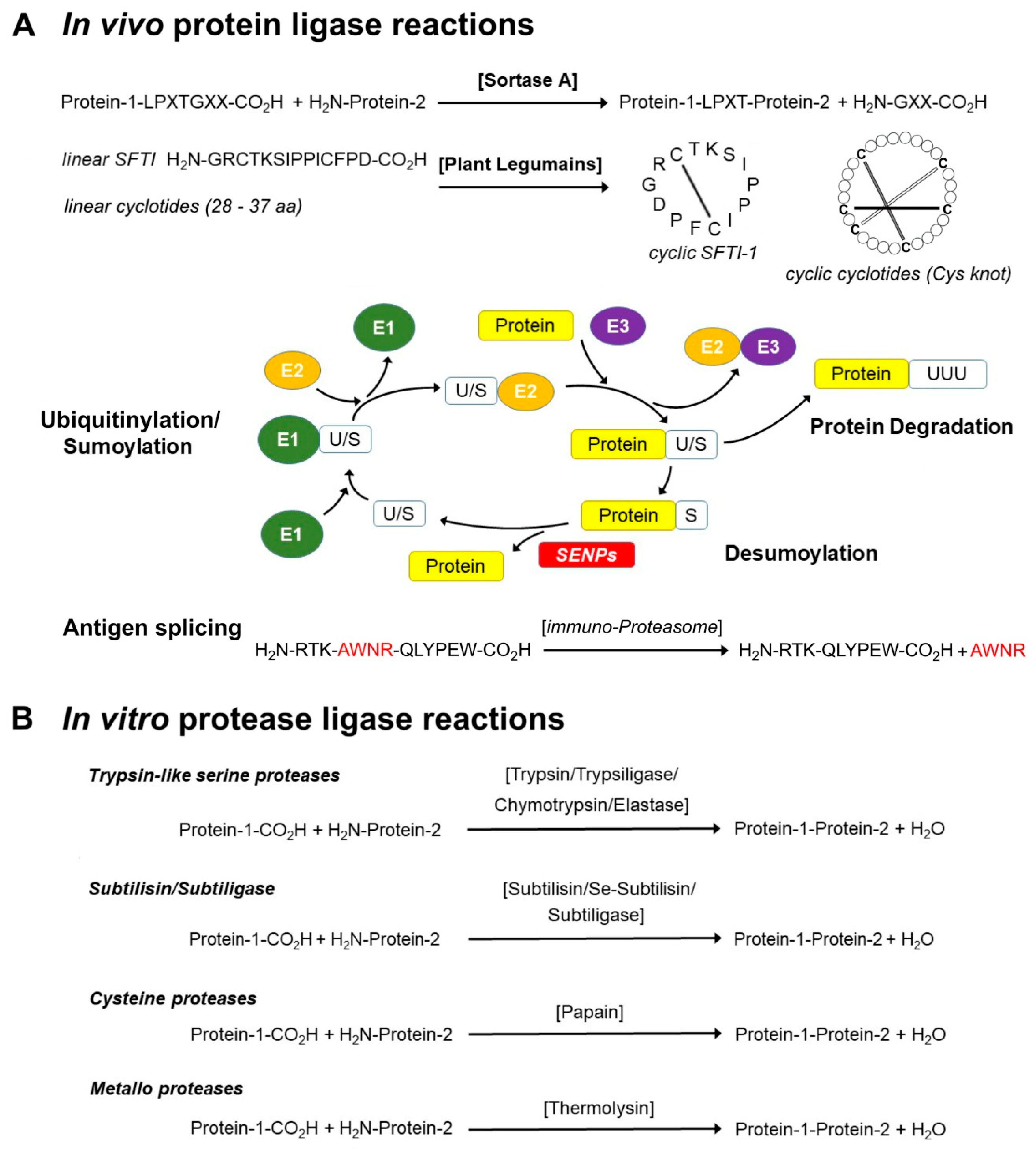
Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal

Prediction of Missed Proteolytic Cleavages for the Selection of Surrogate Peptides for Quantitative Proteomics | OMICS: A Journal of Integrative Biology
