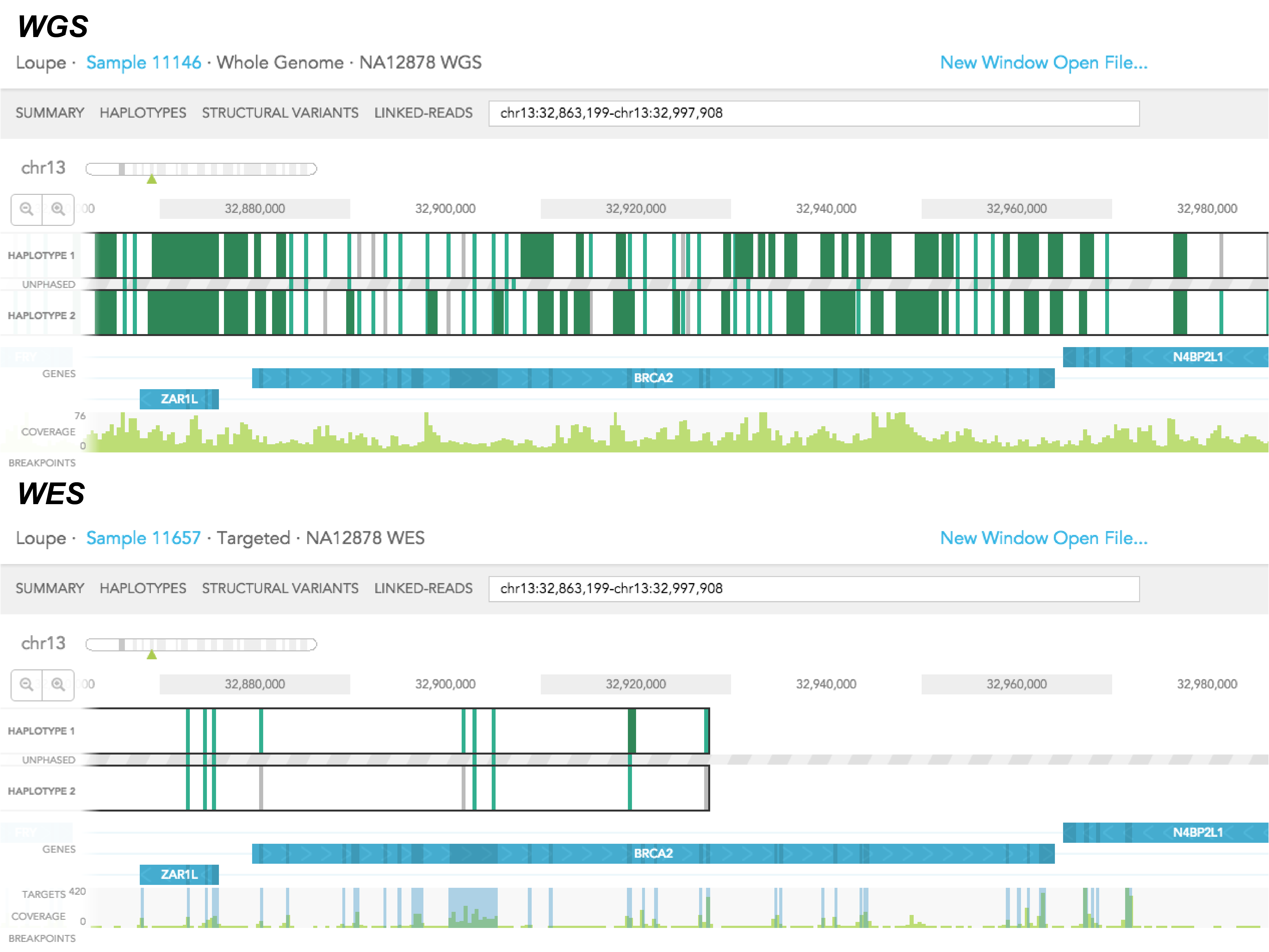
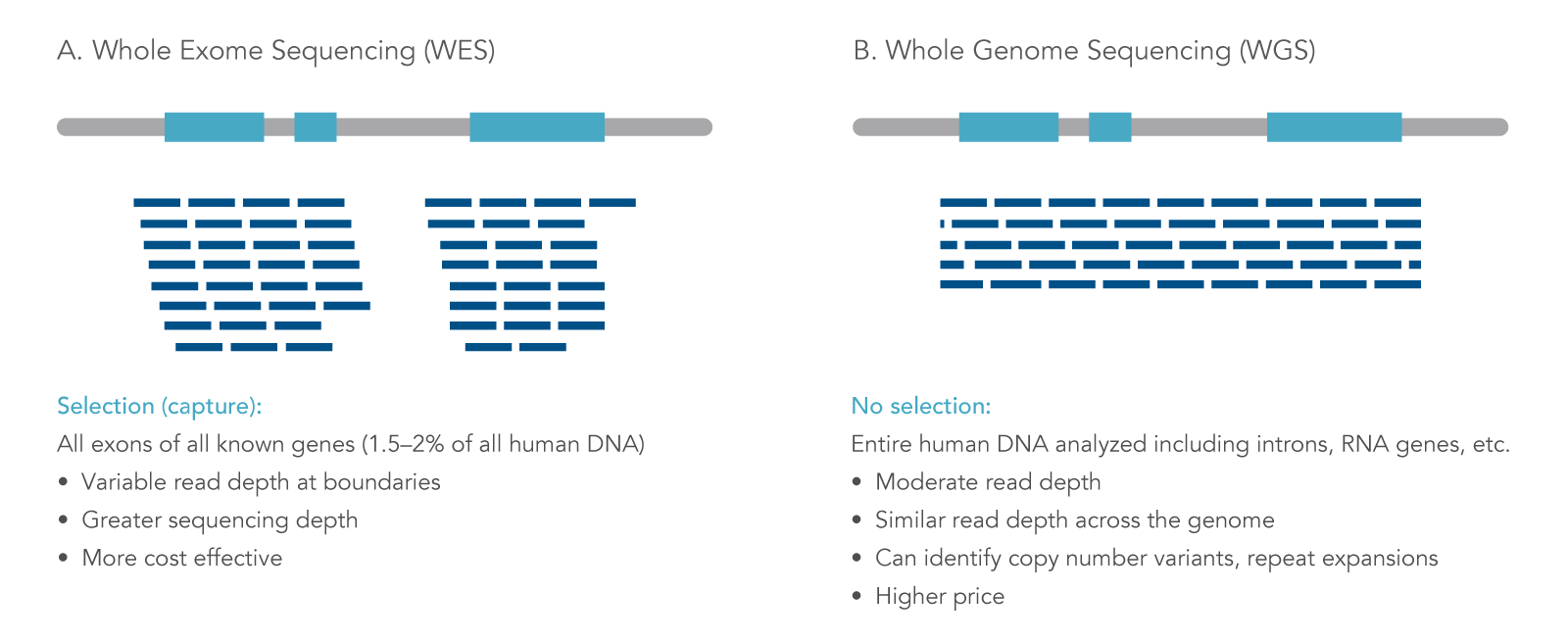
Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants | PNAS
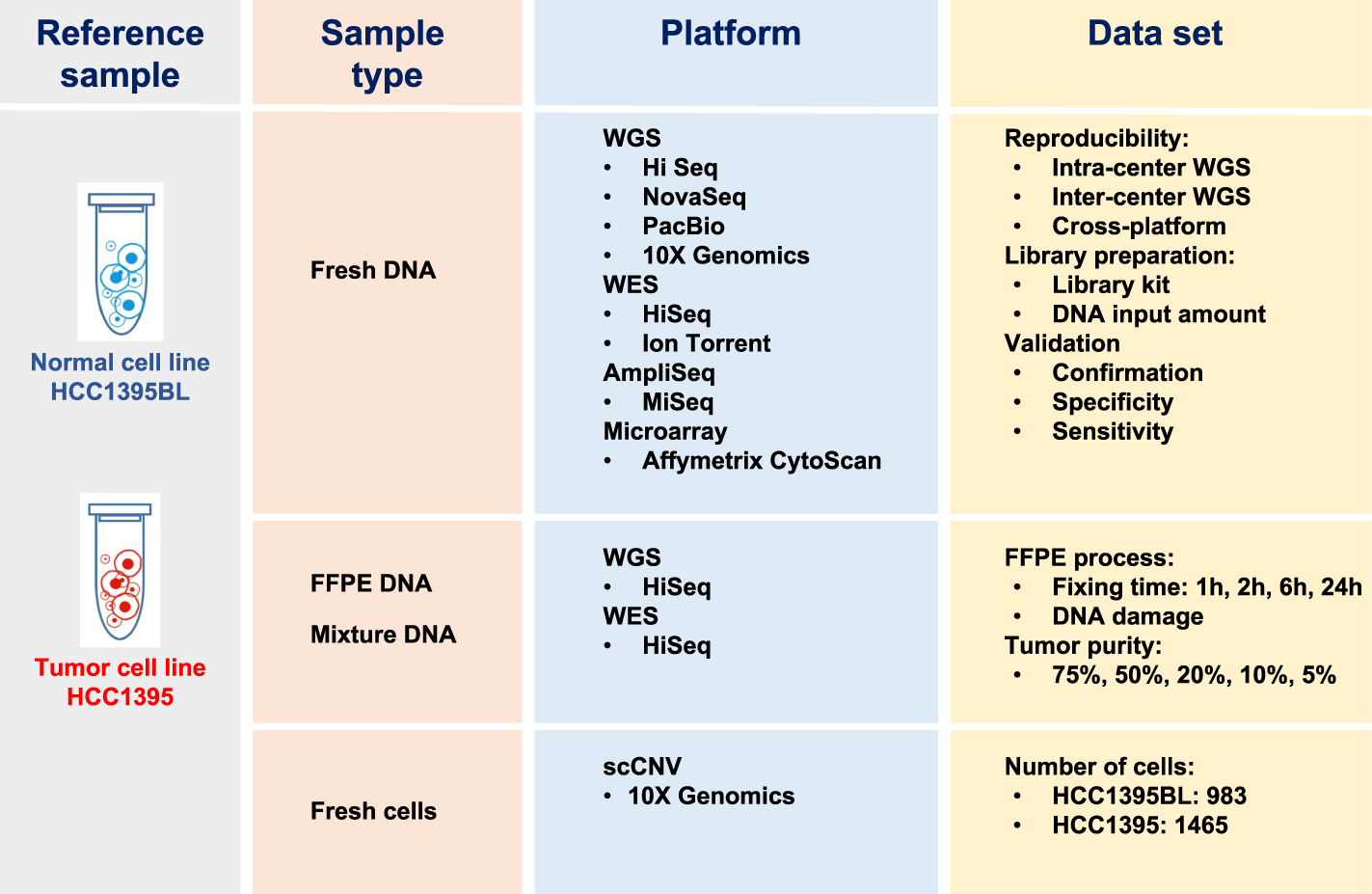
