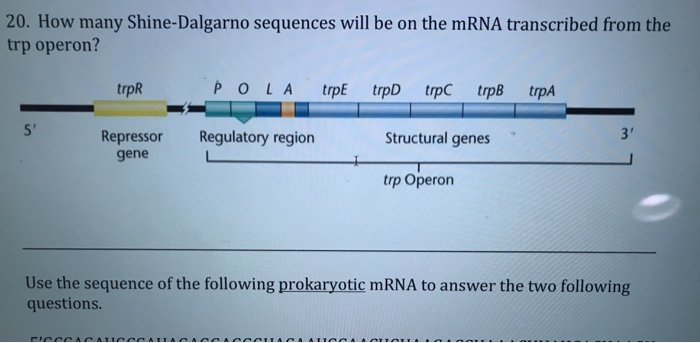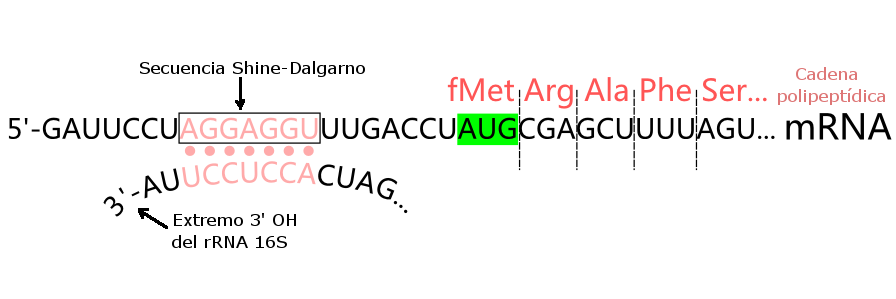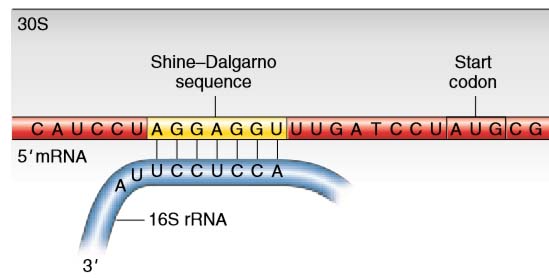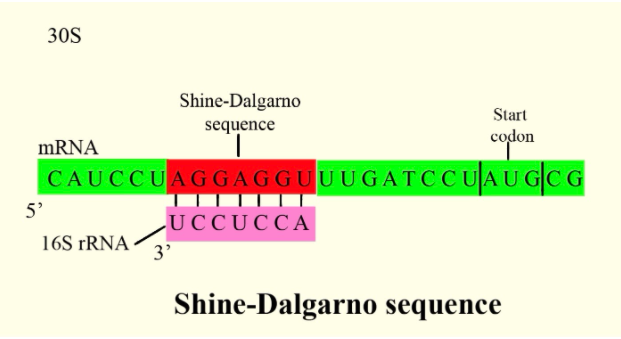
In prokaryotes, the ribosomes binding site on mRNA is called(a)Hogness sequence(b)Shine- Dalgarno sequence(c)Prinbow sequence(d)TATA- box

What is the Difference Between Shine Dalgarno and Kozak Sequence | Compare the Difference Between Similar Terms

Leveraging genome-wide datasets to quantify the functional role of the anti- Shine–Dalgarno sequence in regulating translation efficiency | Open Biology

Extending the Spacing between the Shine–Dalgarno Sequence and P-Site Codon Reduces the Rate of mRNA Translocation - ScienceDirect

The possible dual impacts of Shine-Dalgarno (SD) sequences on protein... | Download Scientific Diagram

Influences on gene expression in vivo by a Shine–Dalgarno sequence - Jin - 2006 - Molecular Microbiology - Wiley Online Library

Shine-Dalgarno Sequences Play an Essential Role in the Translation of Plastid mRNAs in Tobacco | Semantic Scholar

Twitter 上的 BASE mRNA Facility:"We were all taught the Shine-Dalgarno sequence but did you know it was discovered in Australia? The SD sequence is a ribosomal binding site in bacterial #mRNA that

Correlations between Shine-Dalgarno Sequences and Gene Features Such as Predicted Expression Levels and Operon Structures | Journal of Bacteriology

Influences on gene expression in vivo by a Shine–Dalgarno sequence - Jin - 2006 - Molecular Microbiology - Wiley Online Library