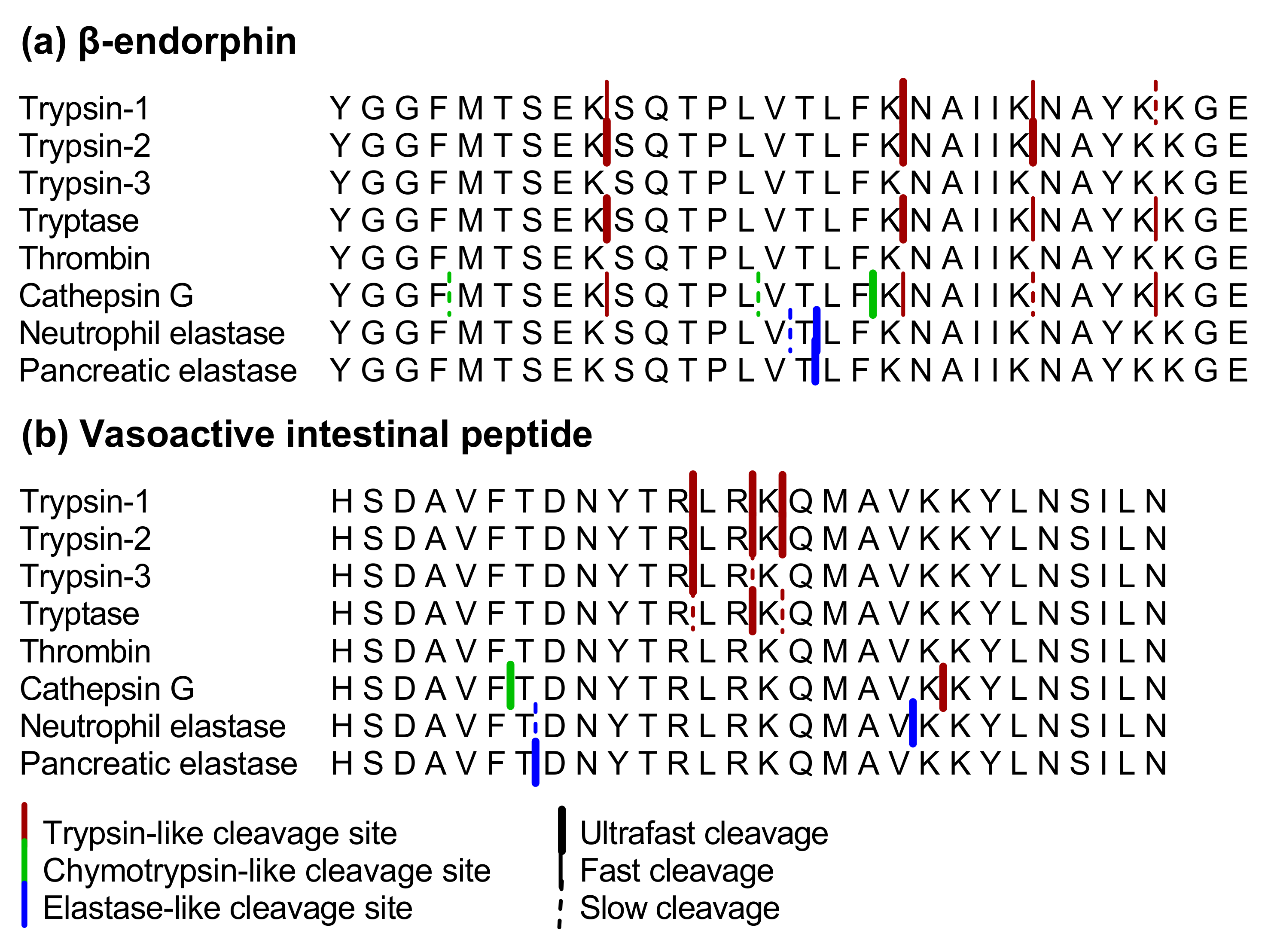
IJMS | Free Full-Text | Proteolytic Cleavage of Bioactive Peptides and Protease-Activated Receptors in Acute and Post-Colitis

Marked difference in efficiency of the digestive enzymes