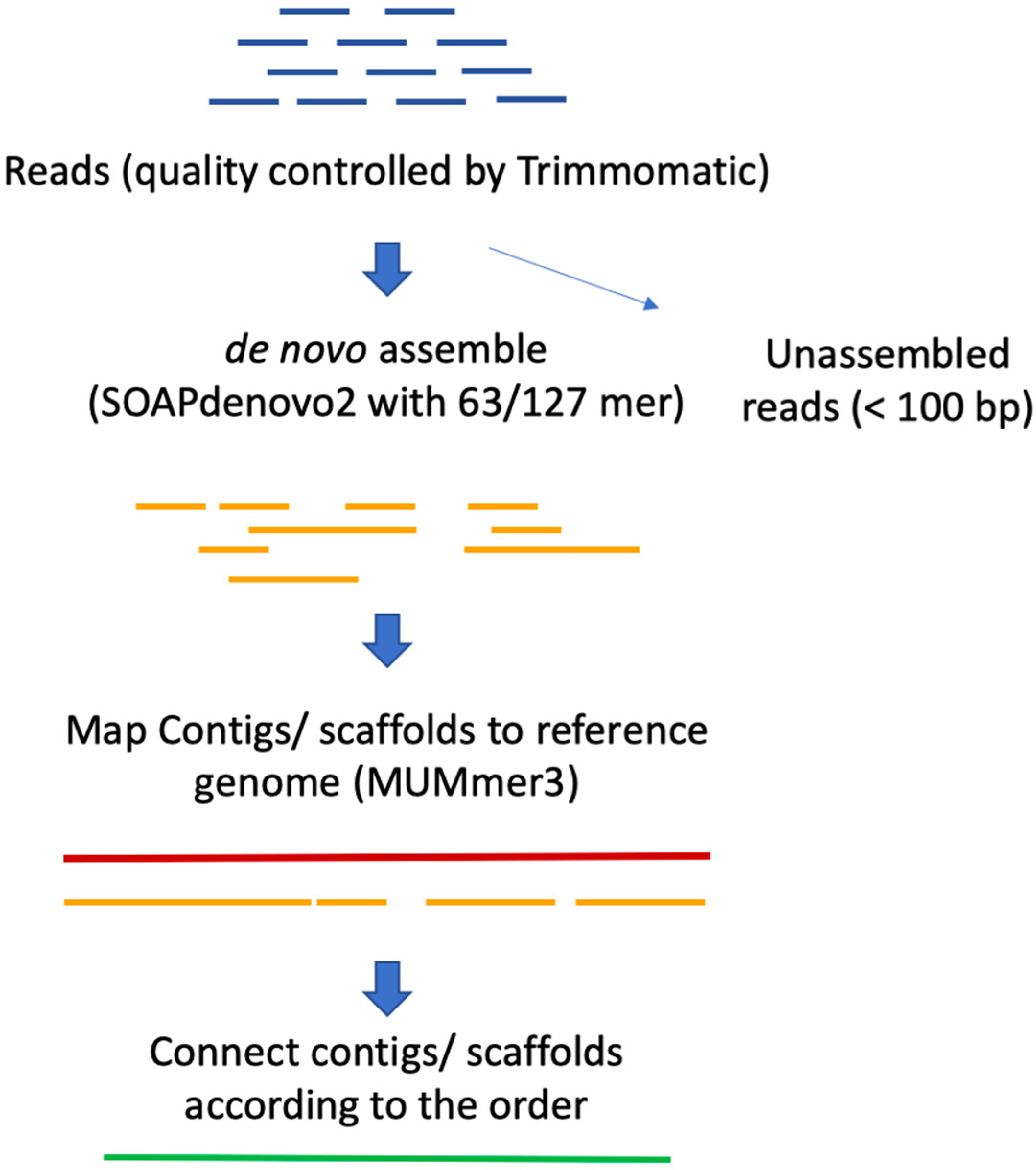
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1
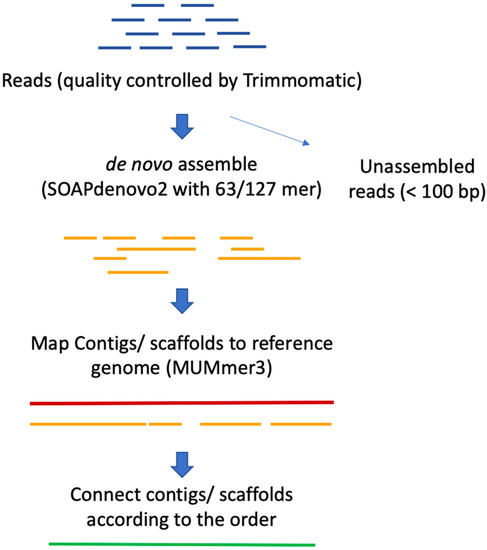
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1

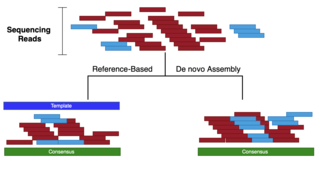
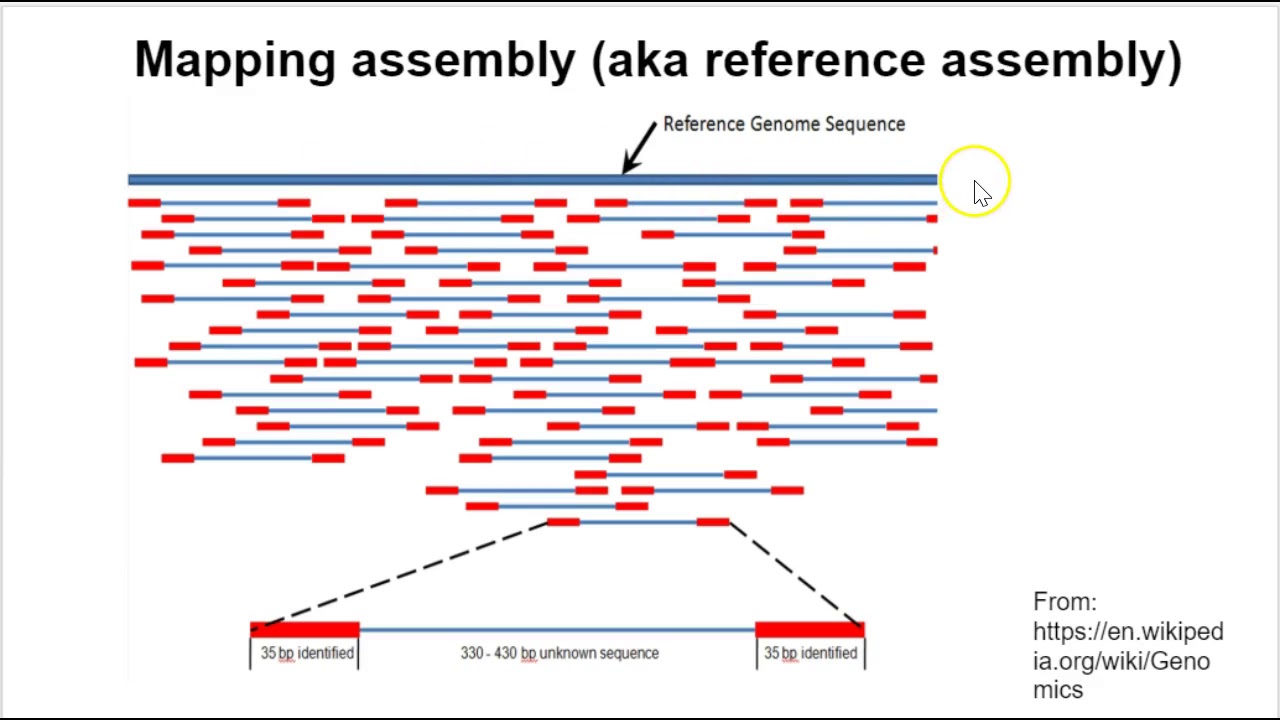
