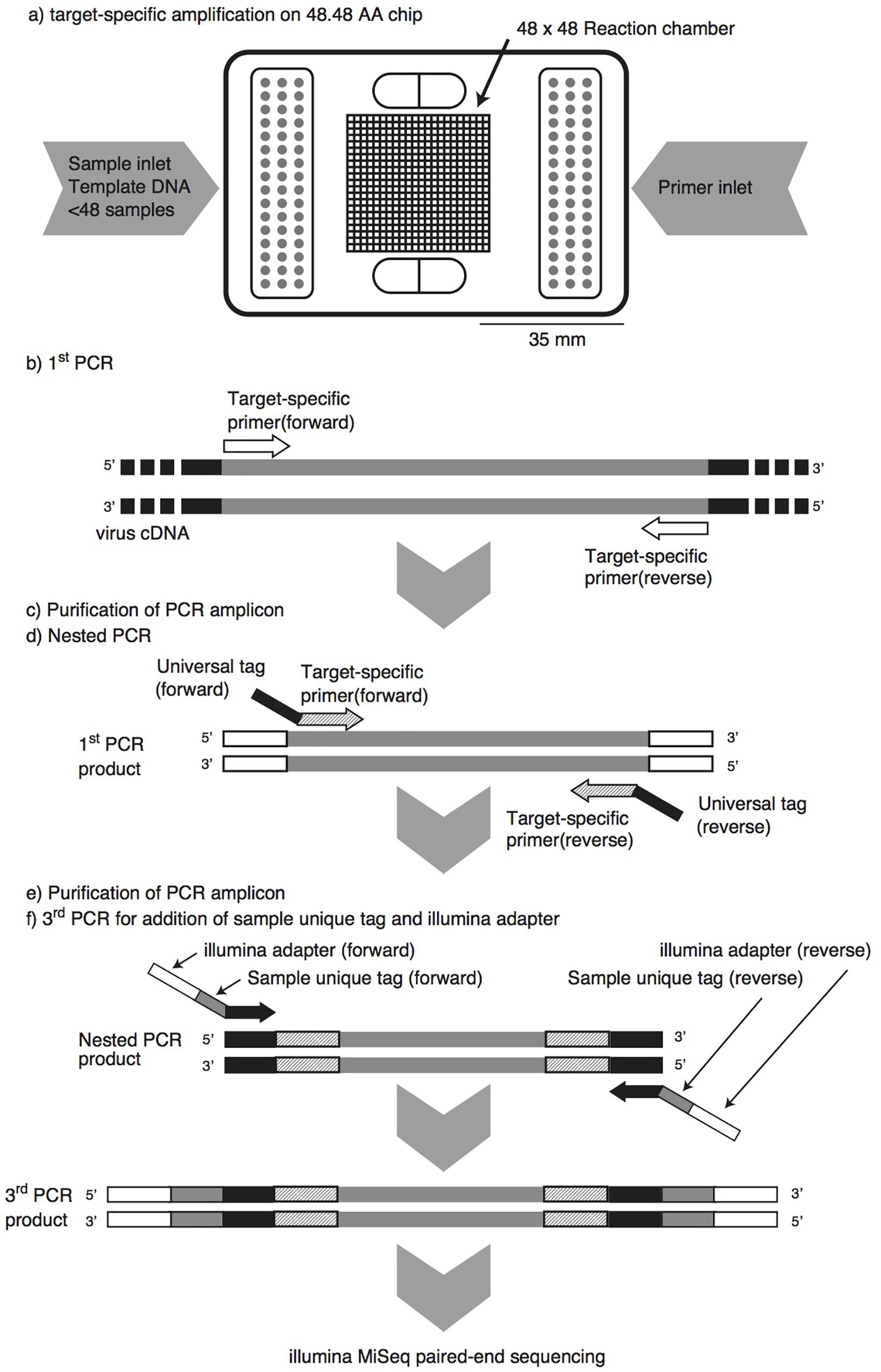
Frontiers | Microfluidic PCR Amplification and MiSeq Amplicon Sequencing Techniques for High-Throughput Detection and Genotyping of Human Pathogenic RNA Viruses in Human Feces, Sewage, and Oysters

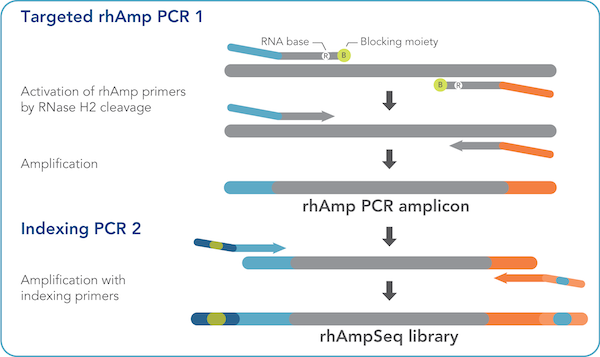
Integrated DNA Technologies on Twitter: "Whether you are investigating thousands of targets or a few, the fast and easy rhAmpSeq workflow generates NGS-ready amplicon libraries for deep, targeted resequencing. Explore rhAmpSeq amplicon