pepsin, trypsin, chymotrypsin, and pancreatic elastase to cleave tightly folded proteins

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal

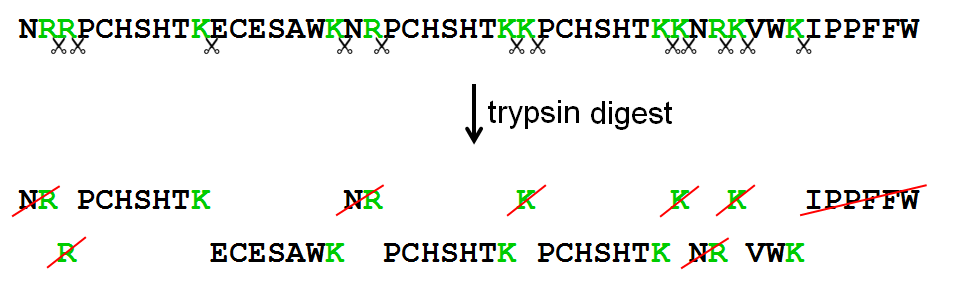
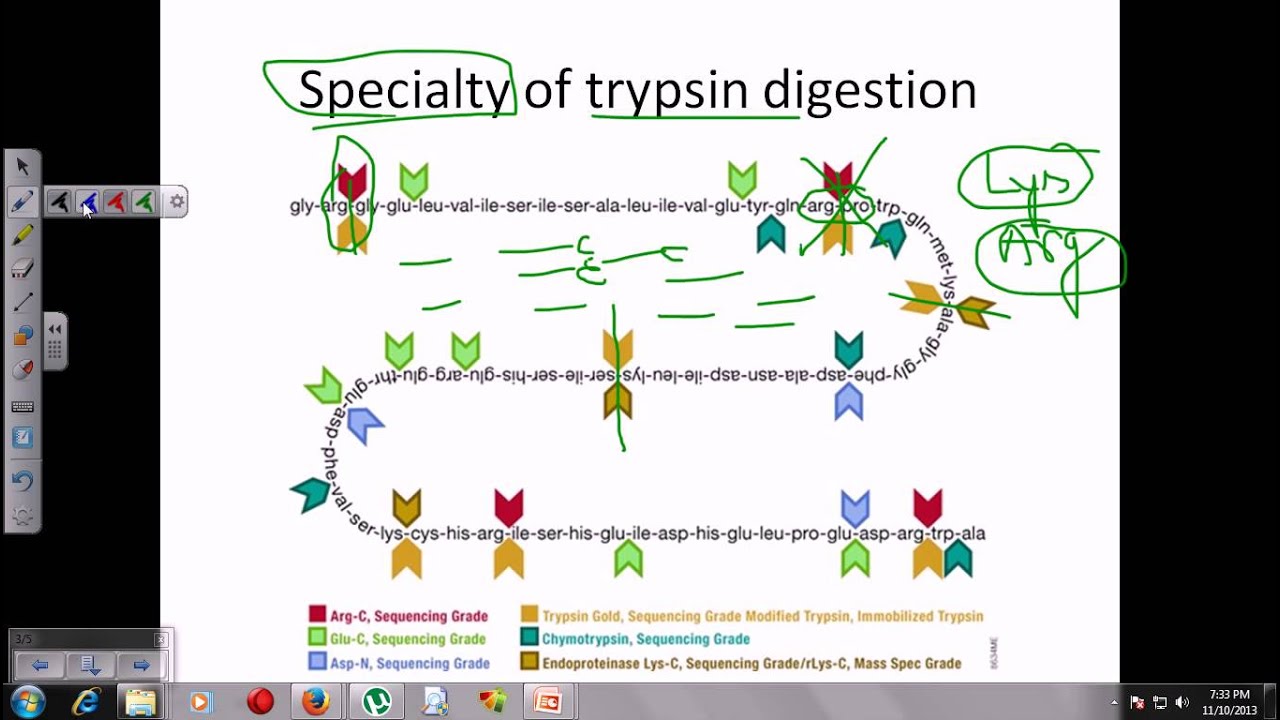