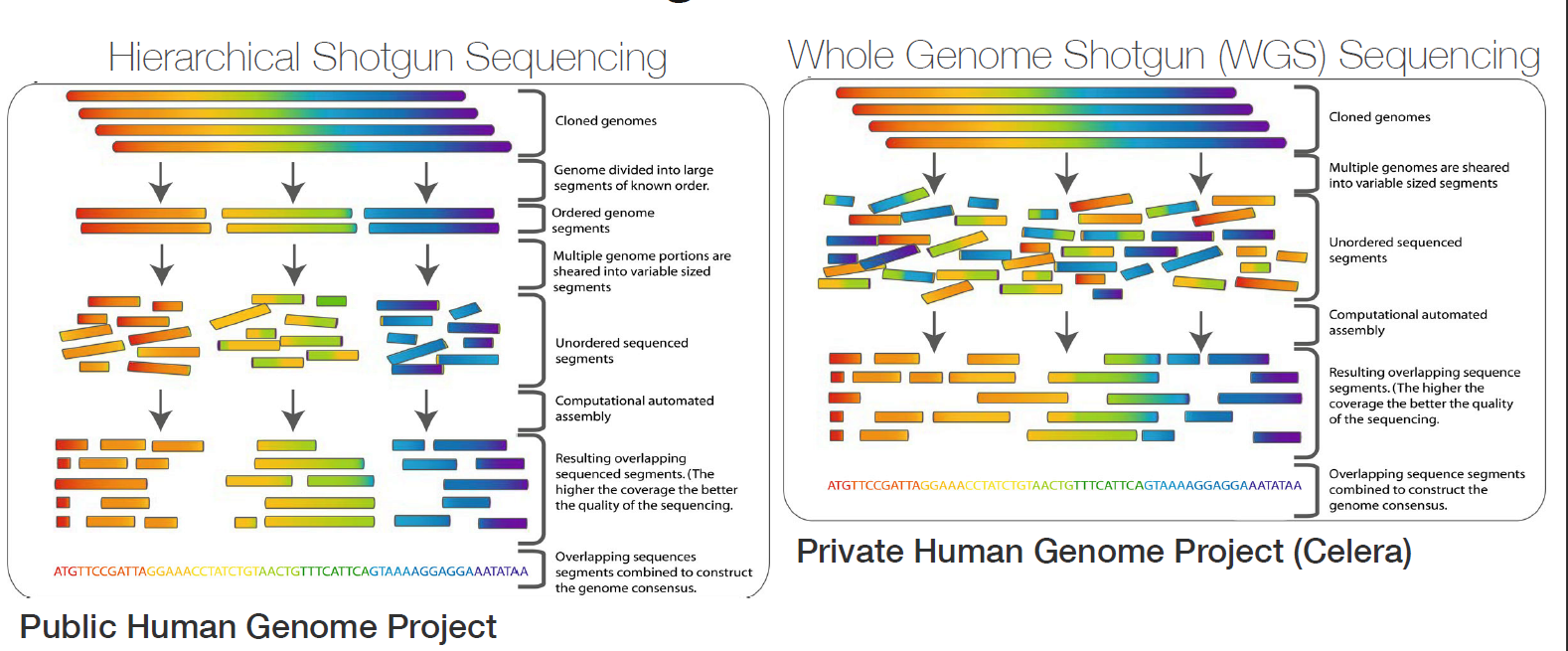
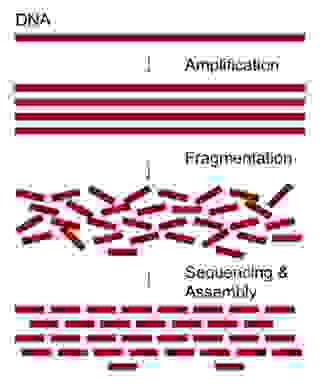
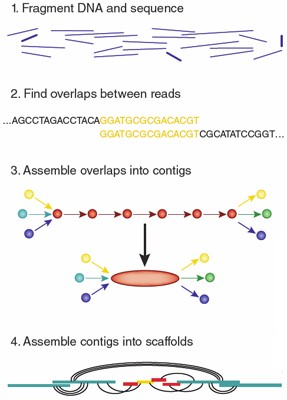
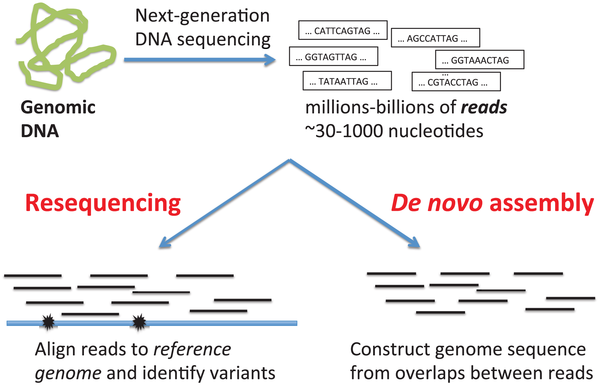
Sequencing from scratch: reference genomes and de novo sequence assembly – HudsonAlpha Institute for Biotechnology