Whole genome and exome sequencing reference datasets from a multi-center and cross-platform benchmark study | Scientific Data

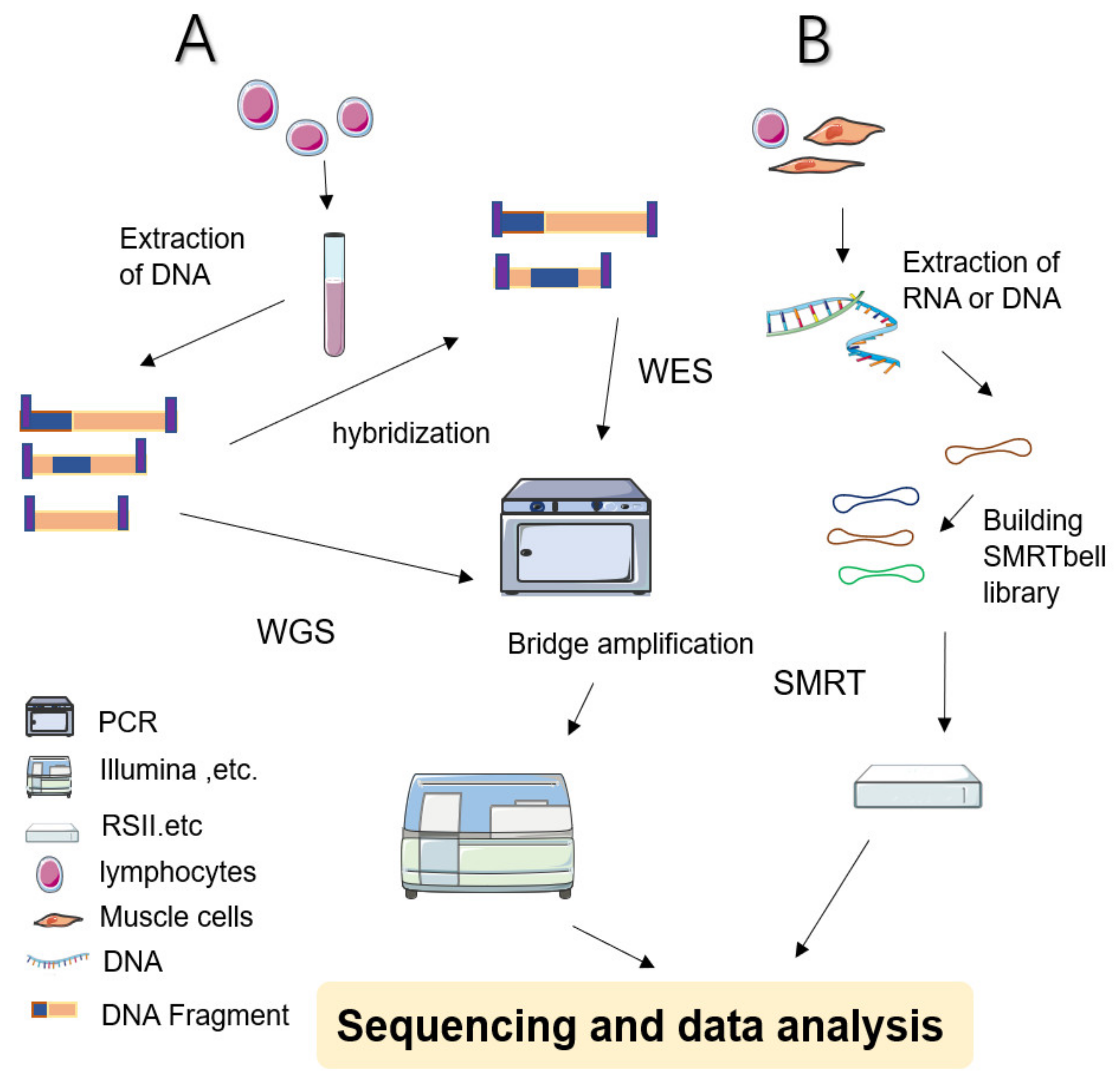