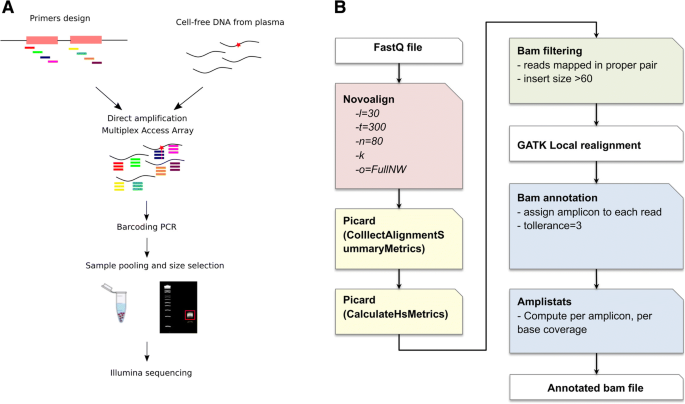
Next Generation-Targeted Amplicon Sequencing (NG-TAS): an optimised protocol and computational pipeline for cost-effective profiling of circulating tumour DNA | Genome Medicine | Full Text
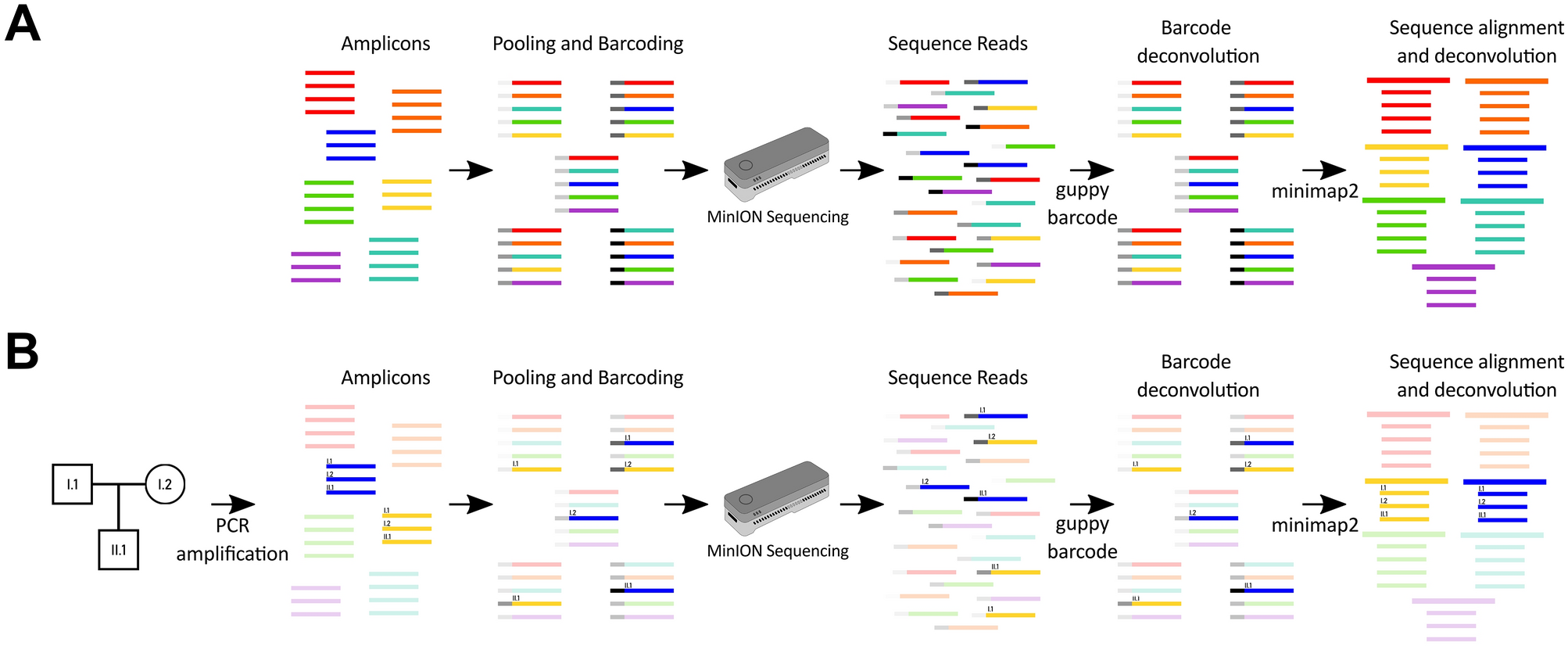
Proof of concept for multiplex amplicon sequencing for mutation identification using the MinION nanopore sequencer | Scientific Reports