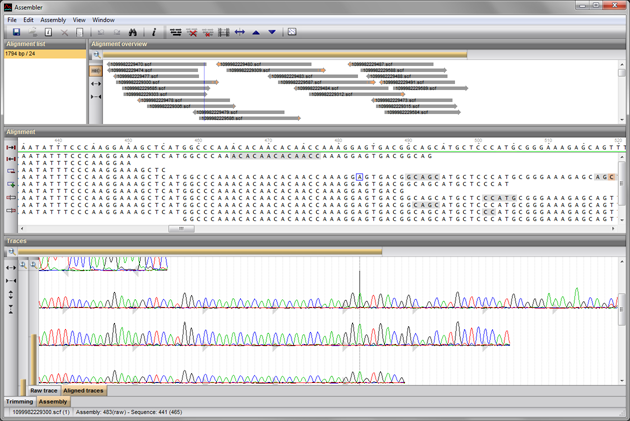
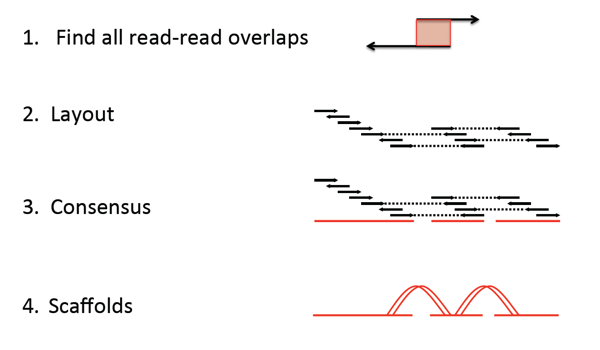
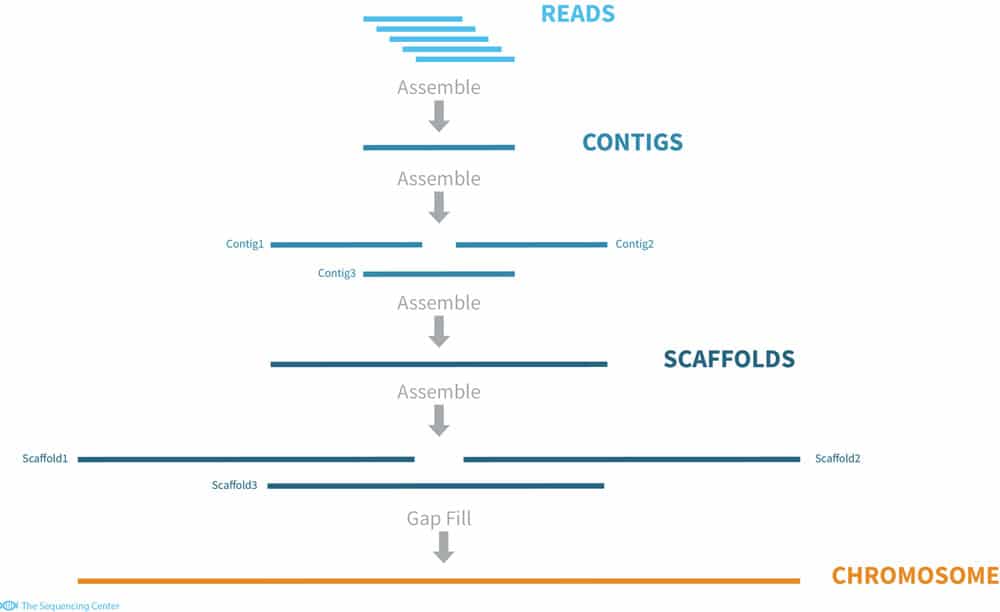
Benchmarking system for genome assembly and/or variant calling software (ikke tilgjengelig) - Institutt for informatikk