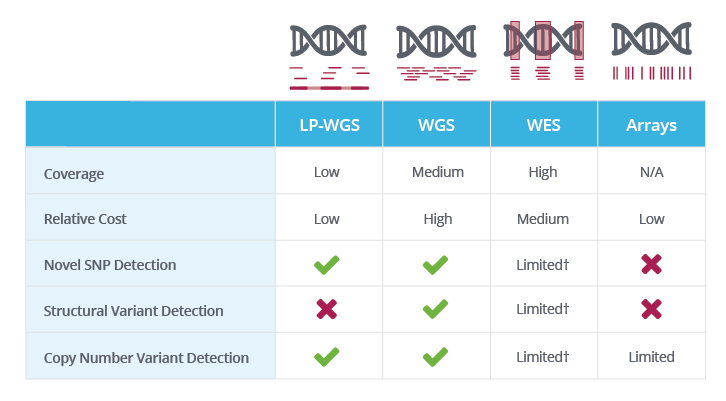
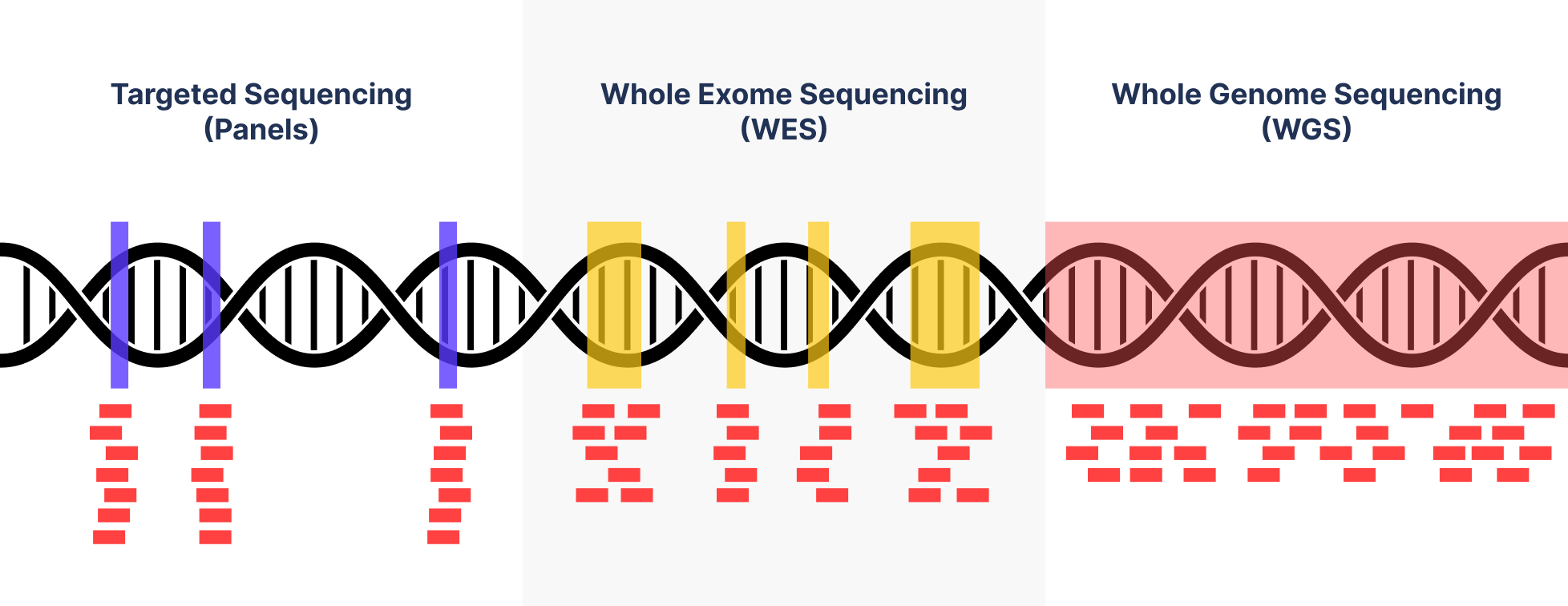
Whole genome sequencing (WGS), whole exome sequencing (WES) and