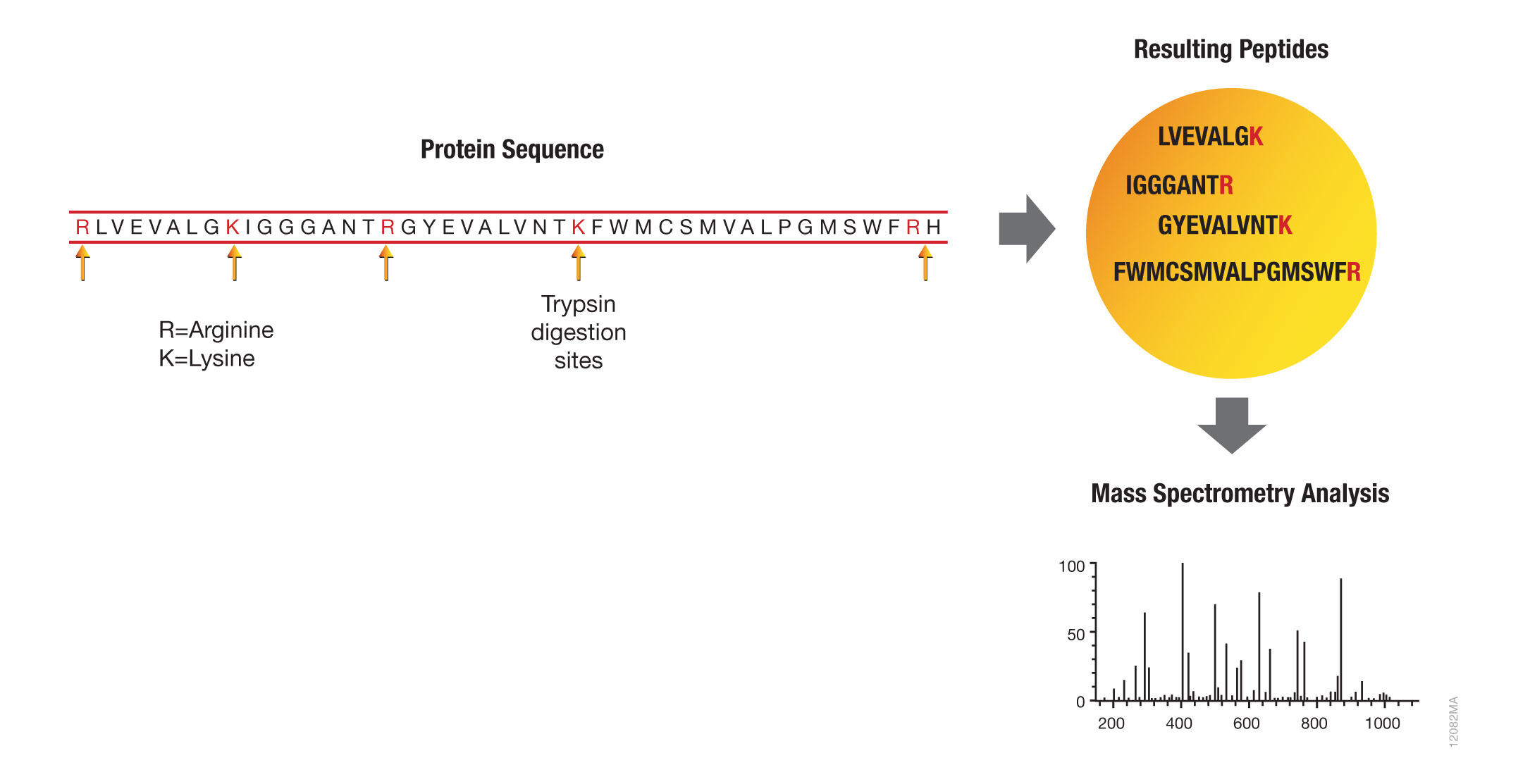
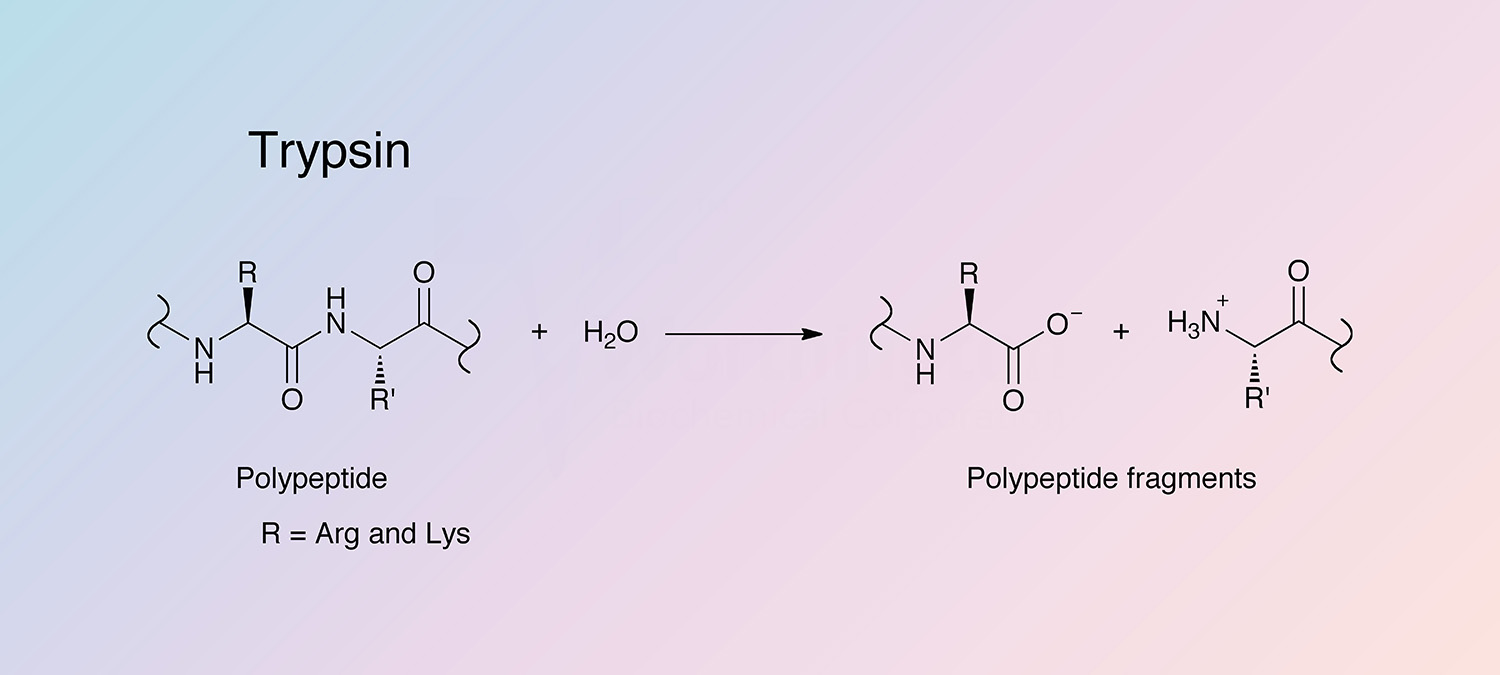
Schematic representation of the major trypsin cleavage sites and one... | Download Scientific Diagram
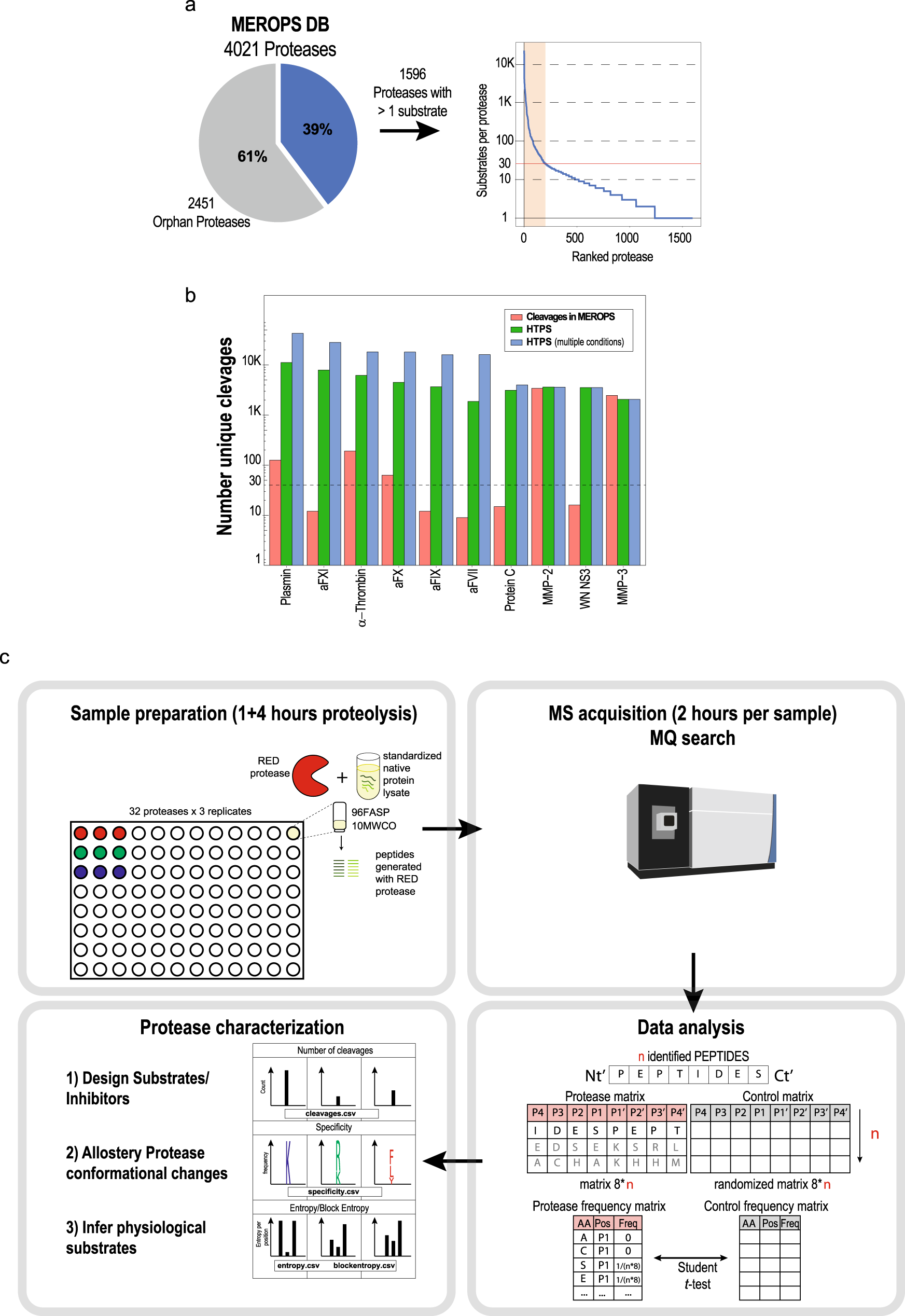
Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen | Nature Communications

Insight into Trypsin Miscleavage: Comparison of Kinetic Constants of Problematic Peptide Sequences | Analytical Chemistry

Amino acid sequence specifications for proteolytic cleavage by trypsin,... | Download Scientific Diagram
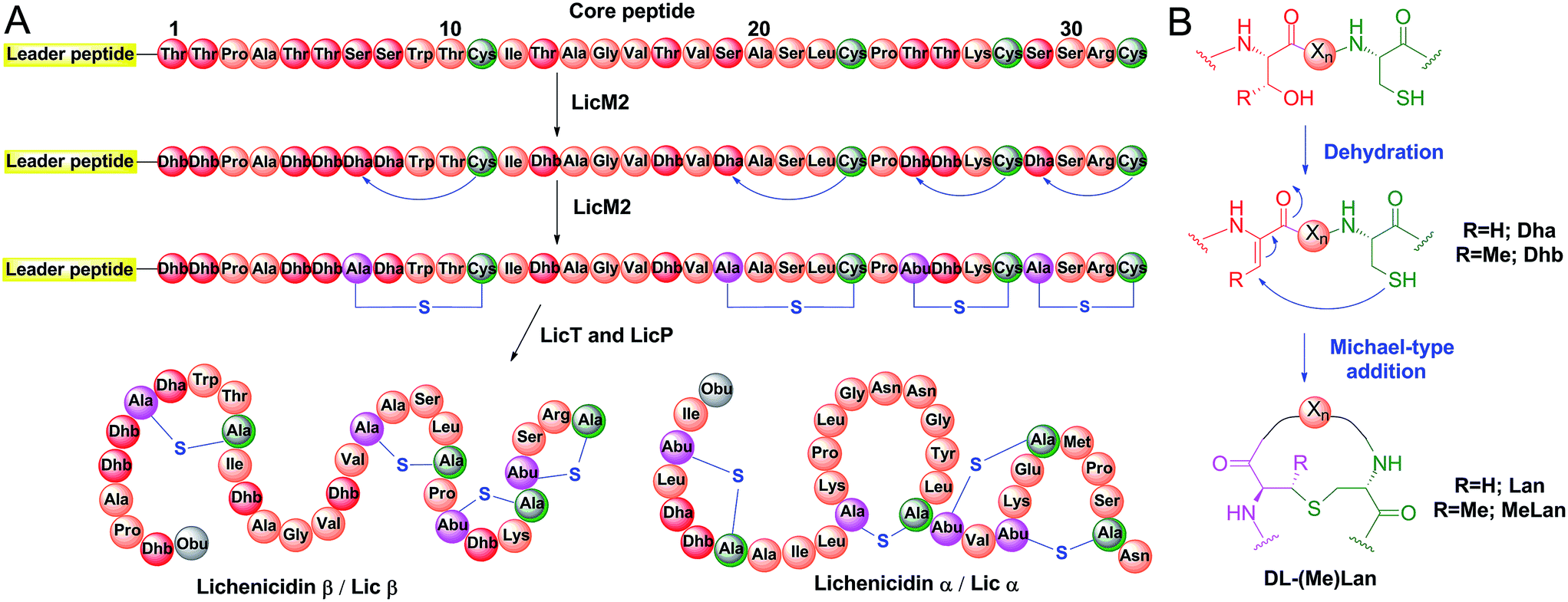
Applications of the class II lanthipeptide protease LicP for sequence-specific, traceless peptide bond cleavage - Chemical Science (RSC Publishing) DOI:10.1039/C5SC02329G

Proteomic Identification of protease Cleavage Sites (PICS) workflow.... | Download Scientific Diagram

Global Sequencing of Proteolytic Cleavage Sites in Apoptosis by Specific Labeling of Protein N Termini: Cell