Sequencing from scratch: reference genomes and de novo sequence assembly – HudsonAlpha Institute for Biotechnology

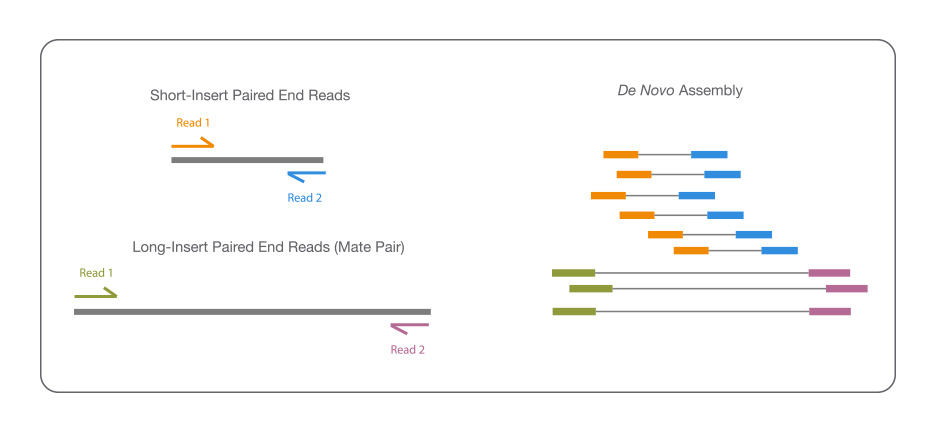
Tools and Strategies for Long-Read Sequencing and De Novo Assembly of Plant Genomes: Trends in Plant Science