Common enrichment options for targeted sequencing. Using amplicon-based... | Download Scientific Diagram
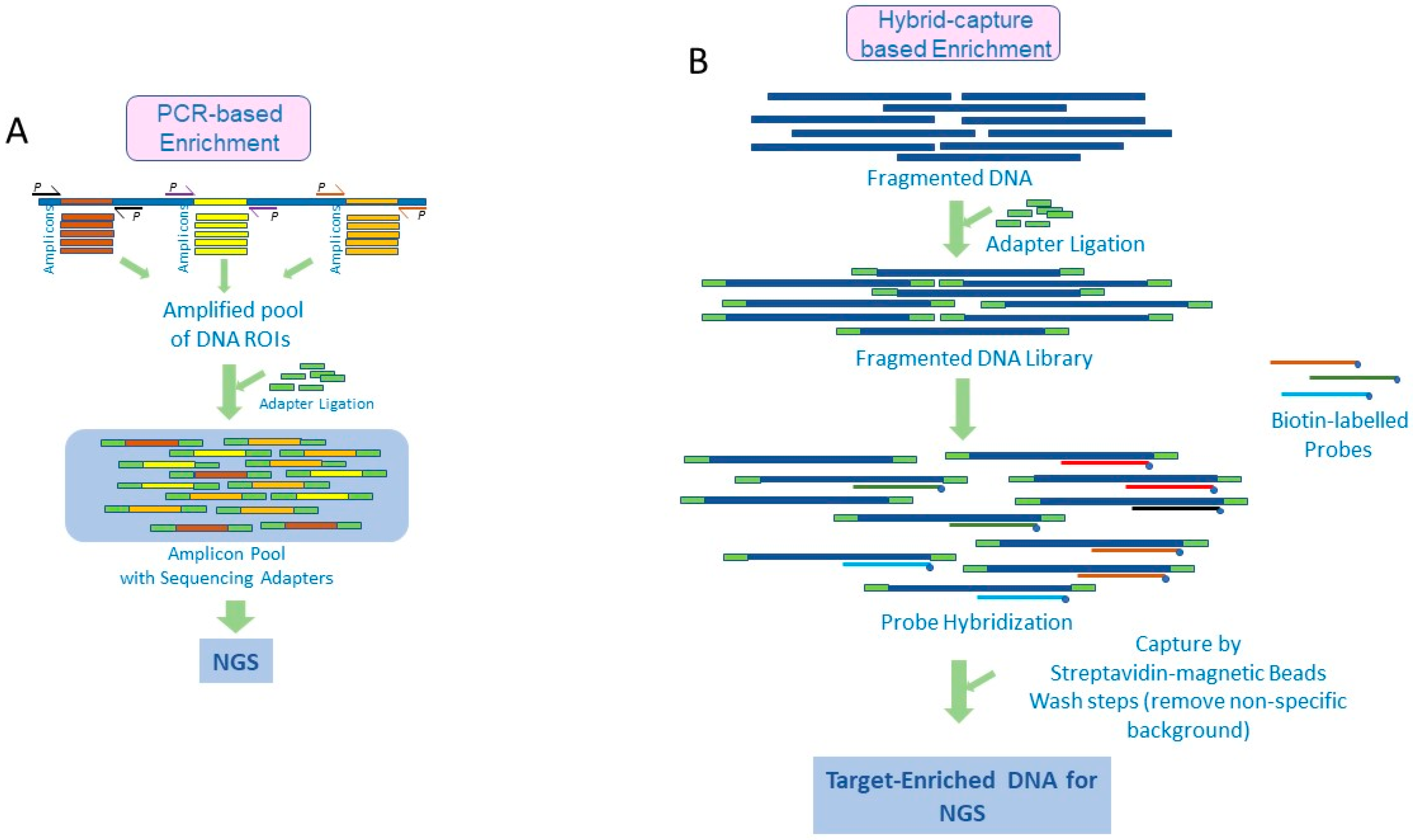
Diagnostics | Free Full-Text | Target Enrichment Approaches for Next-Generation Sequencing Applications in Oncology

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems

Targeted Amplicon Sequencing for Single-Nucleotide-Polymorphism Genotyping of Attaching and Effacing Escherichia coli O26:H11 Cattle Strains via a High-Throughput Library Preparation Technique | Applied and Environmental Microbiology