clinical exome sequencing (CES) in patient care
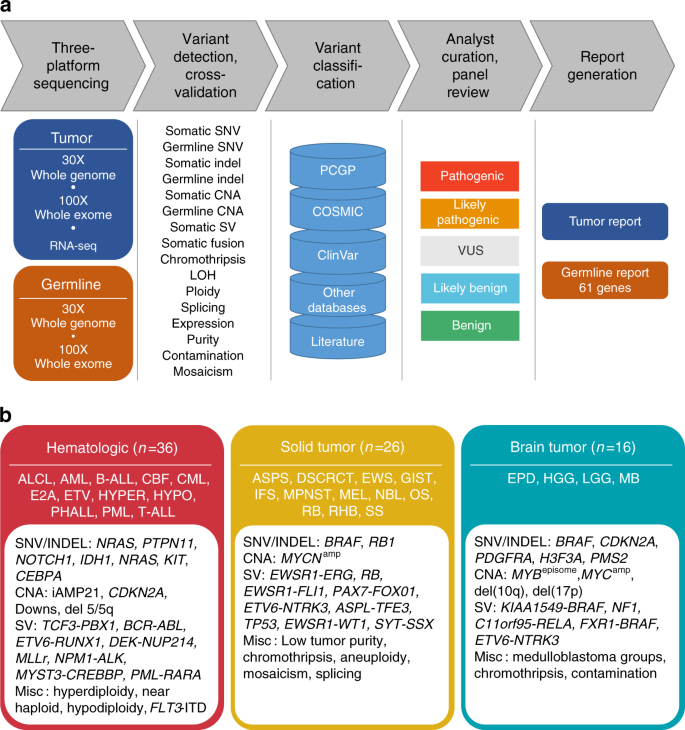
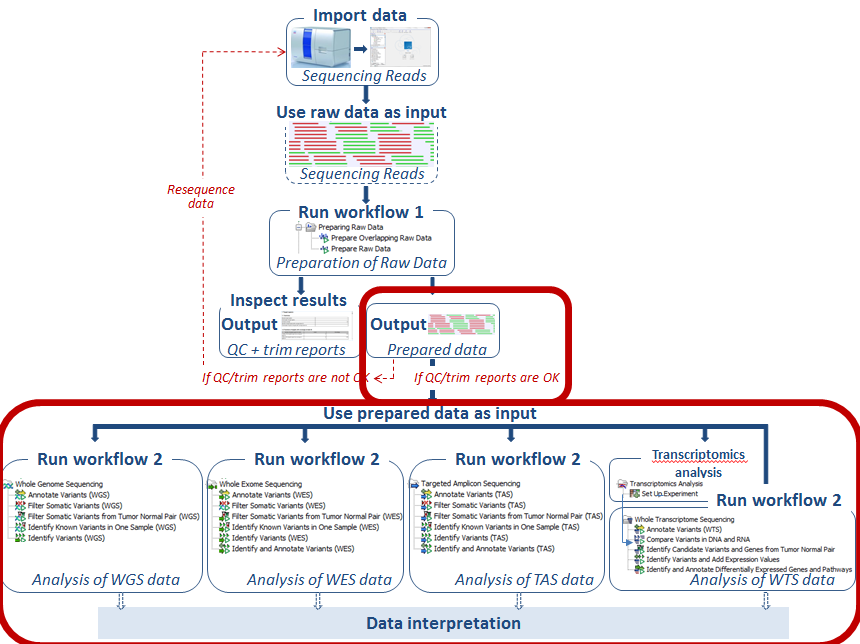
Clinical cancer genomic profiling by three-platform sequencing of whole genome, whole exome and transcriptome | Nature Communications

Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage | Scientific Reports