Direct comparison of RT-ddPCR and targeted amplicon sequencing for SARS-CoV-2 mutation monitoring in wastewater - ScienceDirect
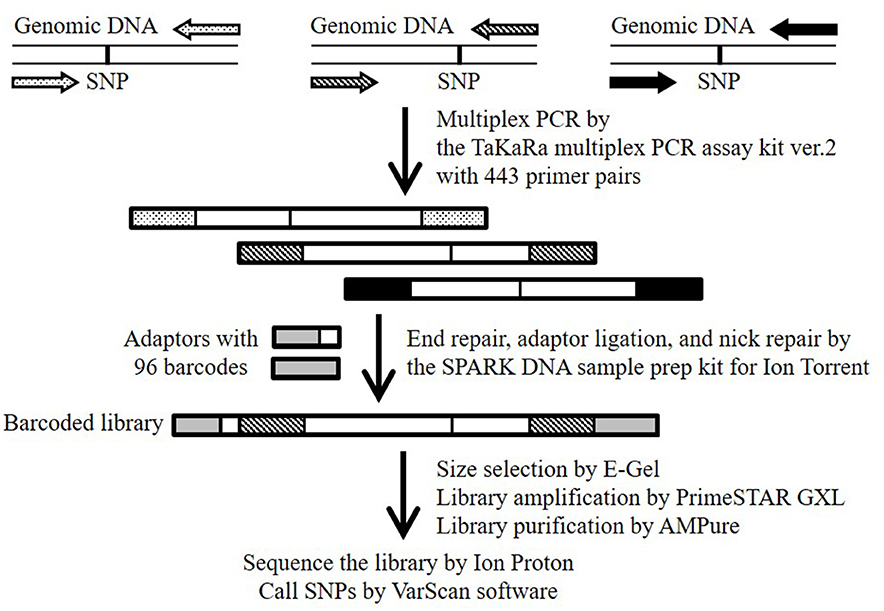
Frontiers | Multiplex PCR Targeted Amplicon Sequencing (MTA-Seq): Simple, Flexible, and Versatile SNP Genotyping by Highly Multiplexed PCR Amplicon Sequencing




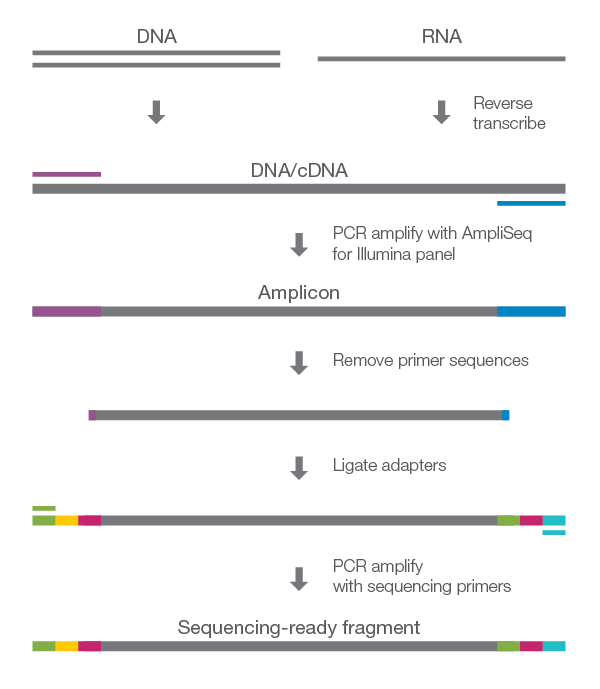
![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)